Team:Bologna/Lab-Notebook
From 2009.igem.org
Revision as of 09:34, 29 September 2009 by Marco.cavina (Talk | contribs)
| HOME | TEAM | PROJECT | MODELING | WETLAB | LAB-NOTEBOOK | DRY LAB | SUBMITTED PARTS | HUMAN PRACTICE |
|---|
Week 1: from 07/20/09 to 07/24/09
- Trasfomation of PSB1K3, PSB3K3, PSB1A2, PSB4A3(2007). Only the last one seems to work. We also use PSB1A2.
- Trasformation and Amplification of A, B, C:
A) PSB3K3(LMC) + 1429 + RBS + LACI
B) PSB1A2(HC) + 2547 + Ox + RBS + GFP + T
C) PSB3K3(LMC) + 1429 + RBS + GFP + T
- Working (using DH5alfa) on:
A + B ---> Evaluation GFP fluorescence imaging comparing with the rRBS one
B ---> Evaluation GFP fluorescence imaging
C ---> Evaluation GFP fluorescence imaging
B+C ---> Evaluation ΣGFP fluorescence imaging
Week 2: from 07/27/09 to 07/31/09
- Sequences suspension to have 100 μM conc. (EP digestion and trasformation in a HCN plasmid with AMP)
| DNA sequences | Plasmid |
|---|---|
| H2O mQ == 4.5 μM | H2O mQ == 19.5 μM |
| Sequence == 20 μM | DNA (miniprep) == 5 μM |
| Buffer == 3 μM | Buffer == 3 μM |
| BSA == 0.5 μM | BSA == 0.5 μM |
| EcoRI enzyme == 1 μM | EcoRI enzyme == 1 μM |
| Pst1 enzyme == 1 μM | Pst1 enzyme == 1 μM |
| TOT: 30 μM | TOT: 30 μM |
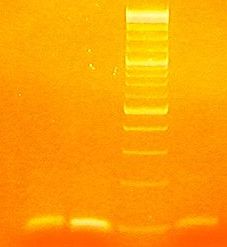
Gel Run:
1.CIS
2.TRANS 4
3.MARKER
4.TRANS 7
5.PSB1A2
Week 3: from 08/3/09 to 08/7/09
- Amplification of CIS, TRANS 4 and TRANS 7 using PSB1A2. Inoculation and digestion.
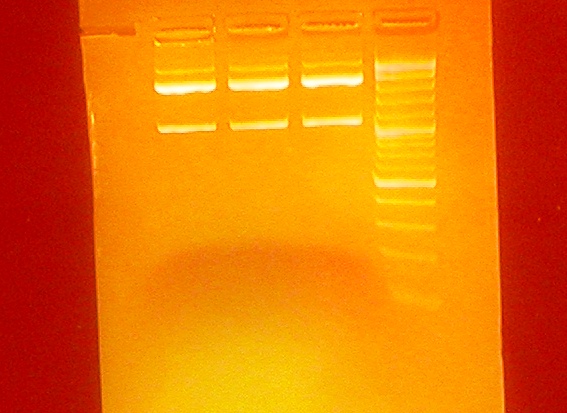
- Unexpeted results from the gel run in all three sequences (3000, 2200, 1000 bp instead of 2100, 2000, 100 bp):
- Done the ligation again whit the same bad results.
- Done some test fluorescence measurements using GFP in plasmids with different copy number.
Week 4: from 08/10/09 to 08/14/09
HOLIDAY!!!
Week 5: from 08/17/09 to 08/21/09
- Done again the amplifications of CIS, TRANS 4 and TRANS 7 using PSB1A2 to undestand which is the problem we have had during the third week. Achieved the same wrong results.
- We decided to sequence the 1000 bp strand to understand what it really is. We sent our material to the Bmr-genomics in Padova for the dna sequencing.
Week 6: from 08/24/09 to 08/28/09
Bad news for our team:
- Re-Annealing of our sequences and after have done the PRC-Colony we discovered that the colonies were green. Probably due to a GFP contamination. The DNA sequencing coming from Padova confirmed our theory: in the 1000 bp strand there was the presence of GFP protein and no sign of our CIS or Trans sequences.
Week 7: from 08/31/09 to 09/04/09
- Done some experiments (trasformation and digestion with the same plasmid, PSB1A2, using different concentration of annealed) to confirm the Gfp contamination. All the gel runs have a 1000 bp strand.
- Investigation to discover where is the contamination.
- Starting use a new plasmid: PSB4A3. Trasfomation and digestion with the CIS sequence. From the gel run came out a 400 bp insert that could be correct (CIS+Primer=100bp+270bp=370bp):
- The same attempt with TRANS 4 and TRANS 7 didn't give us any concrete resulst.
Week 8: from 09/07/09 to 09/11/09
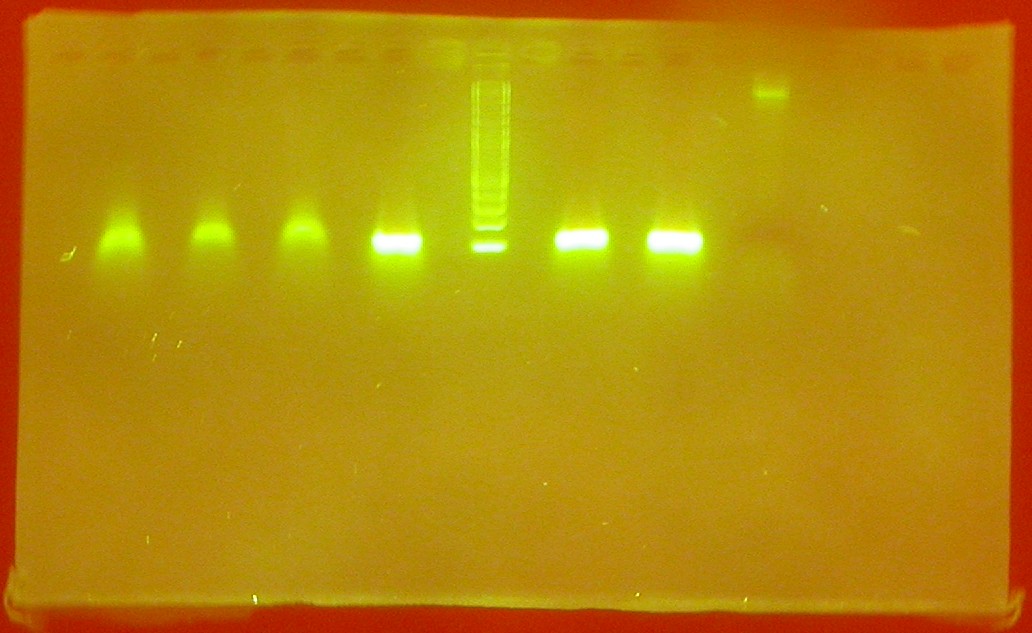
- Primers of our sequences arrived, so we amplified CIS-repressing and TRANS-repressor with a PCR
- Gel run of results of PCR using a stain named GelStar that can detect smaller strains of DNA
- We assumed that some of our ligation problems may be caused by the low enzymes efficiency because of the restriction sites were too near to the extremities of our sequeces. (because of there wasn't enough base pair between the restriction sites and the extremities of our sequeces.)
 "
"