Team:Tsinghua/ELSI
From 2009.igem.org
GuoQiangChen (Talk | contribs) (→Safety Issues Q & A) |
GuoQiangChen (Talk | contribs) (→Innovation and Human Practice) |
||
| (29 intermediate revisions not shown) | |||
| Line 6: | Line 6: | ||
=Terminology= | =Terminology= | ||
We termed our synthesized targeted gene therapy vector "GenSniper" to describe the cell specificity based on the biobrick targeted approach. Unlike the traditional gene therapy vector able to infect a variety of cells, GenSniper will be capable of specifically infecting one type of cells, just as snipers specifically shoot one target at high resolution. | We termed our synthesized targeted gene therapy vector "GenSniper" to describe the cell specificity based on the biobrick targeted approach. Unlike the traditional gene therapy vector able to infect a variety of cells, GenSniper will be capable of specifically infecting one type of cells, just as snipers specifically shoot one target at high resolution. | ||
| - | [[Image:terminology1.png|400px|center|thumb|A sniper in the movie Enemy at Gates]] | + | [[Image:terminology1.png|400px|center|thumb|A sniper in the movie Enemy at Gates. We termed our gene therapy vector GenSniper because the design allows it to target specific cell precisely, just as a sniper does]] |
Apparently, other approaches (especially fusion proteins) have been reported for targeted gene therapy. As for our project, we aim at providing a standardized modification principle for targeted gene therapy, namely the introduction of Targeted BioBrick. With the cell-specific peptides changed by PCR, the synthesized GenSniper will be able to target different type of cells. Targeted BioBrick can also be incorperated onto different proteins for different function into a specific type of cells. | Apparently, other approaches (especially fusion proteins) have been reported for targeted gene therapy. As for our project, we aim at providing a standardized modification principle for targeted gene therapy, namely the introduction of Targeted BioBrick. With the cell-specific peptides changed by PCR, the synthesized GenSniper will be able to target different type of cells. Targeted BioBrick can also be incorperated onto different proteins for different function into a specific type of cells. | ||
