Team:UNIPV-Pavia/Notebook/Week3Jun
From 2009.igem.org

|
|
|
|
|
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
Week from June 15th, to June 21st, 2009
 Previous Week
Previous Week
|
Next Week

|
June, 15th
- This week we wanted to complete the assembly of an aTc sensor and to test if lacZ works. Sequence analysis cannot be efficiently performed on I732017 or on A3 because of lacZ coding sequence length (>3 Kb), so we decided to check lacZ integrity assembling a constitutive promoter upstream of A3 and testing the ligation on X-Gal plates.
- We already had J23100, J23101, A3 and A4 at -20°C. Digestions:
| J23100(S-P) | J23101(E-S) |
| A3(E-X) | A4(X-P) |
- We ran/cut a 1% gel for J23100(S-P), A3(E-X) and A4(X-P). All the desired bands had the expected sizes and had been purified from agarose gel. Unlikely we had a low yield for A4(X-P) extraction...
- We ran a 2% gel to isolate J23101(E-S) insert, but unfortunately we could not see any band at the expected size of 35 bp...
- We planned not to perform any ligation: unlucky day for them!:(
- NOTE: we did not re-try to extract J23101(E-S) because it is too small.
Our A4 ligation is composed by RBS-tetR-TT-Ptet , so it can be considered as a constitutive promoter (Ptet) with an additional "non functional" part (RBS-tetR-TT). So it can be easily used as an insert for assembly because of its 900 bp size, that allows it to be efficiently isolated and extracted from a gel!
- For this reason, we infected 5 ml of LB + Amp with 15 ul of A4 glycerol stock (X4 falcon tubes). The following day we plan to cut A4(E-S) (X2) to use it as a constitutive promoter for A3(E-X), while we plan to cut A4(X-P) (X2) to ligate it to J23100 to constitute an aTc->PoPS inducible device. We performed all the inocula X2 because we wanted to be sure to have a sufficient yield for ligations.
- We stored J23100(S-P) and A3(E-X) at -20°C.
June, 16th
- Miniprep for A4 (X4 overnight cultures).
- Digestions:
| A4(E-S) (X2) | A4(X-P) (X2) |
- Gel run/cut; gel extraction.
- Ligations:
- A5 = A4(E-S) + A3(E-X) in pSB1AK3
