Team:Cambridge/Notebook/Week7
From 2009.igem.org
Categories :
Project :
-
Overview
Sensitivity Tuner
--- Characterisation
--- Modelling
Colour Generators
--- Carotenoids (Orange/Red)
--- Melanin (Brown)
--- Violacein (Purple/Green)
The Future
Safety
Notebook :
Team Logistics :
Week 7
Monday
Wet Work
Amplification
Colony PCR of 8 colonies picked from 391 in pSB3K3 transformants, 8 colonies picked from 374 in pSB3K3 transformants, 8 colonies picked from 384 in pSB3K3, and 1 colony picked from 381 in pSB3K3 transformants confirmed to contain an insert of the right length. Overnight cultures of each of those colonies as well. No 374 constructs were found to contain inserts of the right length, however 4 384 constructs were of the correct length. The 381 construct was confirmed once again to be the correct length.
Overnight,
- Restirction digests of pSB3K3, 380, 382, 385, and 390.
- Colony PCR of 10 new 391 in pSB3K3 transformants, another 374 in pSB3K3 transformants
MelA BioBrick
PCR done of the original MelA and the first MelA intermediate (without the second PstI site), to create the DNA for a PstI restriction digest. After PCR the DNA was run on a CYBRsafe-stained gel and then extracted with the QuiGen kit. A Restriction digest was carried out using the fastdigest buffer and PstI and left for three hours.
We also tested the Primer C again, as it didn't work last time. Three PCR's were done:
- Primer F and Primer C (same aliquot as used last time, to check it wasn't just an experimental error on my part)
- Primer F and a new aliquot of Primer C
- Primer F and revortexed and re-aliquoted primer C from the origonal vial
None of these produced any result - there is clearly an error in the primer that I ordered.
Pyocyanin *NEW*
The following biobricks were transformed into TOP10 and plated out in preparation for pyocyanin biosynthesis:
- I746910 (pSB1A2) - pBAD + B0034
- I723024 (pSB3K3) - PhzM
- I723025 (pSB1A2) - PhzS
- B0015 (pSB1AK3) - double terminator
- B0034 (pSB1A2) - RBS
Dry Work
We found the phzM and phzS pigments in the registry!
- BBa_I723024 (phzM)
- BBa_I723025 (phzS)
This means we can now start working on these pigments, connecting them to processing and logic systems.
Development of Matlab programs to automatically process plate reader data was continued.
Tuesday
Wet Work
Amplification
Colony PCR of 391 in pSB3K3 showed none had the correct insert. Ligations of 380, 382, 385, 390 into pSB3K3, transformed into Top10.
381 and 384 were tranformed into the arabinose strain in preparation for assaying using the plate reader.
Violcein
James has a plasmid that produces cuiR which when stimulated with C6-HSL acts activates the vioA promoter. We put the vio+ cells into the shaking incubator, preparing to make them competant tmorrow.
Pyocyanin
Transformations of I746910, I723025 and B0034 worked, and were put in overnight culture in preparation for miniprepping.
I723024 and B0015 were transformed again into TOP10.
Dry Work
Wednesday
Wet Work
Amplification
- Overnight colony PCR of 380, 382, 385, 390 in pSB3K3.
- Moved 375, 392, 394, 395 into pSB3K3, transformed into TOP10.
- Overnight colonies of 381 in pSBK3K in arabinose strain and 384 in pSB3K3 in arabinose strain in preparation for plate reader run tomorrow, as well as the appropriate controls.
