Team:Cambridge/Notebook/Week6
From 2009.igem.org
Categories :
Project :
-
Overview
Sensitivity Tuner
--- Characterisation
--- Modelling
Colour Generators
--- Carotenoids (Orange/Red)
--- Melanin (Brown)
--- Violacein (Purple/Green)
The Future
Safety
Notebook :
Team Logistics :
Week 6
Models made for switch, latch, and differing experimental conditions - issue of melanin retention explored - one PstI site removed from MelA - pink carotenoid pigment produced - pseudomonas strain recieved - started planning the powerpoint presentation
Monday
Dry Work
Modelling
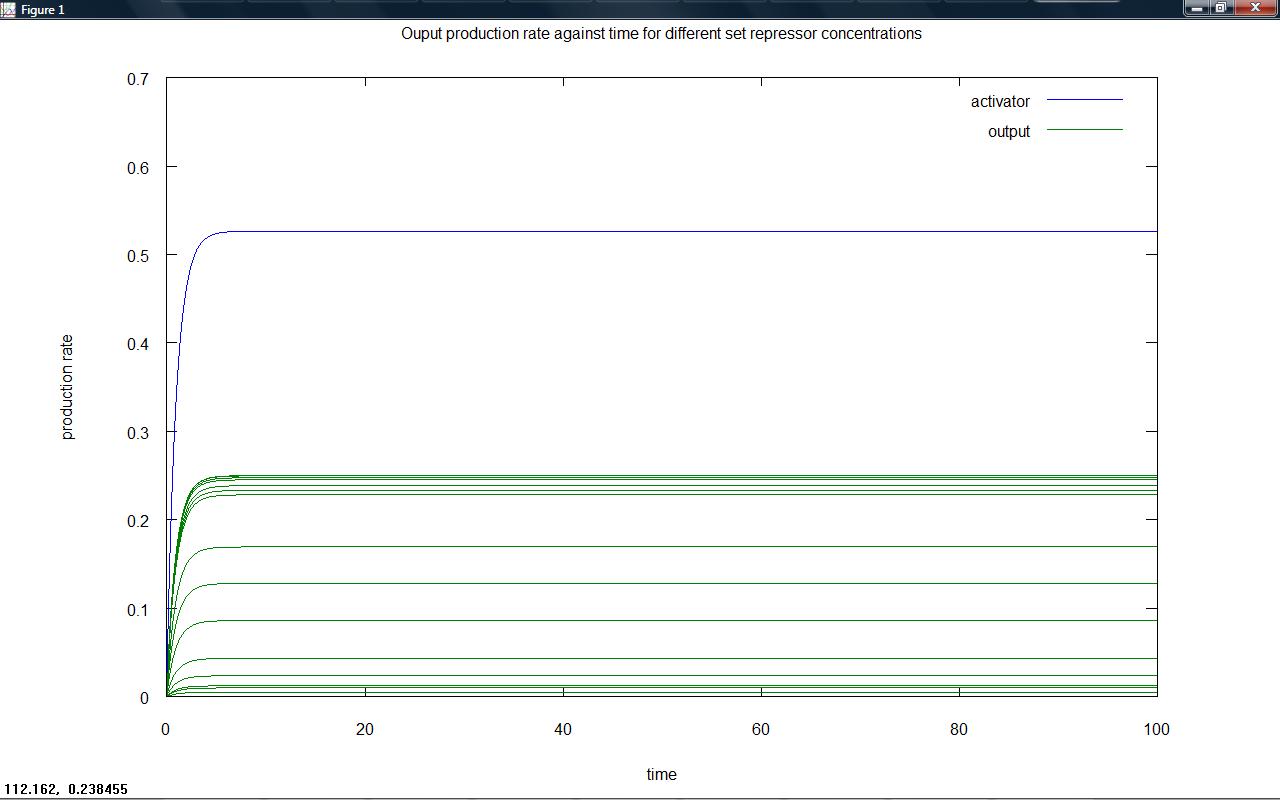
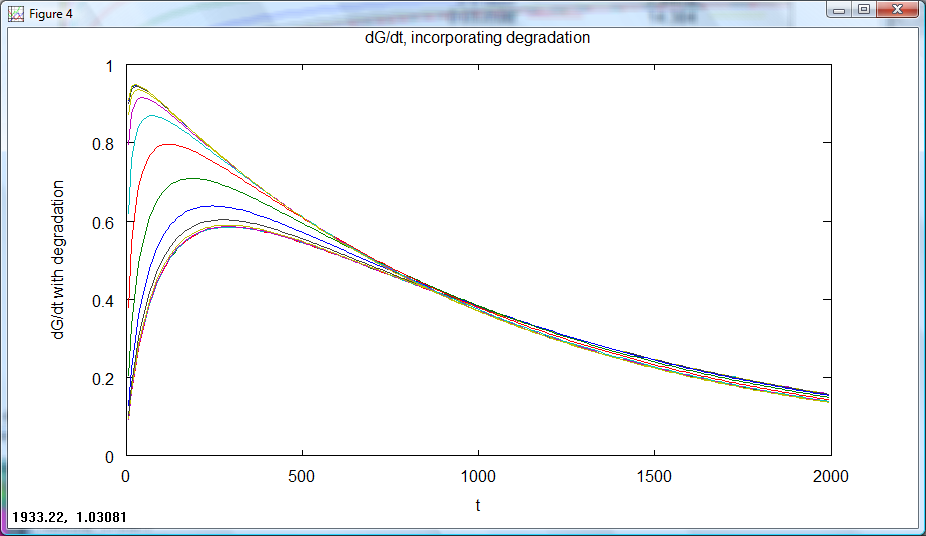
On friday, the two models for a proposed threshold switching system were created. Model 6.2 (competition between activator and repressor for a site on the DNA) was investigated; For different 'arabinose' (or other proposed input) and repressor concentrations the final level of output from the system was plotted. It was hoped that for different input levels (shown in the below graphs by the different green lines of output) the position of switching to low output would take place at different repressor levels due to the competitive nature of the system. Investigation into refining the model will take place. It is important to remember that whilst total output is what is seen, plotting rates of output production is necessary; the 'switching level' could be considered to be the point at which the rate of output production becomes zero. A standard way of designing this switching level is required.