- A6 = J23100(S-P) + A4(X-P) in J61002 without RFP protein generator
- We incubated the two ligation reactions overnight at 16°C.
June, 17th
- We prepared three X-Gal plates (of LB + Amp).
- We transformed the two ligations (1 ul) in TOP10 E. coli.
- We plated A6 transformed bacteria + SOC in a non-X-Gal plate.
- We plated A5 transformed bacteria + SOC in two X-Gal plates: we plated 100 ul in the first plate (called A5) and 200 ul in the second plate (called A5bis) in order to check for blue colonies the following day.
- We also plated 20 ul of E0240 glycerol stock in the third prepared X-Gal plate, in order to have a negative control to check if TOP10 E. coli actually don't express beta-galactosidase.
- Expected results:
- Plates A5 and A5bis should show blue colonies;
- Plate E0240 should remain colorless.
June, 18th
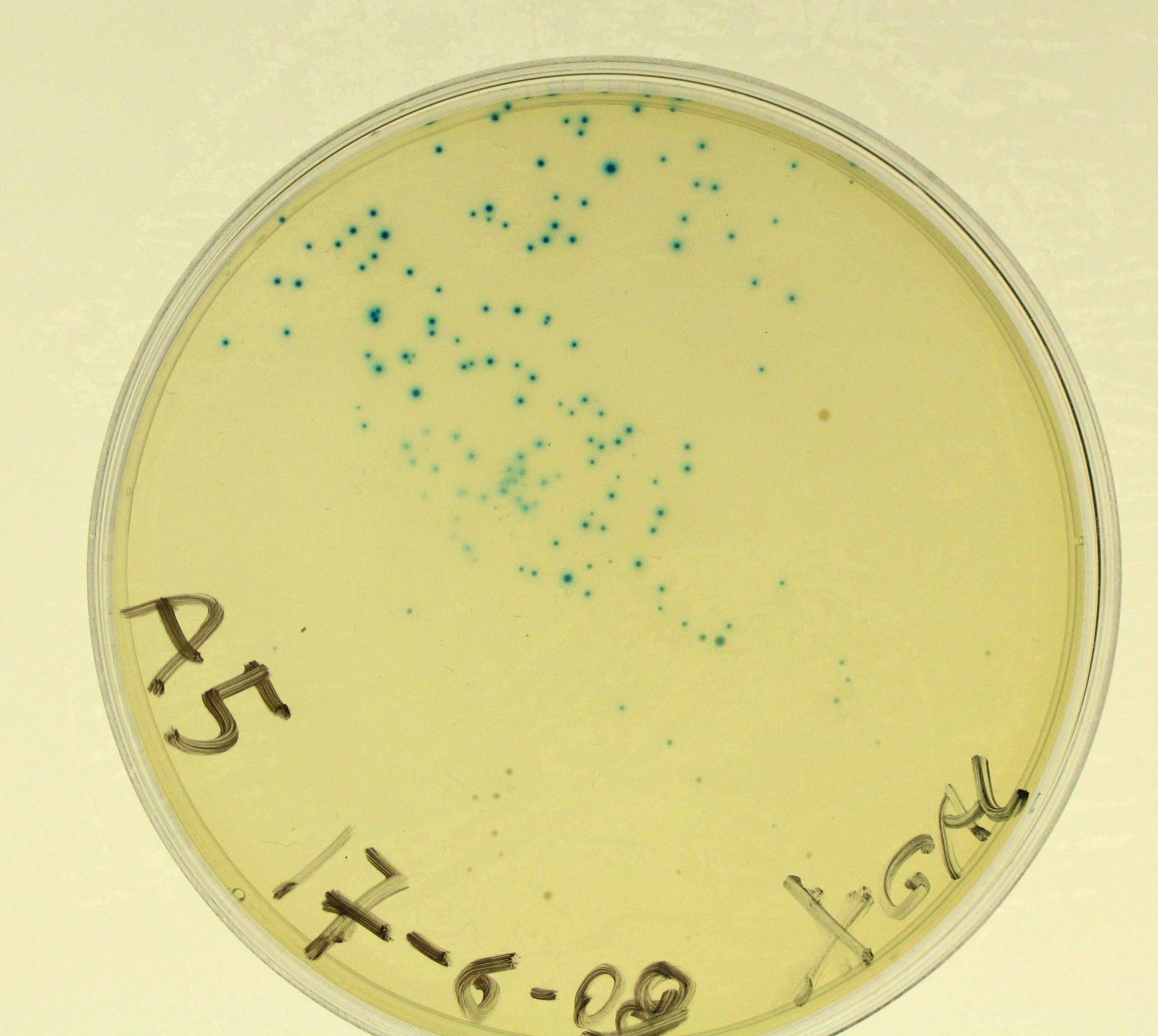
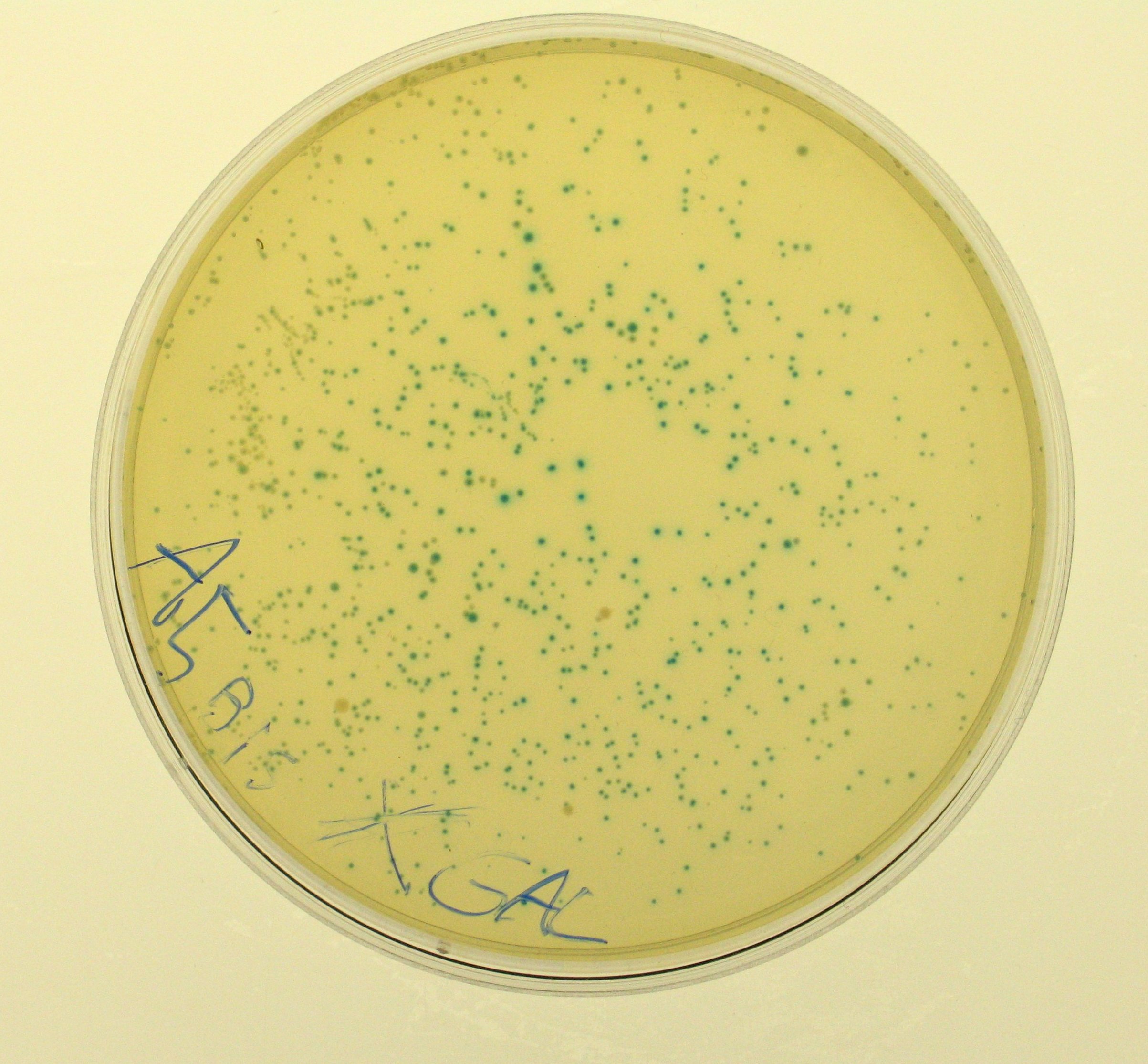
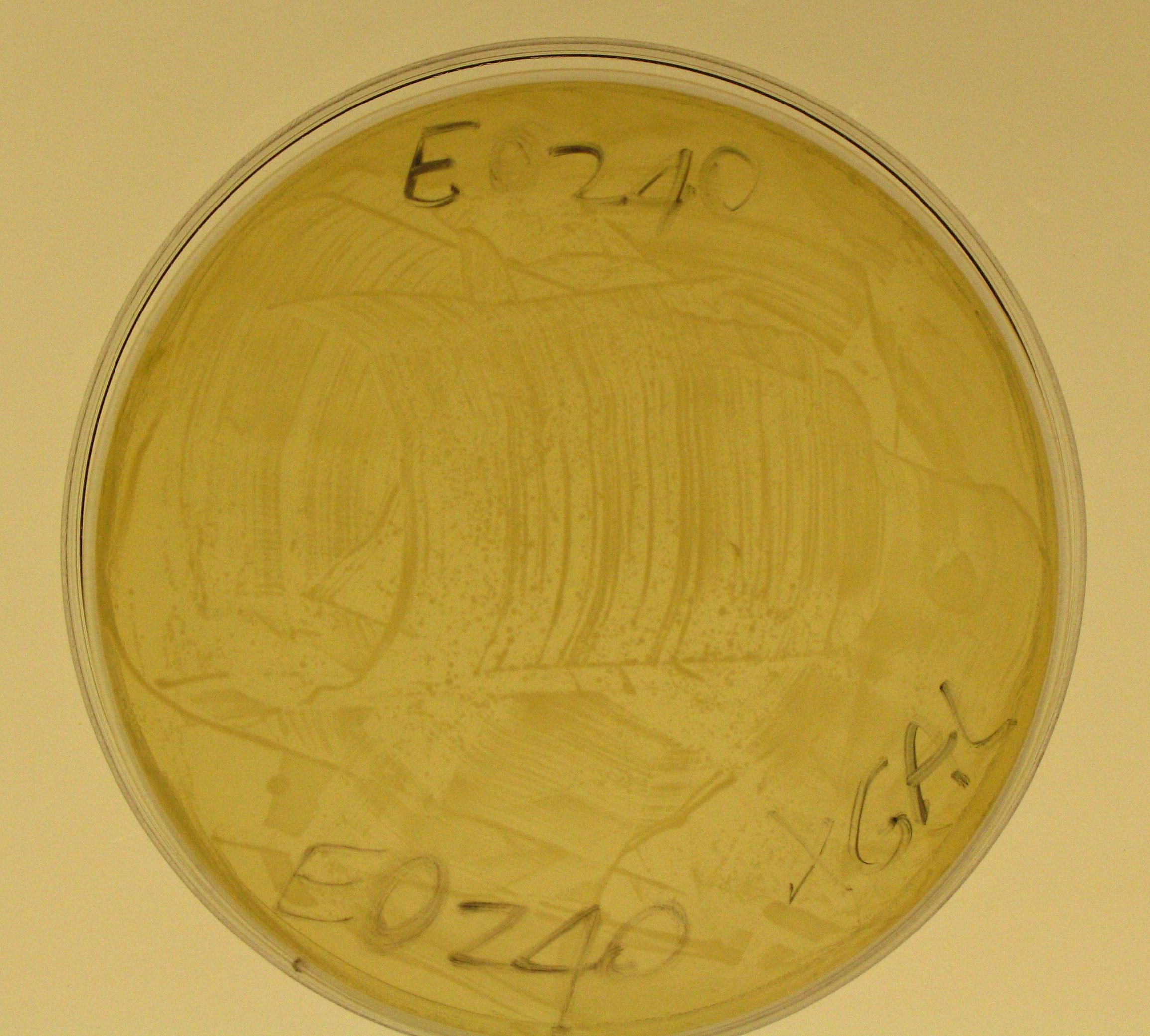
- Plate results:
- A5 showed blue colonies;
- A5bis showed blue colonies;
- E0240 did not show any blue colony.
