Team:Newcastle/Labwork/25 August 2009
From 2009.igem.org
Formal Lab Session - 25th August 2009
Overview
- Stochastic Switch Team - carried out mini-preps of the BioBricks BBa_R0062, BBa_R0079, BBa_C0161, BBa_C0179, and BBa_J44000 (set up yesterday) and also carried out digests on Biobrick BBa_C0178 mini-preps
- Metal Sensor Team - ethanol precipitated the BBa_J33206 midi-prep samples
- Sporulation Tuning/Chassis Team - froze down the cwlD, and sleB/cwlJ mutants for storage following their success
Stochastic Switch team
Summary
Today we decided we should redo the minipreps from last weeks biobricks using the overnight cultures we set up yesterday. For the miniprep procedure we followed the extra guidelines that Chris advised us about. The final rehydrated plasmid minipreps were put into the -20 freezer in the 'iGEM plasmid minipreps' box. We also did a restriction digest of the two miniprep samples for biobrick C0178 and ran this on a 0.8% agarose gel. This went well, both samples having the fragment sizes expected. Tomorrow we will be carrying out another restriction digest of todays minipreps, however to save time tomorrow, we poured a gel and put it into the fridge ready to load the samples tomorrow. In order to make sure the prepared gel did not dry out, a small amount of 1% TAE was poured onto the gel, which was wrapped in cling film, clearly labelled and put into the fridge.
Restriction digest
Today we did restriction digest for a miniprep prepared for BBa_C0178. We prepared DNa ladder to load to the gels. At the end of the day, we prepared our gel to run tomorrow.We followed the restriction digest protocol for our two miniprep prepared for BBa_C0178 and used the followings to prepare the solutions.
- H20, 6.5 ul
- 10x Buffer, 2ul
- DNA from minipreps, 10ul
- EcoRI, 0.5 ul
- PstI, 0.5 ul
- BSA, 0.5 ul (It was diluted 100 times)
We gave a quick pulse to mix everything in the tubes before we put them into 37C incubator for an hour.
Running the gel
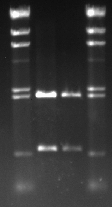
Metal container team had already prepared the gel for us. We placed the gel tray into the gel electrophoresis device and top it up with 1xTAE buffer. 9ul of DNA from each tube is combined with 1ul of buffer and loaded into the wells. DNA ladder was loaded into the 1st and the 4th wells, DNAs were loaded into the 2rd and the 3rd wells. Restriction digest should have cut the plasmid(pSB1A2) into two pieces, plasmid and the the biobrick, from EcoRI and PstI sites. pSB121A2 is 2079bp and BBa_C0178 is 609bp long. As it can be seen from the results below, they match with the HindIII markers.
Preparing the DNA ladder
We diluted the Lambda DNA/HindIII marker 10 times with the loading dye as below and placed the tube containing the mix into the fridge
- 360ul loading dye
- 40ul Lambda DNA/HindIII
Metal Sensor Team
Introduction
Friday's lab session saw the DNA gel electrophoresis of the mini-preps of the BioBrick BBa_J33206 and it's plasmid pSB1A2; both cut and uncut with restriction enzymes. The previous lab session also saw a midi-prep of the BBa_J33206 BioBrick which was then transferred to the freezer for storage.
However not everything is as straightforward as it seems; the bands exhibited when restriction enzyme-treated mini-prep samples were run on the gel on Friday weren't what was expected. Whilst the backbone fragment looked realistic, the small insert fragment looked significantly smaller than it should have done. As suggested in the previous lab session this result could be due to the efficiency of the restriction enzymes, which came from previous aliquots (they did not come straight from the original container meaning they may not work as well)
Friday's lab session also involved Team Newcastle's first attempt at carrying out a PCR reaction. To prove that the plasmid pGFP-rrnB had truely integrated into the amyE gene in the Bacillus subtilis genome (in those B. subtilis which had been transformed), the Metal Sensor team carried out PCR reactions using primers which correspond to regions within the amyE gene in the Bacillus genome. Any PCR products that arise from the different samples we used can determine whether the plasmid has truely integrated.
In any case today's work involves removing the ethanol from the DNA and concentrating it (as observed in the midi-prep protocol). We also hope to make some agarose gel up and run the digests of the midi-prep samples on it through DNA gel electrophoresis. In addition to these tasks, we hope to prepare the PCR 'results' and run them on the gel in DNA gel electrophoresis; once analysis has been made and the verdict given we can decide on where we go from here.
Practical Outline
These are the tasks that the Metal Sensor team are aiming to complete by the end of the day:
- Run the PCR 'products' on agarose gel (DNA gel electrophoresis)
- Carry out ethanol precipitation of midi-prep DNA samples and concentrate them.
- Treat the midi-prep DNA samples with restriction enzymes and run them on the gel also
Procedure
PCR Products analysis
We prepared the PCR reaction products in the way in which we would treat normal DNA samples, which can be found in the DNA gel electrophoresis protocol, i.e. 9 microlitres of DNA to 1 microlitre of dye.