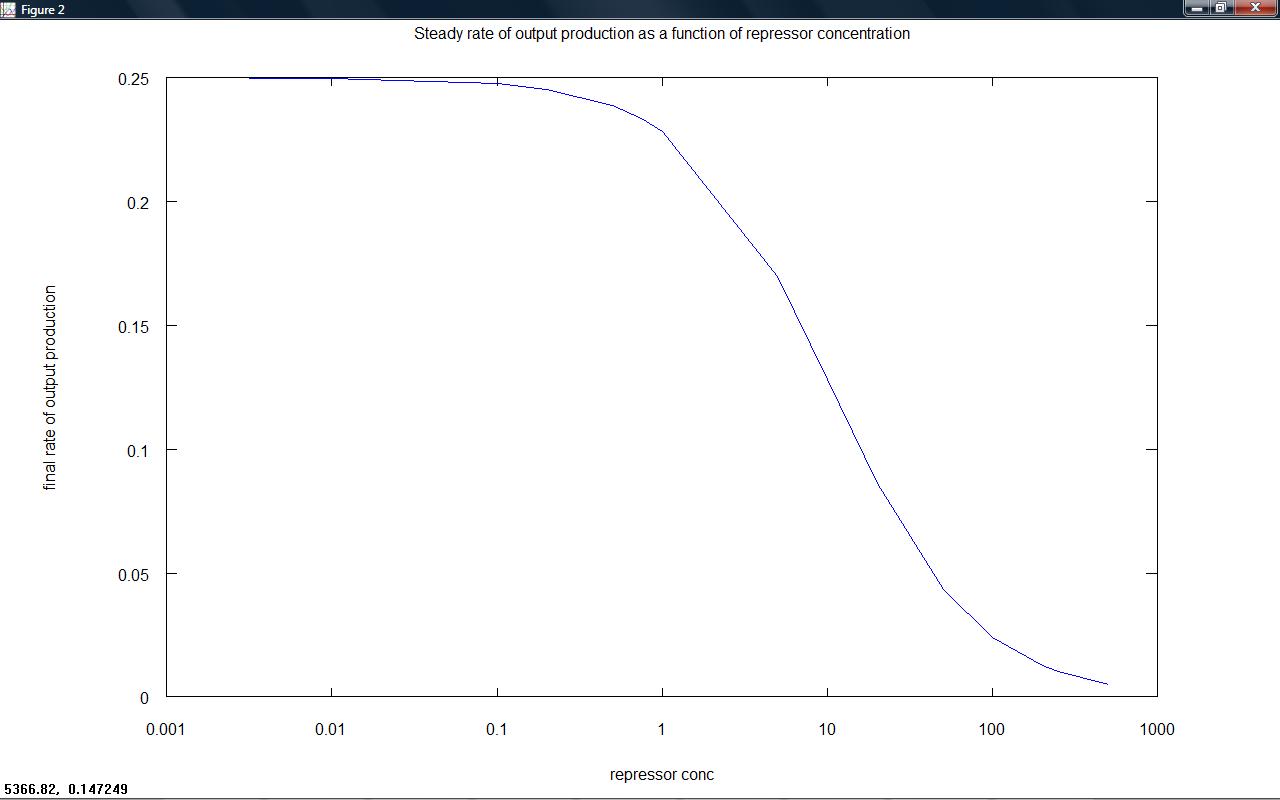
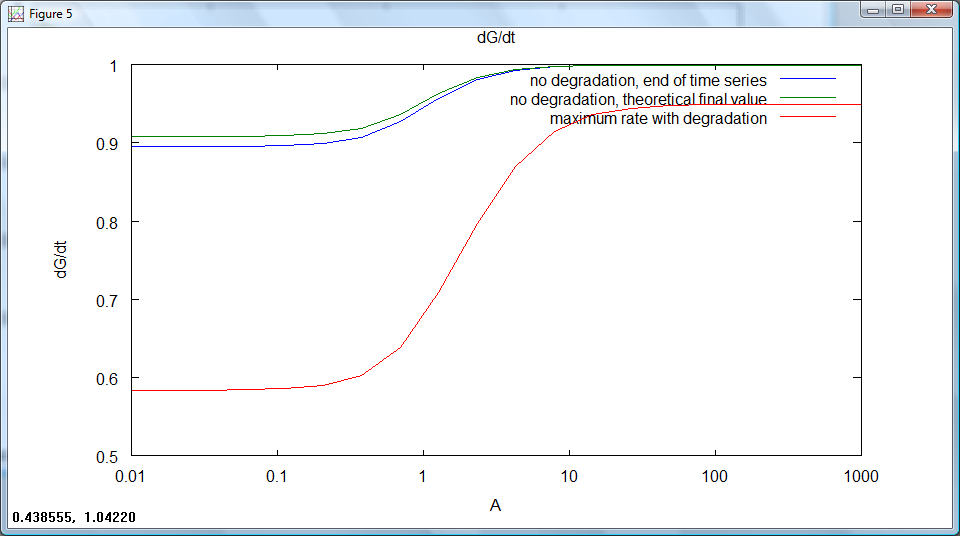
Rate of output production against time for a series of set repressor levels showed that different steady rates of production were rapidly obtained (figure 1 below). The final rate of production (representative of the steady state) was plotted against repressor level, showing that the 'switching level' would occur in the region where the rate falls to zero (figure 2). Ideally, our system would have a well defined switching point, with a much sharper rise to high rates of output. This would likely be achieved by increasing hill coefficients, requiring a system with a greater degree of cooperativity.