- So, we can say that TOP10 E. coli don't express any beta-galactosidase and our lacZ gene works!
|
|
|
|
- We picked a blue colony from A5bis plate and infected 5 ml of LB + Amp. We incubated the inoculum overnight at 37°C, 220 rpm.
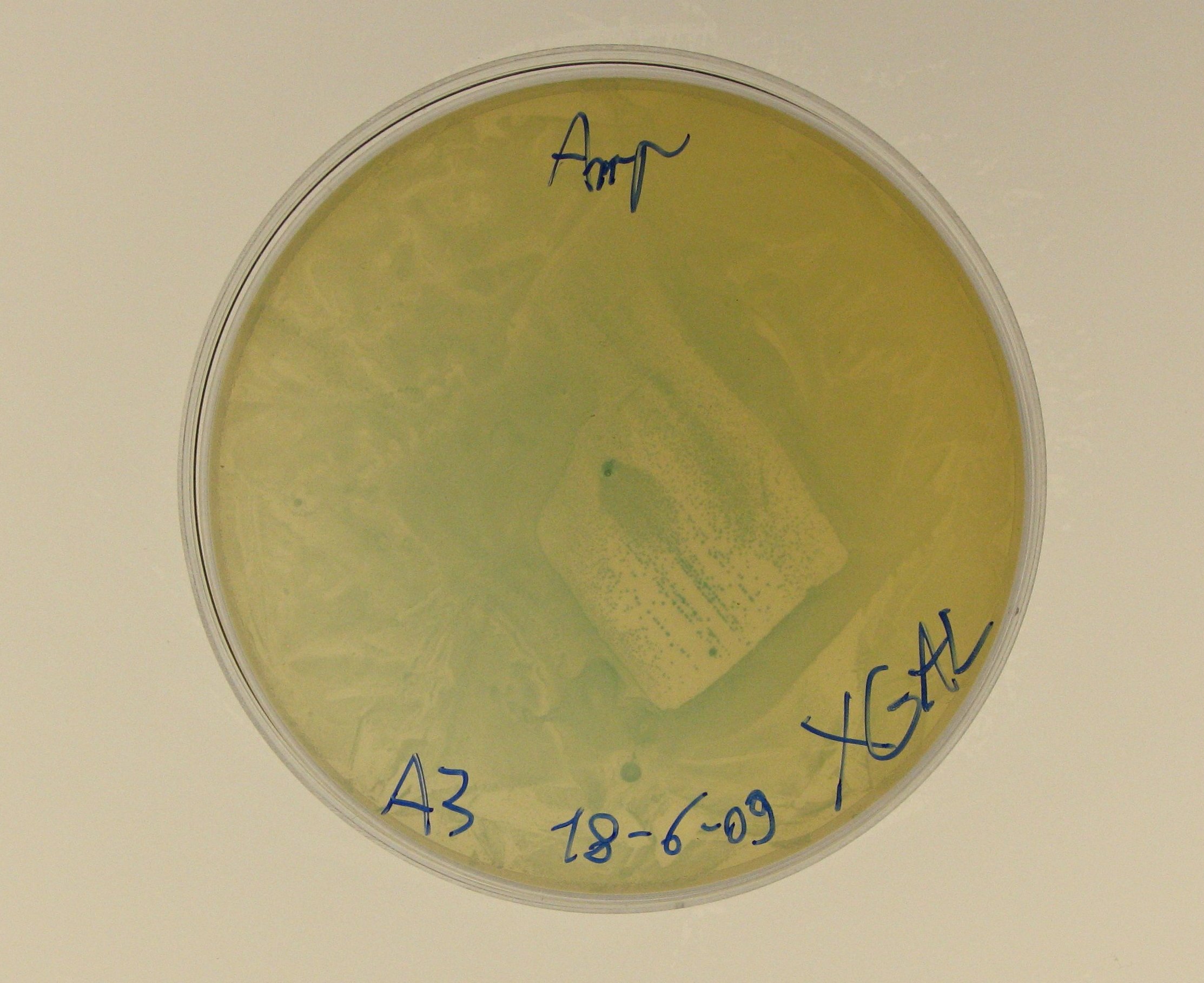
- We decided to perform the X-Gal test on A3, which is the promoterless RBS-lacZ-TT in pSB1AK3:
- we prepared another X-Gal plate (LB + Amp),
- we mixed 20 ul of A3 glycerol stock with 100 ul of SOC medium,
- we plated the diluted A3 culture and incubated the plate overnight at 37°C.