| Line 19: | Line 19: | ||
=Implication on Synthetic Biology= | =Implication on Synthetic Biology= | ||
==Synthetic Genome== | ==Synthetic Genome== | ||
| - | |||
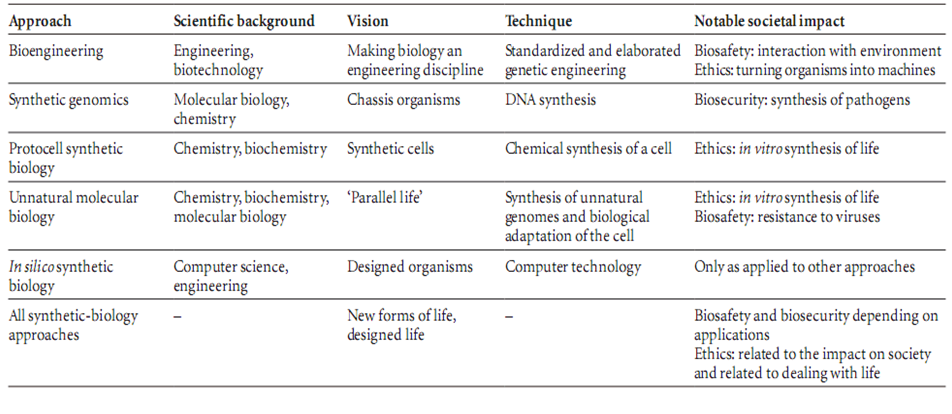
[[Image:synthetic biology approaches.png|800px|center]] | [[Image:synthetic biology approaches.png|800px|center]] | ||
| + | |||
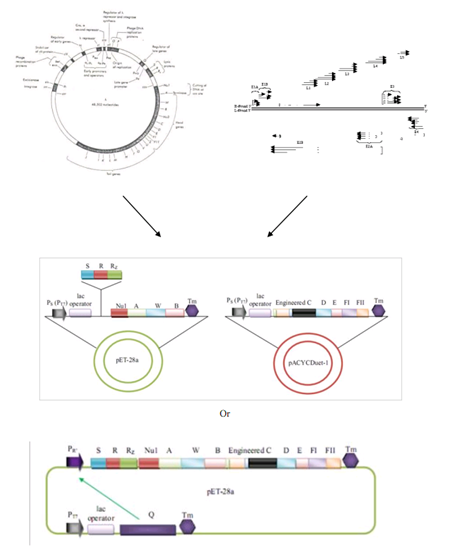
| + | As suggested by the project title, our Project is equivalent to apply synthetic biology concepts and standards at the genomic level, constructing a genome which is neither adenovirus nor bacteriophage lambda genome. This synthetic genome, however, is capable of producing standardized and targeted gene therapy vectors for human clinical practice. Also, we use the abstraction principle in our design of this genome in order to make the synthetic gene therapy vector easy to be further modified, improved and industrialized. This project implicates the development of synthetic biology come to a higher level of living organism, the genome, which meets the trends of synthetic biology innovation. | ||
| + | [[Image:synGenome.png|950px|center|thumb|SynGenome approach]] | ||
==Synthetic Biology in Gene Therapy== | ==Synthetic Biology in Gene Therapy== | ||
| Line 27: | Line 29: | ||
==From Shotgun to Sniper== | ==From Shotgun to Sniper== | ||
Previous attempts on targeted gene delivery may involve the fusion between the gene therapy vector and a receptor protein of specific cell type(s). Before the standardized approach to select specific receptors for any given types of cells, those attempts may not easy to be reproduced. Our project suggests a standardized way of targeted peptide modification. For example, if you change your bullet (cell-specific peptides generated by phage display), you will be able to, in theory, precisely target your desired gene to any given type of cells. Of course, the