Figure 1
figure 2
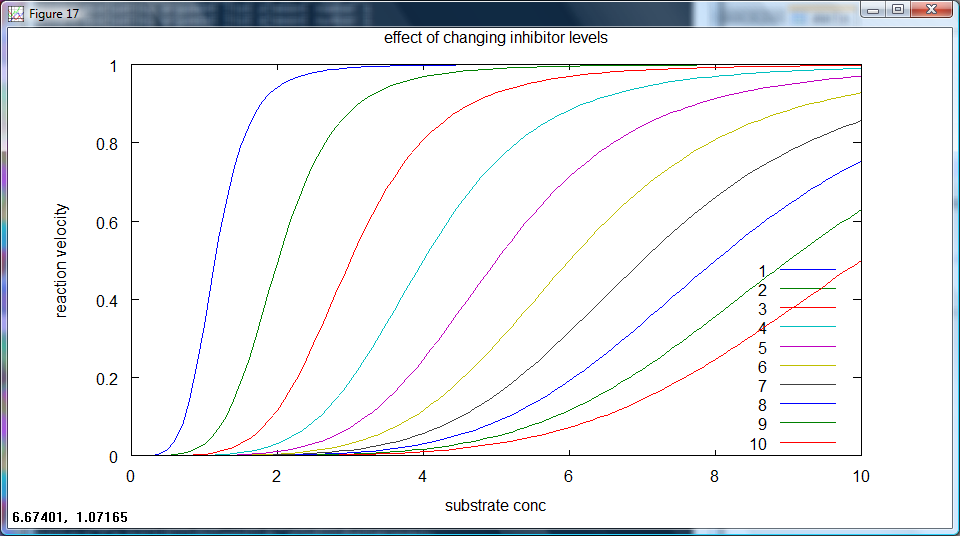
Ideally, for each set repressor level, the output will have a non-zero rate of production at different levels of arabinose input. The most useful plot here is rate of output against input concentration, with different set repressor levels as different lines on the same graph. This is equivalent to the competitive inhibtion of enzyme action, a sample plot of such a case is given below (figure 3).
figure 3
The feasibility of such a system investigated, the question is how to implement such a system?
To continue the modelling work; models need to be refined according to data that begins to come from the plate reader. A return to looking at the latch in terms of rates of output production will possibly be useful, although no such clean switching level is expected to be possible. Also, the preliminary plate reader data shows a decrease in rate of GFP production having reached a peak value. A possible explanation is the large requirements the high rates place on transcription and translational machinery