Team:Heidelberg/Parts Characterization
From 2009.igem.org
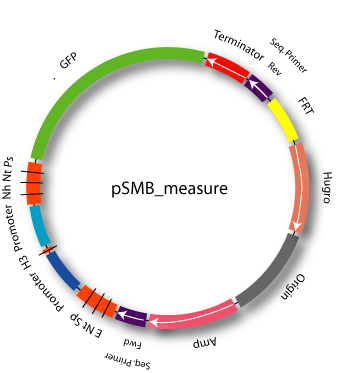
Characterized partsFind below some of our most important and best-characterized parts. Measurement KitPart:BBa_K203100We created a plasmid backbone, pSMB_MEASURE (SMB is for Synthetic Mammalian Biology; Part:BBa_K203100), which should be used for promoter characterization in mammalian cells. pSMB_MEASURE (see Fig. 1) contains a reference promoter, JeT[1] (Part:BBa_K203112), which is flanked by BBb_2 (Tom Knight) sites and can therefore be replaced by the promoter to be measured. JeT is ideal as a reference promoter for a variety of reasons. First, it has an intermediate expression strength; second, it is regulated by a wide variety of transcription factors and relatively low levels of change in fluorescence among different conditions (Fig. 2).
We separated JeT's core promoter from its proximal promoter by a HindIII site; it can therefore be used for for the de novo creation of promoters containing the JeT core promoter or for modifying the strength of a promoter by core promoter swapping. In addition, it contains a FRT site which will allow for stable integration into mammalian cells also containing a FRT site. Thus, it provides the possibility to characterize the promoter in a defined genome and in this way helps to avoid some of the challenges we identified for promoter characterization in mammalian systems. For the same reason, it also contains a mammalian selection marker (hygromycin). For the generation of the plasmid, please see part design. As a reporter gene, it contains GFP, which is followed by a SV40 mammalian terminator. We generated another plasmid pSMB_REFERENCE, which contains mCherry instead of GFP. It can be used for normalizations to transfection efficiency in Flow Cytometry and image analysis. Core promotersPart:BBa_K203113We cloned the CMV core promoter from Part:BBa_I712004 in front of the JeT proximal promoter from part Part:BBa_K203112 to obtain JeT_CMV. This construct can be used to create novel promoters containing the CMV core promoter. We characterized this construct in three different cell lines, HeLa, MCF-7 and U2-OS and found it to have 50%-60% percent of JeT's activtiy depending on the cell line (Fig. 3). Thereby, the HeLa cell line was measured five times and the MCF-7 and U2-OS cell line measurements were performed four times. The fact that variations in the core promoter can be used to vary expression strength of a certain promoter of interest comes in useful if the transfer function of an existing promoter is to be altered. It can also be used to further diversify the synthetic promoters we created.
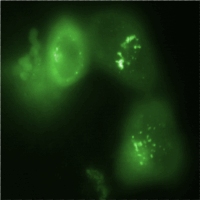
Promoters created by RA-PCRMain article: Synthetic promoters All synthetic promoters contain the core promoter of Part:BBa_K203112 and were screened / measured in Part:BBa_K203100. BBa_K203110, BBa_K203118, BBa_K203111, BBa_K203109 and others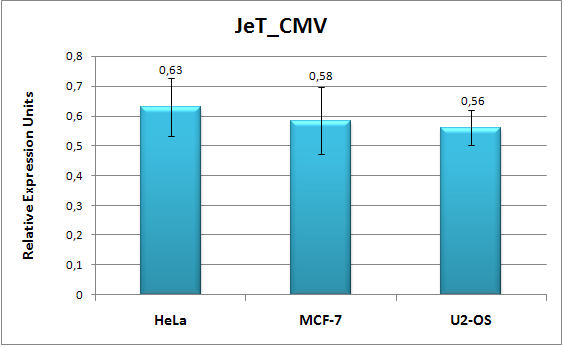 Fig. 3: Flow cytometry measurement data of JeT_CMV (REU) in different cell lines. The three cell lines MCF-7, U2-OS and HeLa were cotransfected with the JeT_CMV promoter coupled to GFP and a reference plasmid including the promoter JeT coupled to mCherry. The relative fluorescence (REU) of GFP was measured 20 hours after transfection. The MCF-7 and the U2-OS cell line were measured four times and the HeLa cell line five times. The standard error of the mean (SEM) is indicated by the error bars. As a first application of RA-PCR, we have created a library of constitutive promoters. We performed RA-PCR on oligos containing binding sites for some well known generally activating transcription factors (Sp1, Ap1,CREB, NF-Y) which we identified from literature search [2],[3]]. We also added NF-κB responsive oligos as NF-κB has non-specific activity and is therefore used by a variety of viral constitutive promoters, e.g. the HIV promoter [4]. We picked 24 colonies, two of which we dismissed after a test digest (not shown). Fig. 5 shows the sequence analysis of some randomly selected clones and demonstrates that RA-PCR is able to generate randomized repeats of oligos. We then measured the activity of the clones we picked by applying the Concept of Relative Expression Units (REU) we developed. Fig. 4 shows that we have been able to create a library of promoters of varying strength, some of which have an expression strength higher than JeT (which was not accomplished by JeT's developers, although attempted [1]). Such a library is of great value for fine-tuning gene expression levels. Selected clones were characterized by an independent method (image analysis) and submitted to the registry.