Targeted Biobrick proposal needs more revision to meet the industrialization and clinical uses. We hope this attempt can draw the attention of scientists working with gene therapy on the convenience of biobricks. | Previous attempts on targeted gene delivery may involve the fusion between the gene therapy vector and a receptor protein of specific cell type(s). Before the standardized approach to select specific receptors for any given types of cells, those attempts may not easy to be reproduced. Our project suggests a standardized way of targeted peptide modification. For example, if you change your bullet (cell-specific peptides generated by phage display), you will be able to, in theory, precisely target your desired gene to any given type of cells. Of course, the Targeted Biobrick proposal needs more revision to meet the industrialization and clinical uses. We hope this attempt can draw the attention of scientists working with gene therapy on the convenience of biobricks. | ||
| + | |||
| + | [[Image:trandition.png|center|700px|thumb|Tranditional gene therapy targeting]] | ||
| + | [[Image:innovation.png|center|700px|thumb|Gene therapy targeting by GenSniper]] | ||
==Targeted Biobrick Platform== | ==Targeted Biobrick Platform== | ||
| - | [[Image:ZFN.png| | + | [[Image:ZFN.png|800px|center|thumb|Zinc Finger Protein Platform]] |
Our defintion of Targeted Biobrick in gene therapy is quite similar with the booming area in zinc finger nuclease and zinc finger proteins. Attempts in zinc finger proteins do reflect a concept of modulization at the protein level, however the standardized principles of restriction site prefix or surfix have not been widely applied in the construction of zinc finger protein platform. | Our defintion of Targeted Biobrick in gene therapy is quite similar with the booming area in zinc finger nuclease and zinc finger proteins. Attempts in zinc finger proteins do reflect a concept of modulization at the protein level, however the standardized principles of restriction site prefix or surfix have not been widely applied in the construction of zinc finger protein platform. | ||