of the cell. The overall effect could be considered to be one of negative autoregulation with potential for oscillations about a steady level, damping as the system progresses through time. Finally, this week we also aim to look at stochastic simulation possibilities using StochSim.
Wet Work
Threshold Device
After succesful PCR of pSB3K3, did a double digest (PstI and XbaI) of pSB3K3 (vector) and the following inserts: I746390, I746391, I746392, I746394, I746395. Ligated vector and each insert, transformed into Top10.
Melanin Retention
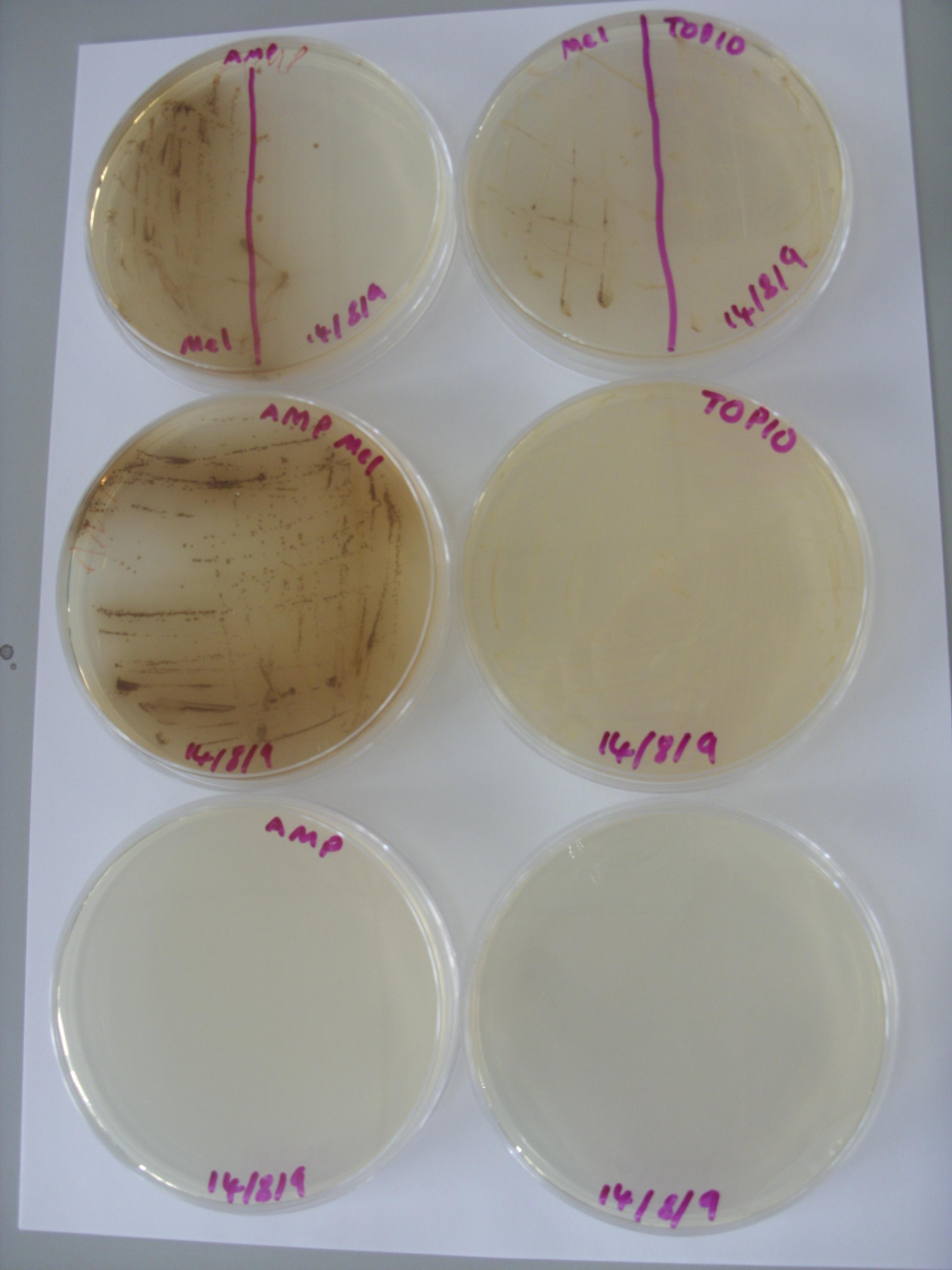
Inspection of the plates from Friday demonstrated no clear reduction of melanin efflux from the strains containing the multi-drug transporter knockouts (see below), thus it appears that the hypothesis that melanin efflux might be controlled by one of these two transporters doesn't stand up.
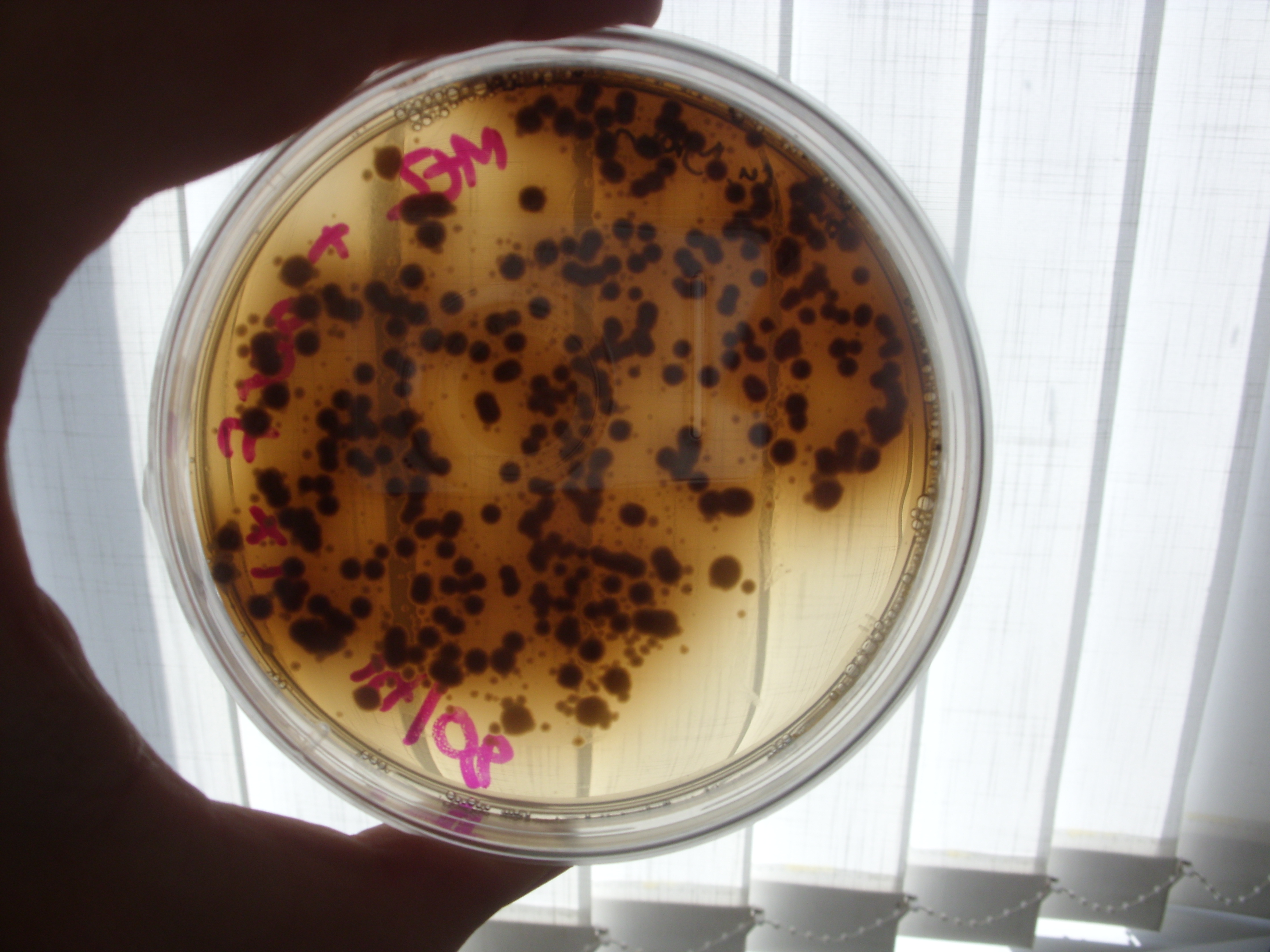
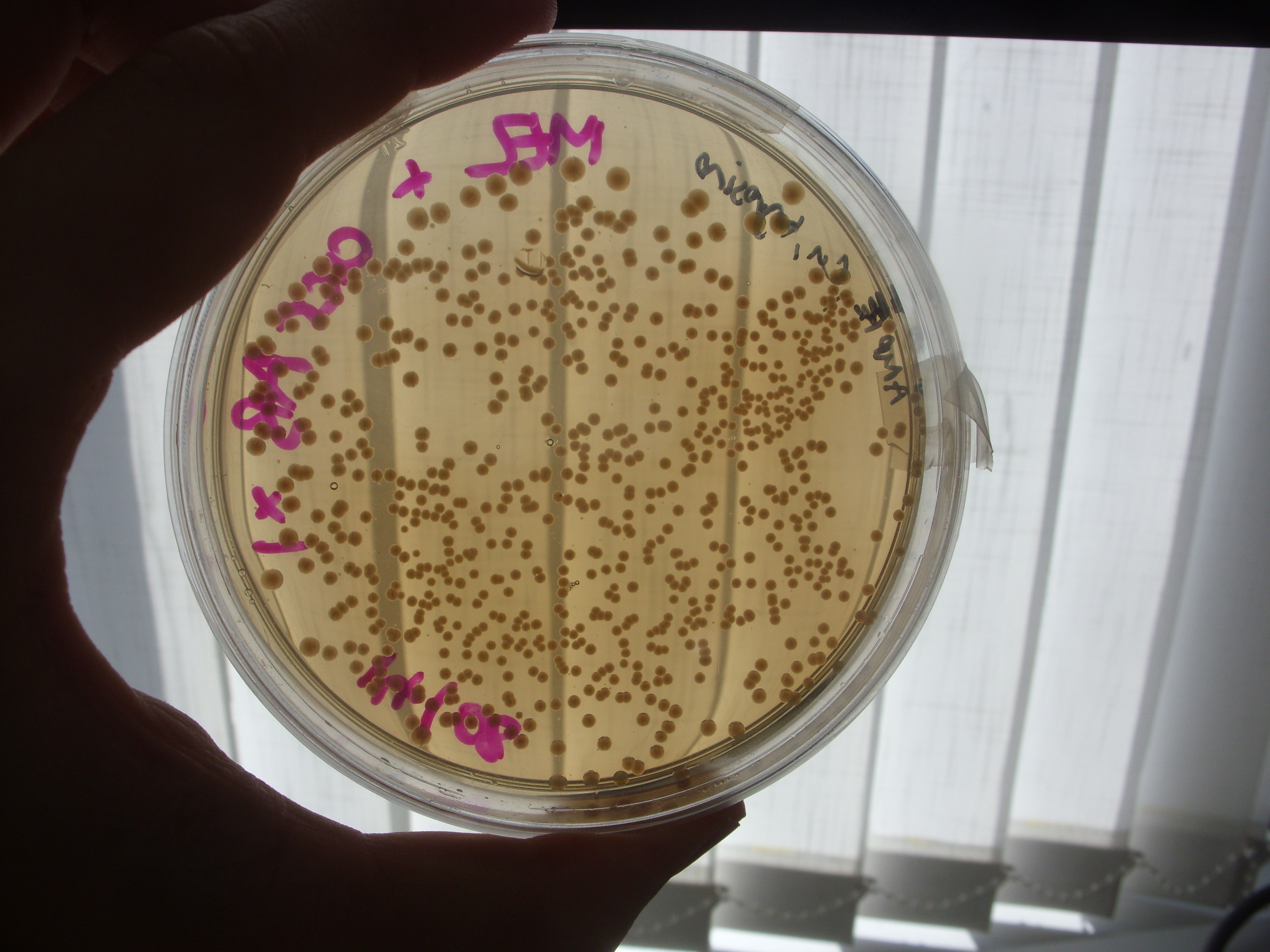
Tuesday
Dry Work
Modelling
Thus far modelling has been rather preliminary. A systematic investigation of varying the parameters in a basic activator model is underway. There are nine parameters that can be changed relating to binding constants, hill coefficients and the rate of degradation of the activator protein.
A set of 'experiments' were performed in silico where these parameters were changed to study the effect on such a system. The results can be summarised as follows:
- Increasing the total repressing 'araC' (or equivalent repressor usually present in the cell) ten-fold reduced the rate of output at a given time and the lowest input concentration from five-fold.
- Reducing K_x one-hundred fold shifted the concentration at which rate of output shifted from low to high, and a reduction to nearly zero of the lowest concentration rate of output was seen. This is possibly because at the time taken the rate of output was not in the vicinity of the steady state maximum. However, reducing K_x implies increased affinity of repressor for its sequence upstream of the promoter, so it would be expected to shift the input concentrations required to get an 'on' response.
