File:Allignments fMT.jpg
From 2009.igem.org
Size of this preview: 800 × 421 pixels
Full resolution (955 × 502 pixels, file size: 157 KB, MIME type: image/jpeg)
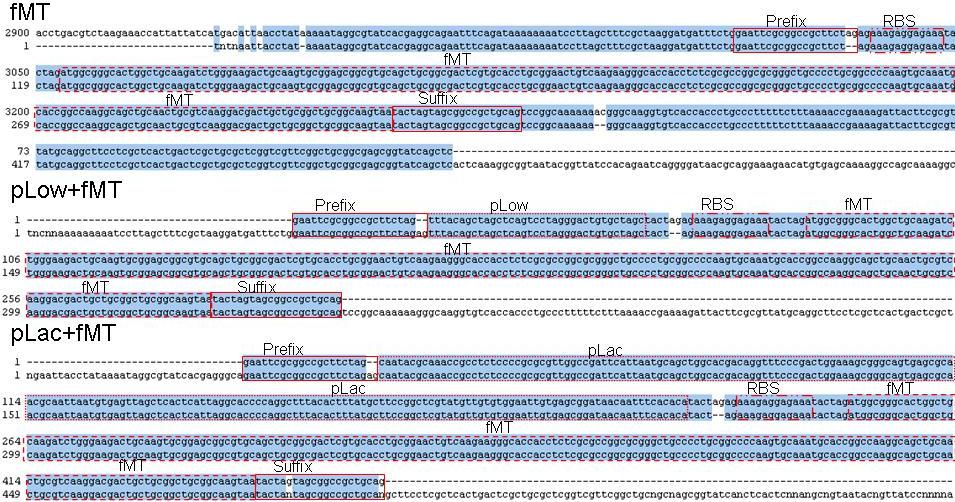
Allignments of the metallothionein fMT constructs, above is In silico construct, below is sequence determined with VF2 primer. The first is without promotor (), second with constitutive promotor () and the third with IPTG inducible promotor (). RBS is . In combining the promotor and the coding sequence the Partregistry seems to add two unexpected additional bases (ag) between the two, missing in sequenced sample. In preparing the In silico construct a different prefix (lacking two bases (ag)), seen in sequenced samples, was used. The n's in the suffix of were determined to be a g as expected when giving a closer look into the chromatogram.
File history
Click on a date/time to view the file as it appeared at that time.
| Date/Time | Thumbnail | Dimensions | User | Comment | |
|---|---|---|---|---|---|
| current | 18:37, 10 September 2009 |  | 955×502 (157 KB) | Wilfred (Talk | contribs) | (Allignments of the metallothionein fMT constructs. The first is without promotor (BBa_K190019), second with constitutive promotor BBa_J23109 (BBa_K190031) and the third with IPTG inducible promotor BBa_R0010 (BBa_K190032). RBS is BBa_B0034. In combining t) |
File links
The following page links to this file:
 "
"