MelA BioBrick
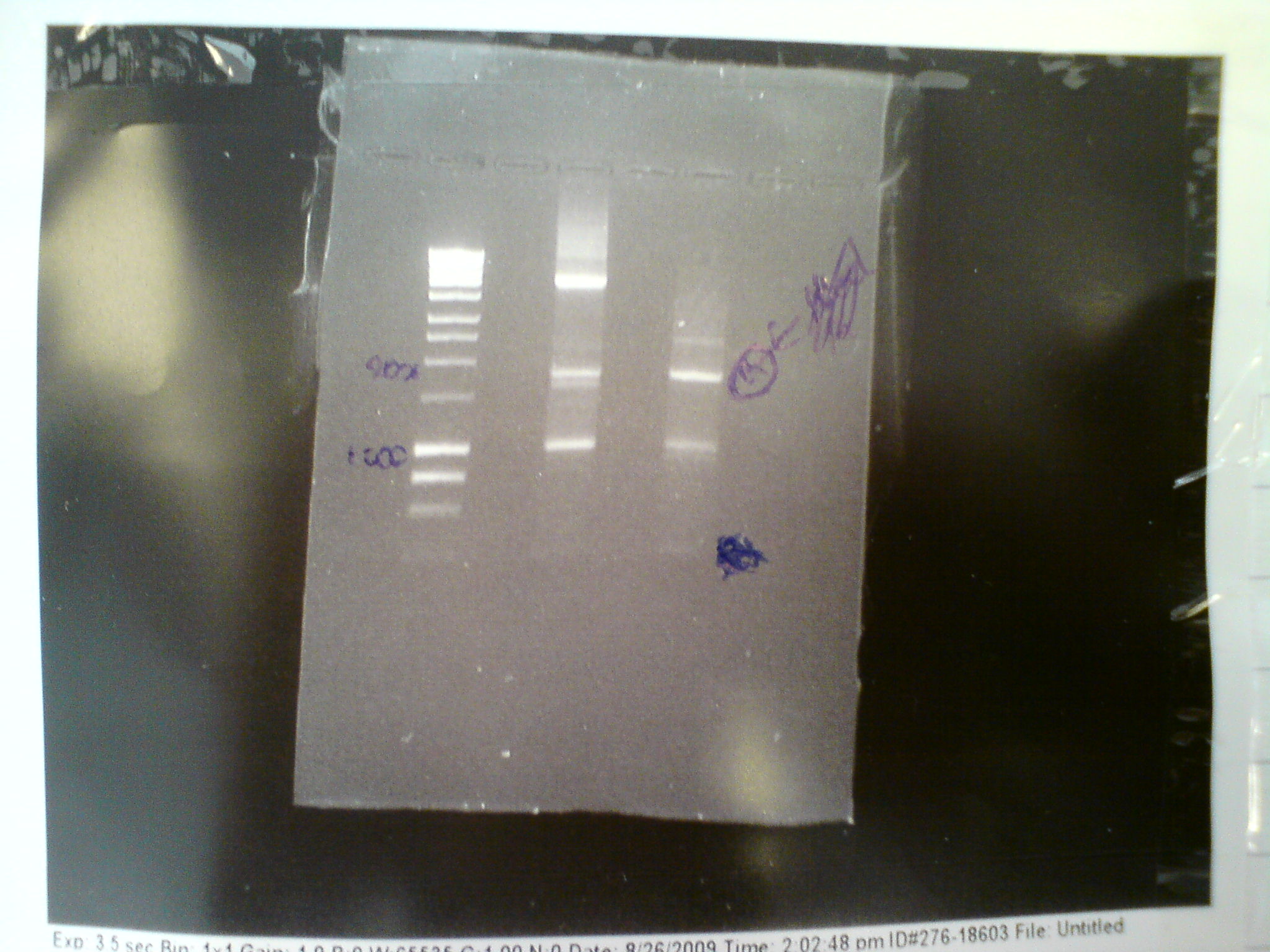
In a final attempt to get Primer C to work, we tried touchdown PCR with the lowest temperatuire 45 degrees C. Primer C and Primer F were used with the origonal melA gene and the end product of the PCR was run on an ethidium bromide gel at 80V for 45 mins:
Product is shown starred in the above gel. The MelA protocol can now be completed.
Violacein
Made competent vio+ cells and transformed them with James's plasmid. Transformed cells were plated on trimethoprim/ampicillin plates and placed in the 37 degree incubator overnight.
Dry Work
Planned standard assemblies for the next month
Thursday
Wet Work
Amplification
- Successfully moved 380, 382, 385, 390 into pSB3K3, cultured colonies overnight in preparation for miniprep tomorrow
- Overnight colony PCR of 375, 392, 394, 395 into pSB3K3.
 "
"