- Changing K_a also shifted the 'switching level' concentration, but towards lower concentrations. Expected this as decrease in K_a increases arabinose-repressor affinity. Also the 'basal' level of output was returned to 0.1 because this system will build to the maximum rate of output for a given concentration faster.
- Decreasing K_p did not shift the switching position (disappointing as it is hoped that different activator-promoter combinations will switch at different input levels), but did raise the rate of output at the lowest concentrations significantly. Decreasing K_p increases activator-promoter affinity in effect, so it would be expected as the system will respond to small levels of activator produced strongly.
- Increase in d_p, the rate of activator degradation, meant that the time taken to reach a steady rate of output production was longer.
The rate of output production chosen was that at the greatest time. Whilst this was not necessarily steady, we are looking at the potential visible outputs and what is important is the visible output after some length of time, not when we can be sure that the system has reached steady state.
Wet Work
Melanin pigment production
Examined the control plates made on Friday:
The picture clearly shows that the pigment production is due to the transformed cells, rather than conditions in the plate, or interactions between cell and the plate.
Making MelA BioBrick
The first PCR was carried out to removve one of the PstI restriction sites. (primers A/B and primers C/F) The first attempt only A/B worked, so this was repeated using Touchdown PCR, with two samples for each primer pair (one with DMSO and one without). The PCR was left running overnight.
Threshold Device
Transformations of ligations were unsuccessful :( Positive ligation control was plated on the wrong antibiotic plate, so will repeat that tonight. Today de-bugged restriction digests. The gels showed about 50% ligation after 3 hours, so have repeated the double digest with another pSB3K3 sample and 380, 381, 382, 384, and 385, this time separating the DNA into 2 separate digests. Gel extraction was successful for 380 and 384; PCR-ed more pSB3K3, as this was lost in the gel extraction (actually doesn't need to be gel extraction, can just be column purified).
Another batch of overnight cultures of the 390, 391, 392, 394 and 395 constructs on the pSB1A2 plasmid were made up in preparation for another set of minipreps on the morrow, following exhaustion of the original miniprep stocks.
Wednesday
Dry Work
Modelling
From preliminary data we saw the rate of output drop off at higher input concentrations after an initial rise to the highest rate of production. One possible explanation was that the transcriptional machinery becomes tied up in processing the huge input and results in a slowing of output rate. This could be seen as a form of negative autoregulation that frequently results in oscillation about a stable value. The key factors responsible here were the direct negative autoregulation of protein production and time lags in the transfer of mRNA out of the nucleus, translation and the possibility of mRNA degradation.