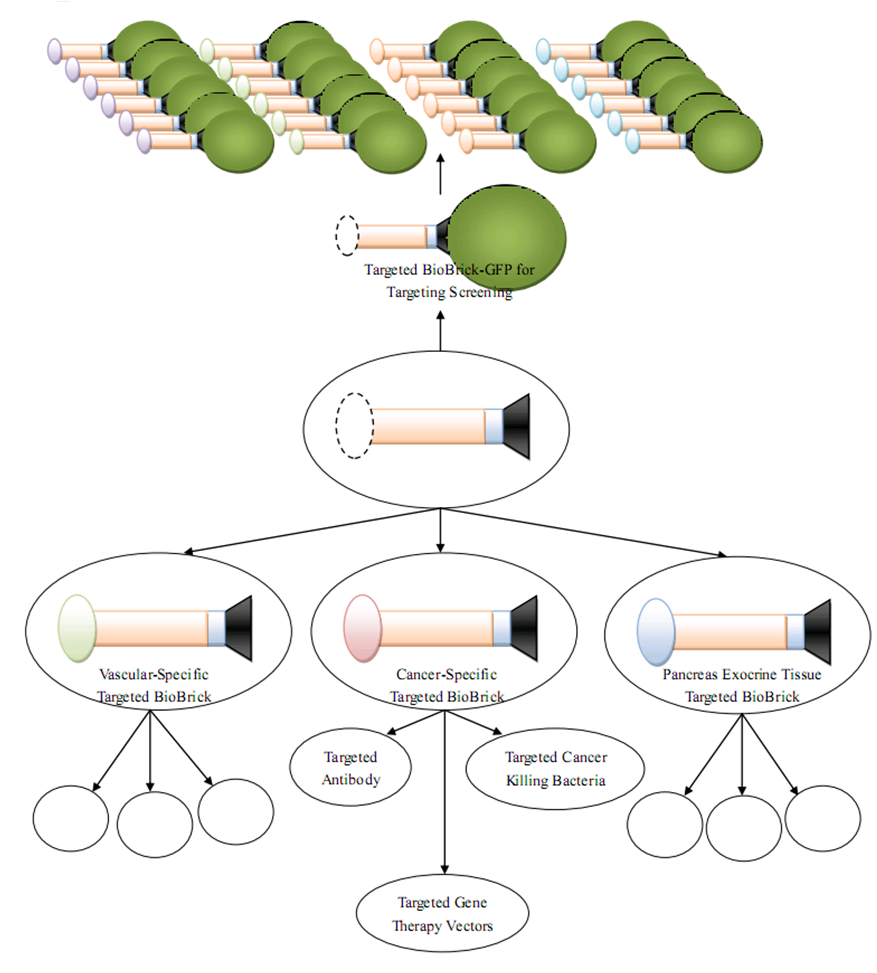
Since our Targeted Biobrick accords the synthetic biology's standardization principle at both the DNA and the protein level, we also proposed a Targeted Biobrick Platform approach for researchers to contruct targeted proteins of different functions based on standardized rules, which will significantly faciliate the application and industrialization of gene therapy and related fields. | Since our Targeted Biobrick accords the synthetic biology's standardization principle at both the DNA and the protein level, we also proposed a Targeted Biobrick Platform approach for researchers to contruct targeted proteins of different functions based on standardized rules, which will significantly faciliate the application and industrialization of gene therapy and related fields. | ||
| + | |||
| + | [[Image:TBP.png|900px|center|thumb|Targeted BioBrick Platform]] | ||
| + | |||
| + | ==Large-Scale Peptide Modification on Specificity Screening== | ||
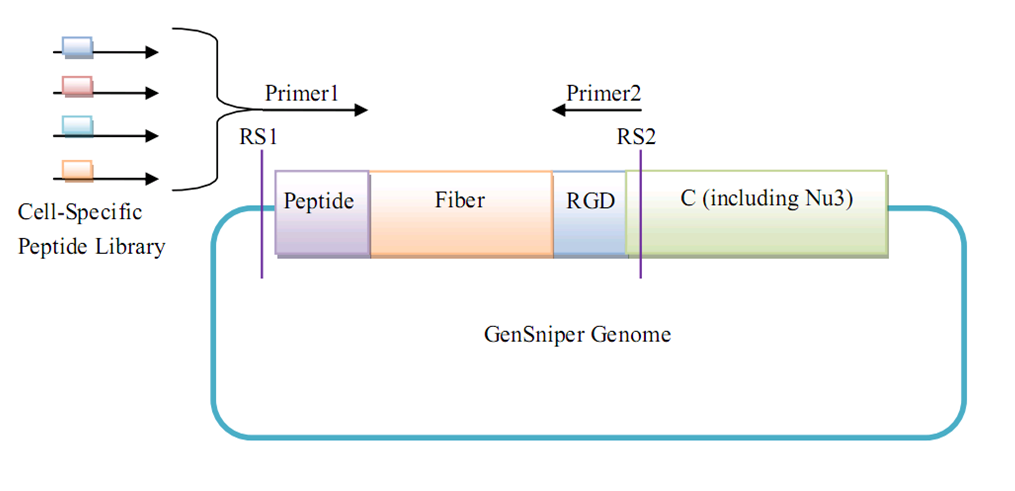
| + | Since the interchangeable module of our synthetic Targeted BioBrick is the small peptide, a large-scale modification of different peptides on the same Targeted BioBrick scanfold is recommended. As for definitive peptides of known specificity, large-scale modification can produce an industrialized library for standardized targeted gene therapy vector or other synthetic machines based on Targeted BioBrick. As for randomized peptides, large-scale modification can be used for the screening of peptides specific for a given type of cells. | ||
| + | |||
| + | Considering the importance of large-scale peptide modification, we contributed a BBF RFC (BBF RFC 46) to standardize future work. | ||
| + | |||
| + | [[Image:largescalemodi.png|700px|center|thumb|Peptide modification procedure for Targeted BioBricks on GenSniper]] | ||
=Safety= | =Safety= | ||
| Line 81: | Line 95: | ||