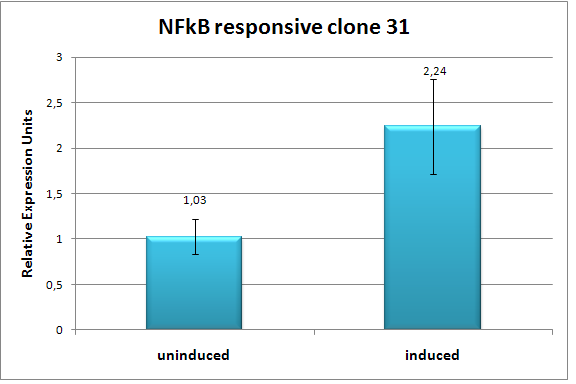
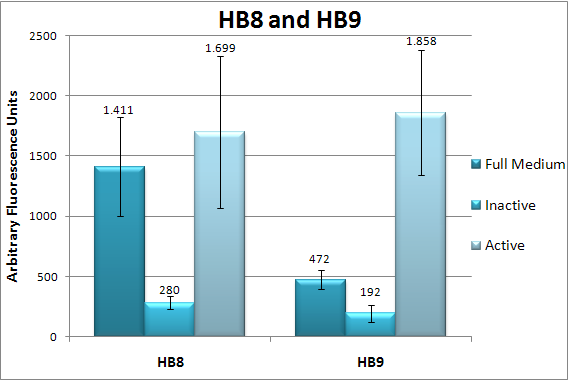
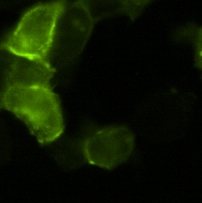
BBa_K203119We characterized clone 31 of our library of NF-κB responsive promoters by flow cytometry (3 independent experiments, each experiment having 3 replicates) and found it to be upregulated by approx. 100% (Fig. 6) in response to TNF-α. We also did a timecourse measurement of induction (Fig. 7). To verify our assumption that promoters created by RA-PCR are specific to the pathway of interest, we tried to induce clone 31 by a wide variety of conditions (full medium, full medium lacking growth factors / serum, minimal medium / starvation and pPARγ induction by Thiazolidinedione) and found no change in indcution levels (Fig. 8). Also, we show that the promoter is active only if p65, a component of NF-κB, localizes to the nucleus [5](Fig. 9).
HEARTBEAT predicted promotersBBa_K203125, BBa_K203126
Promoters predicted by HEARTBEAT underwent rough characterization by automated plate fluorescence reader/TECAN and were shown to be upregulated by Sterol depletion approx. 10-fold, as compared to cells in a surplus of Sterol and LDL.. Targeting SignalsMain article: Outlook and Summary Protein domains for targeting to sub-cellular compartments. We intend to use this to visualize multiple pathways. All parts below were characterized in devices consisting of BBa_I712004 (CMV), the targeting signal, BBa_K203103 (GFP) and BBa_K203102 (PolyA signal/terminator). BBa_K203000Shown to localize proteins to the plasma membrane.
BBa_K203101Shown to localize proteins to the ER membrane. An expression device consisting of Part:BBa_I712004 (CMV) Part:BBa_K203103 (GFP), Part:BBa_K203101 (Sar-1) and Part:BBa_K203102 (terminator) was assembled for this characterization.
References[1] Tornoe J. Generation of a synthetic mammalian promoter library by modification of sequences spacing transcription factor binding sites. Gene 297: 21-32 (2002). [2] Edelman G. M., Meech R., Owens G. C. & Jones F. S. Synthetic promoter elements obtained by nucleotide sequence variation and selection for activity. PNAS 97: 3038-43 (2000). [3] Ogawa R. Construction of strong mammalian promoters by random cis-acting element elongation. Biotechniques 42: 628-632 (2007). [4] Rattner A. NF-κB activates the HIV promoter in neurons. EMBO 12: 4261–4267 (1993). [5] Geng H., Wittwer T., Dittrich-Breiholz O., Kracht M. & Lienhard Schmitz M. Phosphorylation of NF-kB p65 at Ser 468 controls its COMMD1-dependent ubiquitination and target gene-specific proteasomal elimination. EMBO reports 10.4: 381-386 (2009).
|
|
 "
"