It was realised that the simple amplifier system can be solved analytically. A new Matlab file was made to show the results of this, also implementing vectorisation techniques. For degradation of the reporter, numerical integration was necessary.
Wet Work
Threshold Device
Positive ligation control grew successfuly overnight, confirm that the ligation was successful. Possibly lack of vector DNA is why the other ligations didn't work. Did a double digest of pSB3K3, ligated it to pre-cut 381 and 384. Transformed pSB3K3+381 and pSB3K3+384 into Top10 and the arabinose strain, and incubated overnight on kanamycin plates.
Crisipian AND MEGAN (and Alan!) very kindly miniprepped all the activator constructs again so we can continue to consume the DNA at a shocking pace.
MelA BioBrick
Started the protocol to remove restriction sites. PCR with primers A/B worked, however, no results were present for PCR with primers C/F. As both A, F and the enzyme have been shown to work, it was decided that this was probably an issue with Primer C. It was decided to move on and attempt to remove the second resitriction site, using primers A/D and E/F.
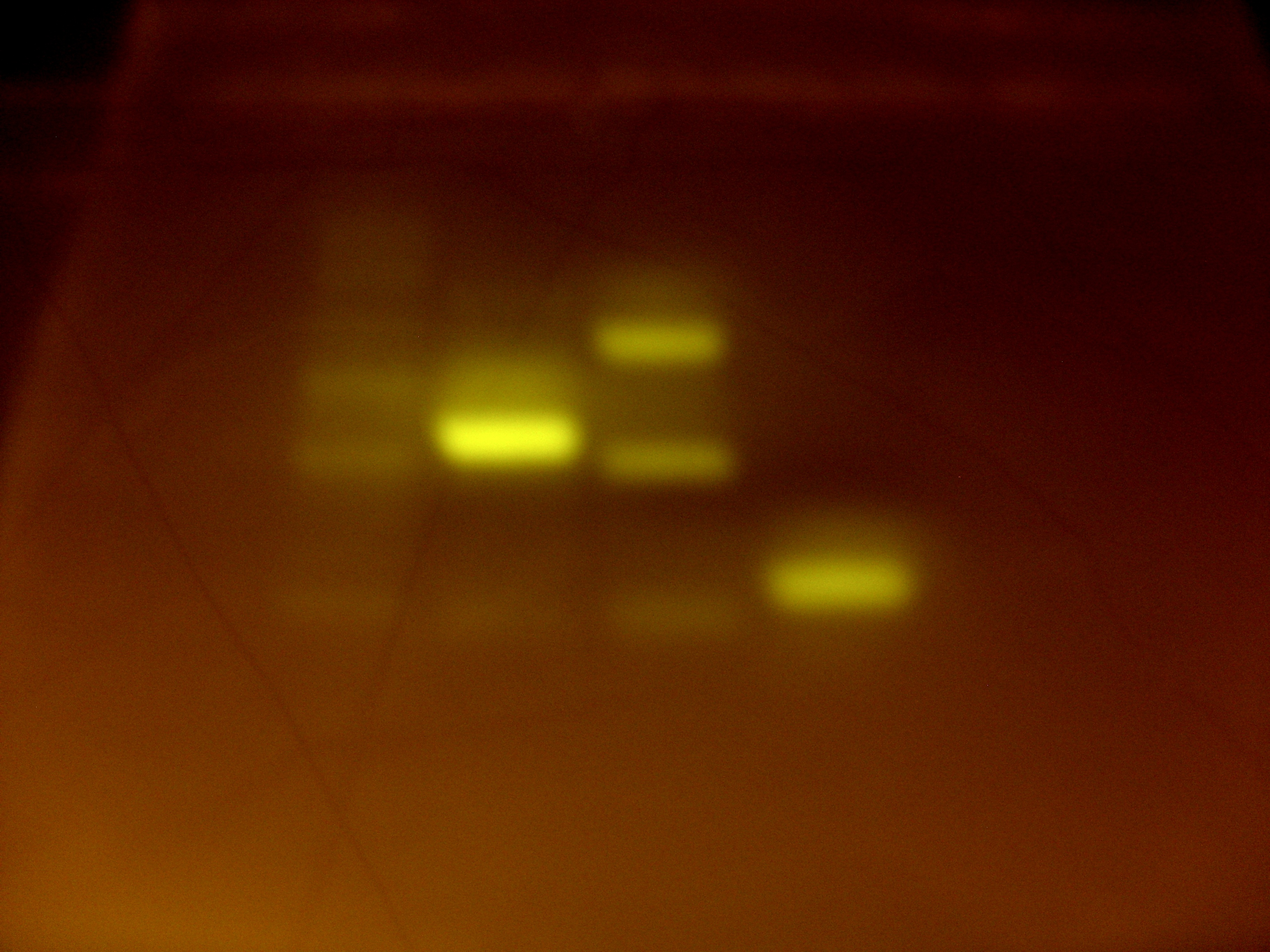
The same Finnzymes reaction mixture was used, along with the Touchdown PCR procedure. Both sets of primers worked! The gel was put in the fridge for DNA extraction and the continuation of the MelA BioBrick formation tomorrow.
From left to right the bands show: 2-log ladder, reaction mix with primers A and D and DMSO added, reaction mix with primers A and D and no DMSO, resction mix with primers E and F with no DMSO. The second and fourth lanes had the DNA at the correct length for the fragments we were expecting, and so these were used for DNA extraction.
Thursday
Wet Work
Threshold Device
Transformation of Top10 with I746381 in pSB3K3 and I746384 in pSB3K3 was successful. Overnight cultures were made of these in preparation for miniprep, glycerol stocks, and streaking out single colonies tomorrow. Overnight culture of as well as I746340 in order to miniprep. We want to move I746340 into pSB3K3 as well in order to characterize it against 1RPU. Overnight colony PCR of single colonies of I746381 in pSB3K3 and I746384 in pSB3K3
Restriction digest of more pSB3K3 and I746391 and I746374, ligated each activator construct to pSB3K3 vector and transformed into Top10, incubated overnight, fingers crossed again.
PCR-ed more pSB3K3.
Carotenoids
PINK PIGMENT IS HERE!:
On the left: Amp EB1 Roche Strin, 1/10th 18/8
On the right: Amp EB1Y ligation control
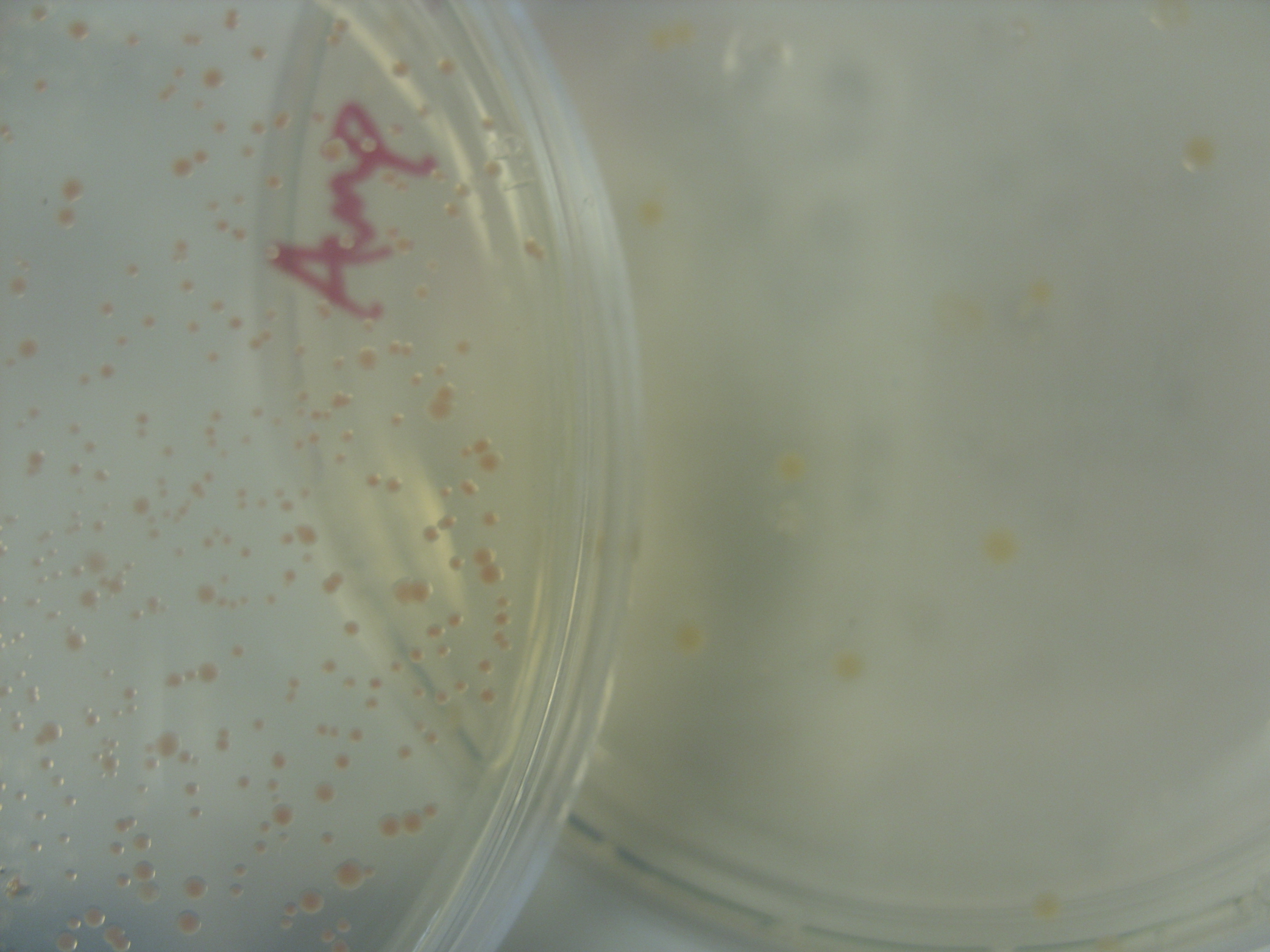
MelA BioBrick
Did DNA gel purification of the AD and the FE segments then ran a quick E-gel to confirm both segments were present. While the F/E fragment is the correct size, the A/D fragment is too small. A closer look was taken at the gel run the other day, and the correct sized fragment was identified for gel extraction.