We have documented Targeted BioBrick in the Registry for safety issues. However, as for basic research, this BioBrick will not raise safety issues to researchers. As for clinical uses, this BioBrick should be carefully characterized as other engineered proteins for clinical use. | We have documented Targeted BioBrick in the Registry for safety issues. However, as for basic research, this BioBrick will not raise safety issues to researchers. As for clinical uses, this BioBrick should be carefully characterized as other engineered proteins for clinical use. | ||
| + | |||
| + | |||
| + | ==Safety Characterization Approaches== | ||
| + | As for the possible safety concerns, we outlined the characterization system for safety at two levels, cellular level and individual level. | ||
| + | |||
| + | ===Cellular Safety Characterization Approach=== | ||
| + | As for cellular level, we suggest researchers to use the GFP-fused Targeted BioBrick to characterize the localization of Targeted BioBrick in the target cells. For cell viability, MTT staining is required. FACS should be applied for apoptosis, autophagy and senescence identification to ensure that the Targeted BioBrick based synthetic machine will not affect the fate of the targeted cells. Gene chip and Western blotting are recommend for reprogramming approaches based on Targeted BioBrick. Western blotting can also be used to identify whether contain oncogenes (such as ras, myc) has been upregulated. | ||
| + | |||
| + | [[Image:Safety1.png|800px|center|thumb|Cellular safety characterization approach]] | ||
| + | |||
| + | ===Individual Safety Characterization Approach=== | ||
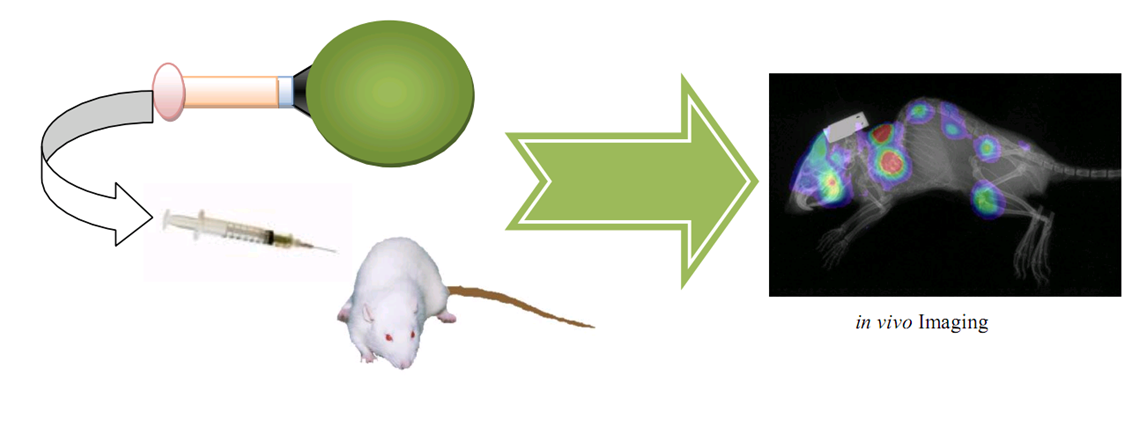
| + | As for individual level, Targeted BioBrick-GFP can be used to test whether the desired machines (no matter protein complexes, drugs or gene therapy vectors such as GenSniper) have been delivered to contain tissues. If no, they may cause potential side effects. | ||
| + | |||
| + | [[Image:Safety2.png|800px|center|thumb|Individual safety characterization approach]] | ||
=Ethics= | =Ethics= | ||
| Line 117: | Line 145: | ||
=Innovation and Human Practice= | =Innovation and Human Practice= | ||