The gel used was 0.8% agarose gel which had been made on Friday's lab session and been stored in the fridge for today's use. It contains 2 sets of wells which means that there is more than 24 wells for the samples to loaded into. Once the gel + tray had been placed into the electrophoresis apparatus the DNA was loaded into the wells. The gel was then run for 45 minutes under 90mv.
The results of the gel can be seen in the results section.
Ethanol Precipitation and Concentration of midi-prep DNA
To work on the midi-preps of the BBa_J33206 and pSB1A2 DNA without the risk of any present ethanol disturbing the process, the ethanol was removed and the DNA concentrated according to GenElute's MidiPrep protocol. This is a summary of the steps taken:
- 0.1ml of 3.0 Sodium acetate was added to the midi-prep tube before 0.7ml of isopropanol was also added. The solution was mixed well by inverting the Eppendorf tube before centrifugation at 13,000g for 30 mins was carried out.
- The supernatant was removed gently before 1.5ml of 70% ethanol was added. Centrifugation for 10 minutes then followed.
- The supernatant was poured off the sample gently before the Eppendorf tube underwent drying in the Speed Vac machine (the protocol can be found here). Once dried out the DNA was resuspended in 50ul of water.
With the midi-preps concentrated, the DNA was then transferred to the -20C freezer for storage. The Metal Sensing team wanted to get the digests carried out today but time ran out.
Unfortunately because of time restrictions the Metal Sensing team did not get to carry out the enzyme digests of the midi-prep DNA. Facing this scenario we decided to make the agarose gel today and then store it in the fridge. The restriction enzyme digests of the midi-prep along with analysis through DNA gel electrophoresis will be resumed tomorrow
Results
PCR Product analysis
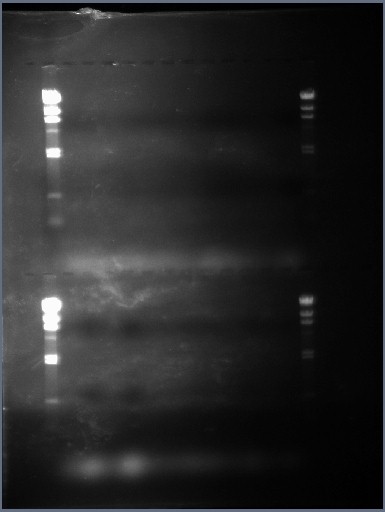
The image above is a photograph taken of the gel (under UV light) with the PCR prodict samples in the wells. It is clear that none of the PCR reactions yielded any results at all, at least that's what the gel is telling us. The HindIII DNA ladder, which is a comparison tool to measure the band sizes formed in the gel by DNA samples, was present in the 4 cases it was inserted so there was no problem with nuclease activity within the gel.
Another thing to note is that there are two white, hazy continuous bands within the gel; one of them is located at the bottom of the first set of wells and the second band is located at the bottom of the second set of wells. These are thought to be the primer dimer complexes formed by the dNTP's. This means that the dNTPS hadn't been accidently missed out.
What to do next?
In light of the PCR reaction failure the Metal Sensing team will need to carry out the PCR reactions on the transformed B. subtilis genome again. This time though, the genomic DNA will be prepared firstly using on the kits found in the lab. This process extracts the genomic DNA of the Bacillus subtilis cells meaning that nothing can get in the way of the PCR reaction from happening. This theory arises from the fact that B. subtilis is Gram positive; it's cell wall may interfere with the PCR reactions. Additionally, the PCR reactions will work better in pure DNA samples.
That will form tomorrow's task. In the meantime, the metal sensor team prepared the overnight cultures of the wild type Bacillus subtilis cells to be used in the genomic DNA preparations tomorrow. To 2 x test-tubes 1.5ml of LB was added and, under aseptic technique, 2 colonies of wild type B. subtilis was extracted from the plate and added to each of the tubes. These test-tubes were then transferred to the shaking incubator at 37C for an overnight growth.
Sporulation Tuning/Chassis Team
Following the success of the treatments of the cwlD, and sleB/cwlJ spores, we decided the freeze down the strains observing Phil's protocol for freezing strains into the TPA collection.
| Recordings | |
|---|---|
| | |
| 2486 | cwlD mutant (treated with lysozyme) |
| 2487 | cwlD mutant (treated without lysozyme) |
| 2488 | sleB/cwlJ mutants (treated with lysozyme) |
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
News
Events
- 20 – 21 June 2009 - Europe workshop (London)
- 23 – 24 June 2009 - UK iGEM meetup (Edinburgh)
- 23 October Practice Presentation (Newcastle)
- 23 October T-shirts are ready
- 27 October Practice Presentation (Sunderland)
- 27 October Poster is ready
- 30 October – 2 November 2009 - Jamboree (Boston)
Social Net
 "
"