Pseudomonas
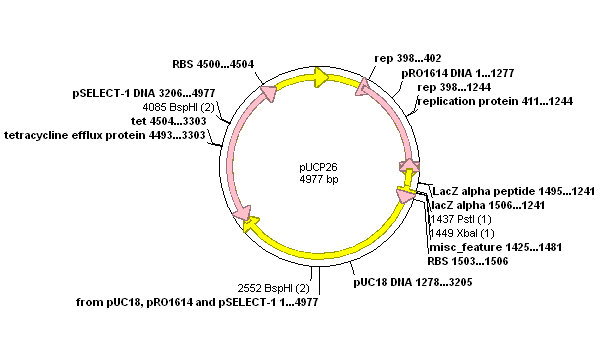
Received E.coli clones containing pUCP-M (containing PhzM) and pUCP-MS (containitn PhzM and PhzS). Both genes are held on the pUCP26 vector (shown below):
Ideally we would convert both the PhzM gene (from pUCP-M) and the PhzM/PhzS construct (from pUCP-MS) into two different biobricks, giving the colours red and green respectivly.
Looked into purchasing phenazine-1-carboxylic acid (PCA), CAS #2538683, as it is the precursor for pyocyanin production that can be exogenously added to" a final concentration of 0.3 mg/ml from a 25 mM stock solution in 5% (wt/vol) NaHCO3" for liquid culture.
Dry Work
Having looked at the sample data sheet for part F2620 in the registry, a re-think of our data gathering and analysis method for the amplifiers is suggested. As a guide to what needs to be done, suggested that the information with F2620 is read on the parts registry.
Modelling
The next step with the basic phage activator system is to fit a Hill function to the data once it has been gathered so that effective constants can be found. We aim to characterise the activator systems with the three key parameters; K, Vmax and n. K gives the 'Switch Point' (point at which 50% of maximum output rate is reached), n is the hill coefficient and Vmax is the maximum rate of output production.
Friday
Wet Work
MelA BioBrick
Ran a gel of the PCR done overnight on Thursday (the two fragments of MelA with one restriction site removed with primers A and B). Achieved a correct size band! The band was cut out and the DNA extracted using the QuiGen gel extraction kit, andstored at -20 degrees. Next week a sample will be digested with PstI to test whether the site has been removed or not.
Threshold Device
Overnight transformations were successful, did a colony PCR of the transformants of 381 in pSB3K3 and 384 in pSB3K3. No 384 constructs showed an insert of the correct length, but one 381 construct did!
Dry Work
Advice on powerpoint presentations
- Present problems and solutions
- problems to solve - need pigments, amplifiers, etc.
- show biosensors - ex. arsenic - then highlight poor output
- show our solution - in effect, start with the end product!
- then delve into what we did
- Should present MAINLY the concept, then what we did and what was achieved.
- first five minutes should be a straight pitch, without any technical information
- with pigments, highlight the research behind it - where the genes are from, etc.
- Need a name and a graphic
Colours and Futures Workshop
In the afternoon we attended a workshop organised by Daisy Ginsberg and James King, which explored the potential future applications and issues arising from the ability to easily produce colour from bacteria. Thia was a good introduction to thinking about issues of Human Practice in synthetic biology as they related to our project. Specifically we explored four main areas which synthetic biology and bacterial pigment production could significantly affect in the future:
- Products
- Services
- Groups
- Law
 "
"