By comparing the '''structural and functional similarity''' of bacteriophage lambda and adenovirus, this project applies the synthetic biology strategy at the '''genomic level''', transplanting the complex process of viral gene therapy vector production into prokaryotic cells for mass production. This approach makes the industrial production of gene therapy vector '''easier to standardize and manipulate''', which coordinates the standardization, abstraction and decoupling principles of synthetic biology. As for the synthetic engineering of the Targeted Biobrick, we decouple the process of viral infection into two sub-steps, attachment and internalization, which are ensured by the synthetic targeted peptides and RGD domain respectively. Viewed from the overall project, this approach is equivalent to '''synthesizing a viable form that actually contributes to clinical use'''. This reflects the synthetic biology’s extension in human practice by '''learning from nature'''. | By comparing the '''structural and functional similarity''' of bacteriophage lambda and adenovirus, this project applies the synthetic biology strategy at the '''genomic level''', transplanting the complex process of viral gene therapy vector production into prokaryotic cells for mass production. This approach makes the industrial production of gene therapy vector '''easier to standardize and manipulate''', which coordinates the standardization, abstraction and decoupling principles of synthetic biology. As for the synthetic engineering of the Targeted Biobrick, we decouple the process of viral infection into two sub-steps, attachment and internalization, which are ensured by the synthetic targeted peptides and RGD domain respectively. Viewed from the overall project, this approach is equivalent to '''synthesizing a viable form that actually contributes to clinical use'''. This reflects the synthetic biology’s extension in human practice by '''learning from nature'''. | ||
| - | |||
| - | |||
An academic summary is given in our [https://2009.igem.org/Team:Tsinghua/Design1 conclusion]. | An academic summary is given in our [https://2009.igem.org/Team:Tsinghua/Design1 conclusion]. | ||