- Colony PCR for A6 plate: we screened six colonies. We let the picked colonies grow in 1 ml of LB + Amp, 37°C, 220 rpm, waiting for the screening results.
- We had a problem with the thermal cycler, it gave an error after about 20 min of the reaction and we had to restart it. At the end of the reaction we noticed that the cap of A6-2 vial had been melted and almost all the reaction evaporated...What a beautiful PCR! isn't it? @_@(...)
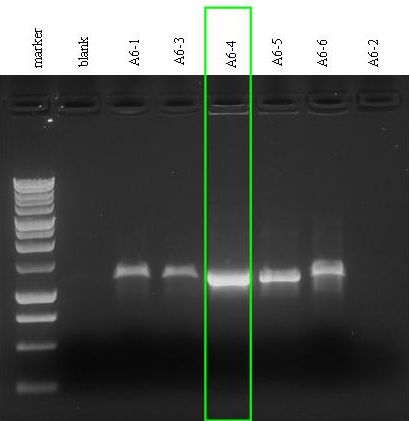
- Electrophoresis for the PCR vials.
|
|
- Gel results: the almost evaporated reaction (A6-2) did not show any amplicon (lane 8). All the other reactions showed the amplicon with the expected size (~1100 bp).
- We chose the 4th colony (A6-4): we filled A6-4 incubated culture with LB + Amp to reach a total volume of 5 ml and let it grow overnight at 37°C, 220 rpm.
- We infected 5 ml of LB + Amp with 15 ul of these glycerol stocks (to prepare samples for sequencing):
| K131009 | A3 | A4 |
- So, in summary, we have 5 overnight cultures:
- K131009 (from glycerol)
- A3 (from glycerol)
- A4 (from glycerol)
- A5 (from A5bis overnight plate, colony randomly chosen)
- A6 (from A6 overnight plate, after colony PCR)
Preparation of experiment with Tecan F200
- We picked two colonies from E0240 overnight plate and infected two falcon tubes containing 5 ml of LB + Amp. We incubated the inocula at 37°C, 220 rpm for 2 hours before the test.
June, 19th
- We prepared glycerol stocks for A5 and A6.
- A3 plates incubated yesterday showed blue colonies, even if LacZ was not preceded by any promoter in the plasmid.
- This result is strange, but we noticed that Caltech iGEM 2008 team had a similar outcome, performing the beta-galactosidase assay on a similar BioBrick https://2008.igem.org/Team:Caltech/Project/Lactose_intolerance: they found significant levels of beta-galactosidase in a high copy number plasmid containing RBS-lacZ-TT. Caltech wrote that it may be caused by 'spurious transcription' amplified by a strong ribosome binding site and high copy plasmid (from their wiki).
- Anyway, we now know that our lacZ sequence is correct and it will be put under the control of the desired promoter during the following weeks. Unfortunately, because of the spurious expression of lacZ, we won't be able to perform a blue/white colony screening after the ligations that involve lacZ gene. For this reason, we are not sure that the blue colony picked on June 18th from A5bis plate is correctly ligated! We decided to purify it and to sequence it anyway. (the plasmid contained in this colony will be called "A5?")
- Miniprep for A3, A4, A5, A6 and K131009.
- We sent the purified plasmids to BMR Genomics for sequencing.
- We received a gift from Bologna iGEM team: J23118-B0034-C0012-B0015 in pSB1A2! (we will call it BOL1) Thank you very much!!!;)
June, 20th
Spring Workshop in London.
June, 21st
Spring Workshop in London
 Previous Week
Previous Week
|
Next Week

|
 "
"