| + | |||
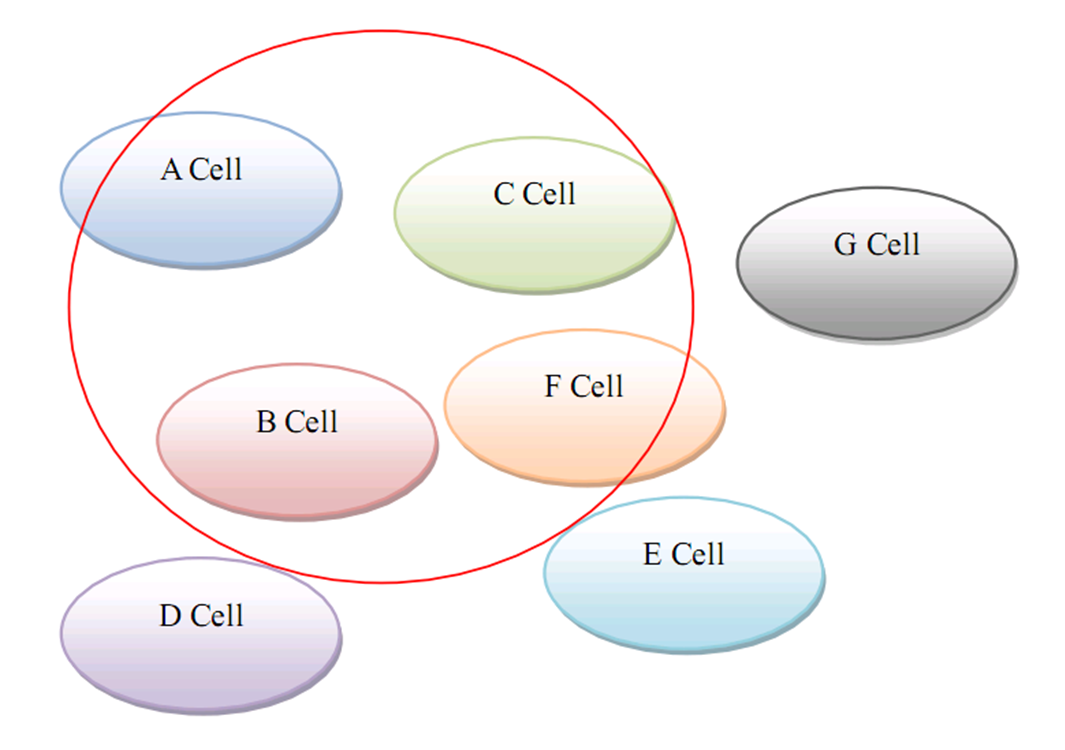
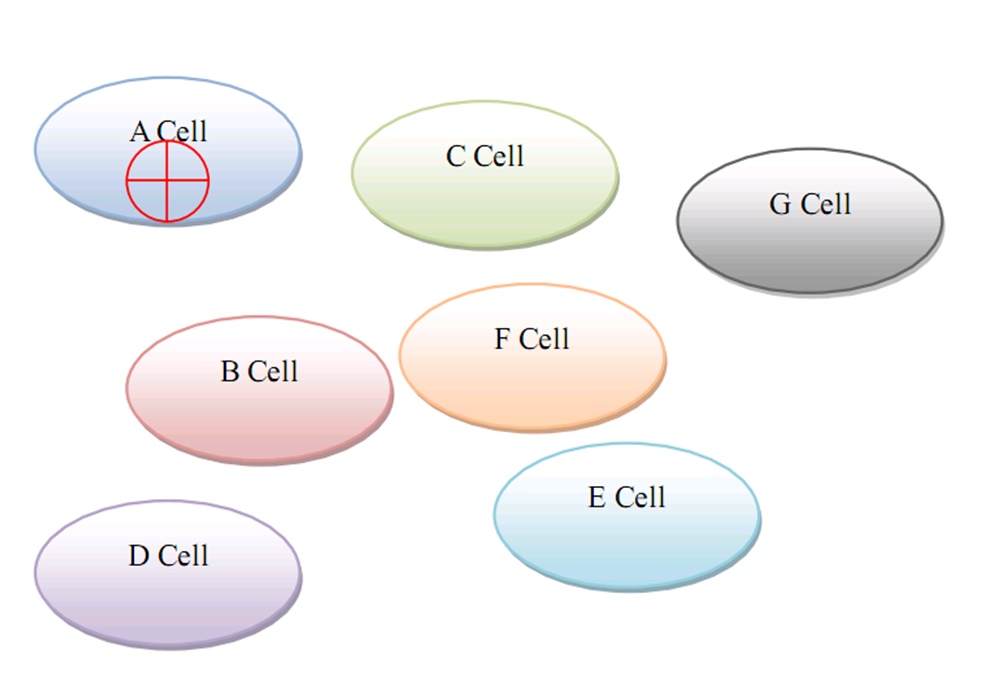
| + | ==Targeted Gene Therapy for Various Diseases== | ||
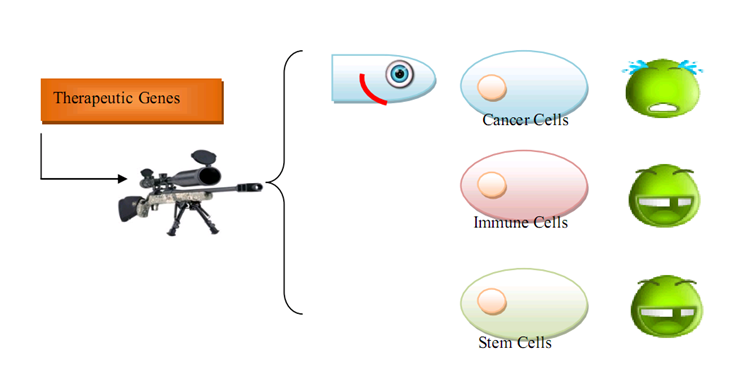
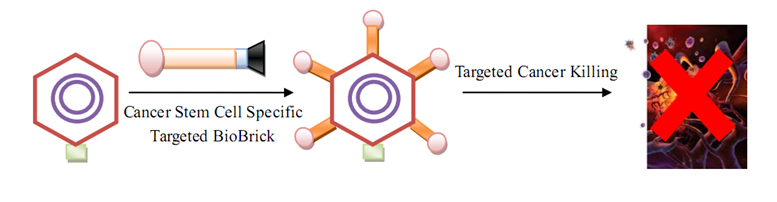
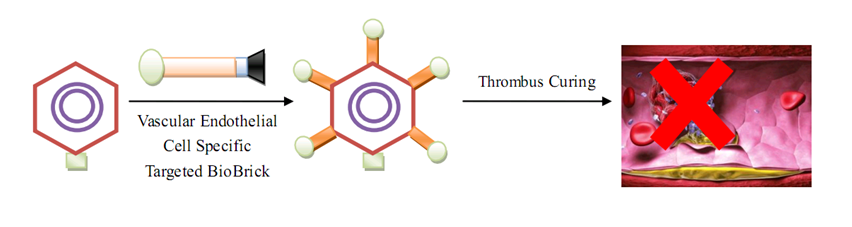
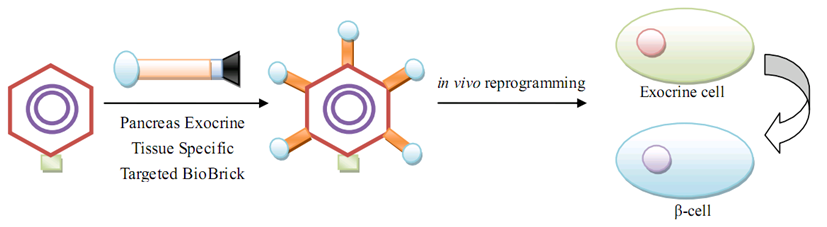
| + | As for human practice, our GenSniper with Targeted BioBirck can faciliate the development of targeted gene delivery, a bottleneck problem of in vivo human practice approaches regardless of the specific genetic network design after delivery. For example, if modified with cancer stem cell specific Targeted Biobricks, GenSniper will be able to specifically kill cancers. If modified with vascular endothelial cell specific Targeted Biobricks, GenSniper will be able to cure thrombus. Given the in vivo reprogramming results, if modified with pancreas exocrine cell specific Targeted Biobricks, GenSniper will be able to cure diabetes via a more safe and precise reprogramming from exocrine cell to beta-cells. | ||
[[Image:humanpractice1.png|800px|center|thumb|GenSniper for cancer therapy]] | [[Image:humanpractice1.png|800px|center|thumb|GenSniper for cancer therapy]] | ||
| Line 129: | Line 158: | ||
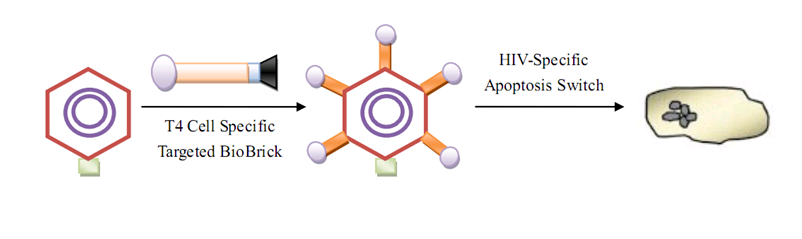
[[Image:humanpractice4.png|800px|center|thumb|GenSniper for AIDS therapy]] | [[Image:humanpractice4.png|800px|center|thumb|GenSniper for AIDS therapy]] | ||
| + | |||
| + | ==Industrialized Production and Purification of GenSniper== | ||
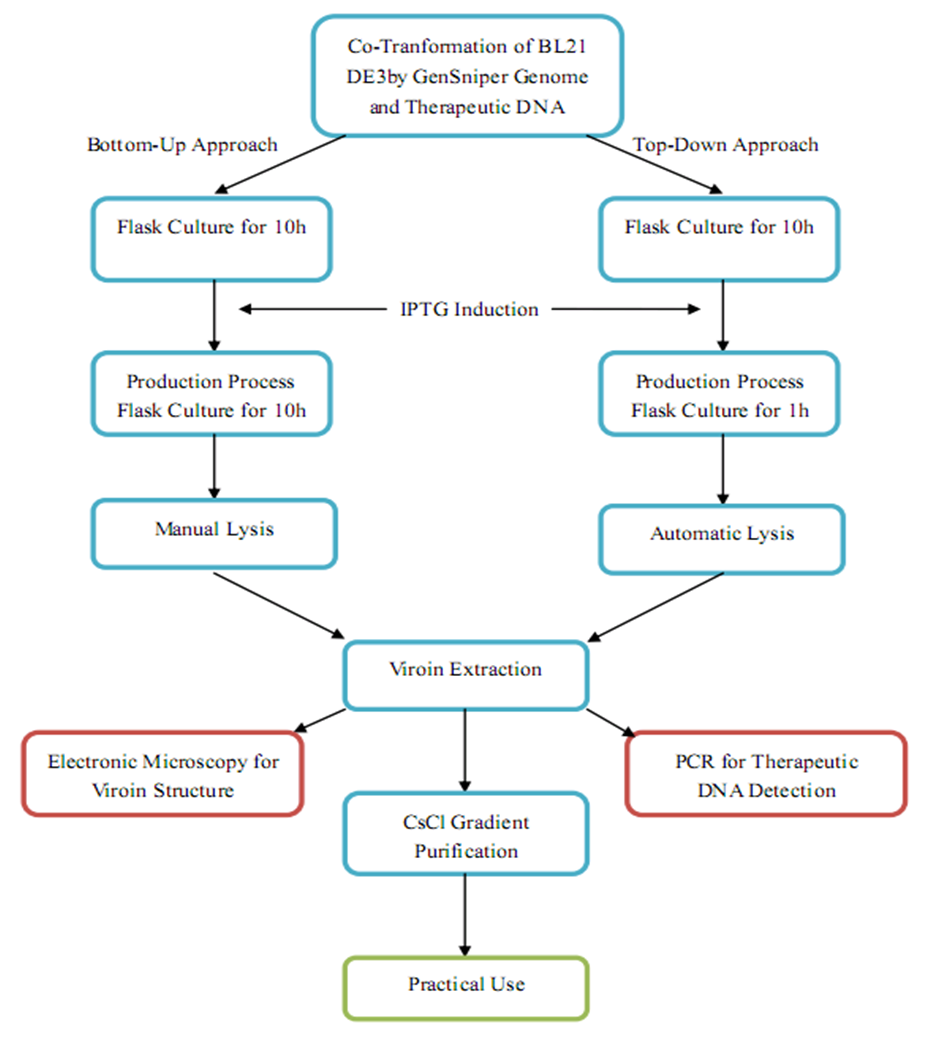
| + | Another main innovation of our project is the transplantation of eukaryotic production system to prokaryotic production system, which, if fully achieved in the future, may be revolutionary for mass application of gene therapy approaches as well as gene delivery systems for lab work. According to our wetlab and drylab results, we outline and detail a procedure of GenSniper production. | ||
| + | |||
| + | [[Image:productionap.jpg|center|thumb|800px|Production Scheme of GenSniper]] | ||
| + | |||
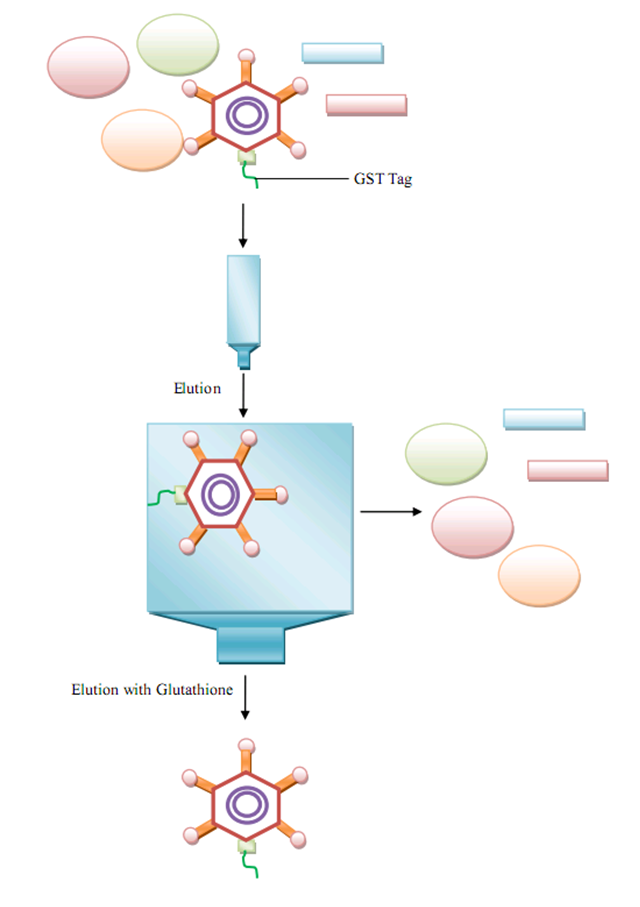
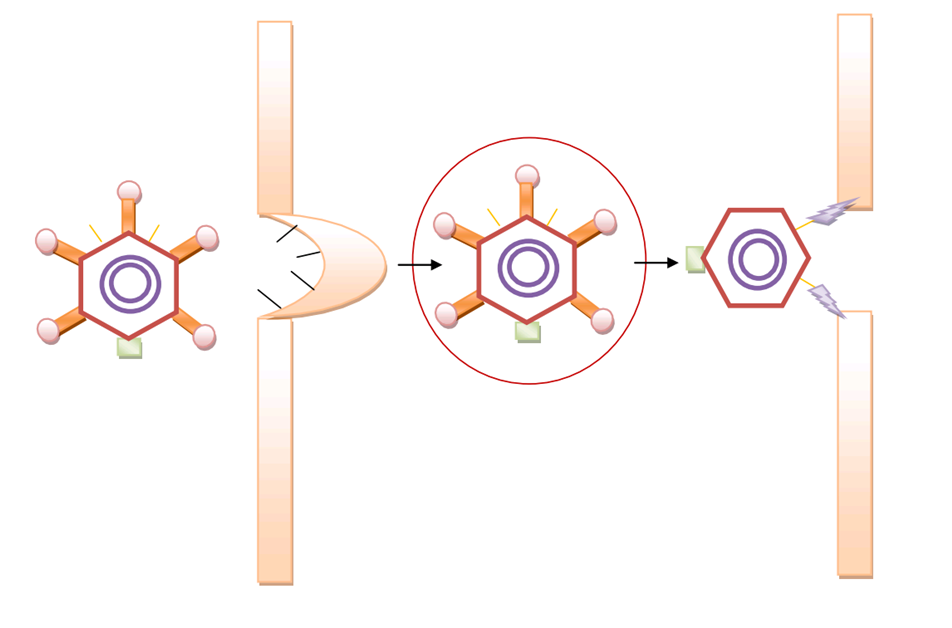
| + | As for further purification, which is not mentioned in our wetlab design, we also propose an easy approach based on another protein modification. This modification is for the residue of the tail structure of bacteriophage lambda, namely FII protein. With a GST tag or other tags available, GenSniper viroins can be easily purified on a large scale, which extend our synthetic biology project to industrialization. | ||
| + | |||
| + | [[Image:productionap2.jpg|center|thumb|600px|Purification Scheme of GenSniper by GST-tag modified FII]] | ||
| + | |||
| + | ==Extension of GenSniper Genome to Faciliate Theraputic Genes Expression== | ||
| + | As suggested in modeling, E from the GenSniper genome is highly recommended to be modified by NLS, which in turn faciliate the recruitment of Theraputic DNA into the nucleus of the target cells. | ||
| + | |||
| + | [[Image:interce4.jpg|700px|center|thumb|GenSniper Intracellular trafficking model can be ensured by display NLS sequence on E]] | ||
| + | |||
| + | In addition, researchers can also incorporate Mu gene from adenovirus genome into the synthetic GenSniper genome. Since, Mu is responsible for DNA binding and targeting in wildtype adenovirus, this extension will further faciliate the localization of transcription active sites in the nucleus. However, since GenSniper must be produced in prokaryotic system, whether or not Mu is functional needs further investigation. | ||
| + | |||
| + | [[Image:Mu.jpg|700px|center|thumb|GenSniper Nuclear Localization by Mu protein from adenovirus]] | ||
= References= | = References= | ||
| Line 163: | Line 210: | ||
[16]http://www.ndsu.nodak.edu/instruct/mcclean/plsc431/students/amanda.htm | [16]http://www.ndsu.nodak.edu/instruct/mcclean/plsc431/students/amanda.htm | ||
| + | [17]http://www.molbio.princeton.edu/index.php?option=com_content&task=view&id=253&Itemid=249 | ||
{{Tsinghua/Pic}} | {{Tsinghua/Pic}} | ||
Latest revision as of 23:37, 21 October 2009
| |||||||||||
 "
"