Team:Cambridge/Project
From 2009.igem.org
Categories :
Project :
-
Overview
Sensitivity Tuner
--- Characterisation
--- Modelling
Colour Generators
--- Carotenoids (Orange/Red)
--- Melanin (Brown)
--- Violacein (Purple/Green)
The Future
Safety
Notebook :
Team Logistics :
Project
Introduction
Improving Biosensors
The Parts Registry's repertoire of input-sensitive devices is incredibly varied. Teams have engineered E. coli to be sensitive to a wide range of environmentally significant compounds, including arsenic, mercury, lead, cyanide, etc., to genetically engineer biosensors as an alternative to other technologies. The Cambridge 2009 iGEM team identified two stumbling blocks to biosensor design.
- Output: Previous iGEM biosensor projects have used pH, electrical conductance, and fluorescence as output. However, these reporter mechanisms require further steps to read the output. While this is acceptable for First World applications, for biosensors to have true Third World applications, a simpler output is necessary.
- Response to Input: By utilizing an input-sensitive promoter, the biosensor is limited by the sensitivity of the promoter. For example, the promoter might be sensitive to input concentrations which have no real world meaning. The promoter's sensitivity could be too high, so it reports concentrations below levels of real-world interest. Alternately, the promoter's sensitivity could be too low, so it reports concentrations above those which mark the boundary between "safe" and "dangerous." A second limitation is the the behavior of the PoPS output from the promoter; for example, output may vary linearly with input. This type of response is incompatible with a digital "safe" or "dangerous" output.
Our Solutions
- A Sensitivity Tuner: To avoid being limited to the sensitivity of the promoter and in order to be able to detect distinct concentrations of an inducer using just one promoter, we see the need for a set of sensitivity tuners. These devices allow you to "tune" your biosensor, such that it reports meaningful concentrations of the inducer appropriate to the biosensor's application. The sensitivity tuner also modifies the PoPS output from the promoter's native behavior to a sigmoidal "on" or "off" response pattern.
- Colour Output: What if we could "see" the concentration of an inducer in a sample by a change in colour of the biosensor? Many prey species are brightly coloured, showing off clearly the fact that they are poisonous or otherwise harmful to potential predators, a phenomenon known as aposematism or warning colouration. Humans use colour as a means of conveying information as well - we can "see" if a child has a fever using a thermometer strip and we can "see" the pH of a solution using a pH indicator. Colour can be a meaningful but simple output solution for biosensors, adapting nature's idea of warning colouration.
Project Details
Design
The Product
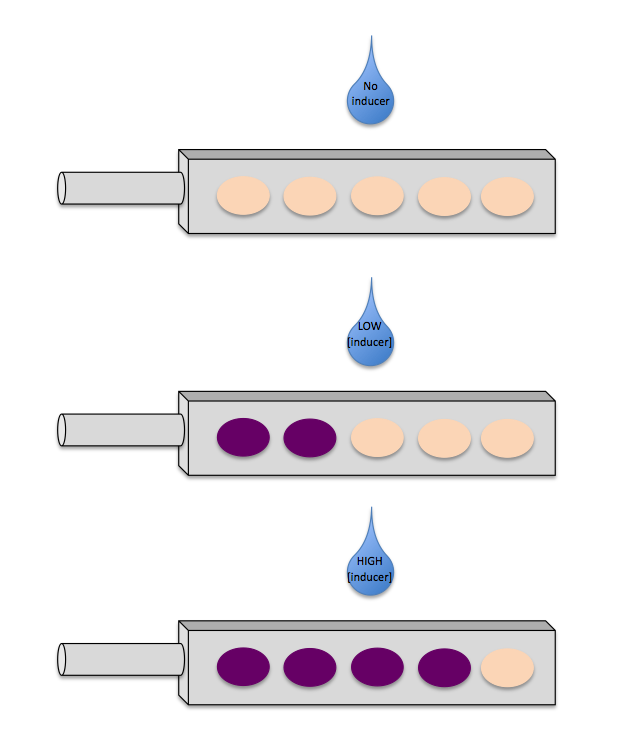
We envisioned a marketable product that reports the concentration of an inducer by colour. Imagine a dipstick with wells, each of which contain pigment-expressing bacteria in response to an inducer. However, each strain is sensitive to a different concentration of the inducer.
The Genetically Engineered Machine
Each bacterial strain is a machine built from a three part system:
- Sensor: Our sensor system is sensitive to different concentrations of an inducer.
- Threshold device: The threshold device is responsible for the sensitivity to the inducer, and acts as an "on" switch to activate pigment production once the inducer has reached a threshold.
- Colour: Pigment production.
Components
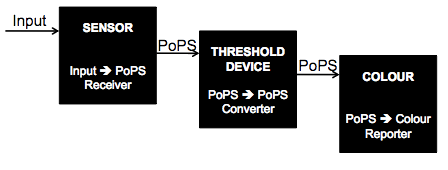
The three part system can be abstracted by the diagram below:
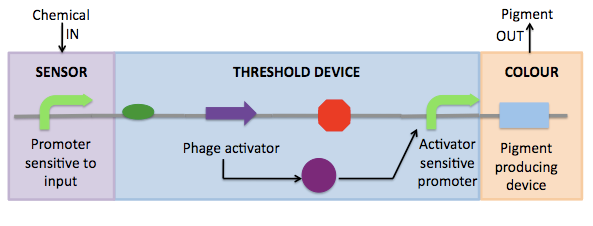
The components of each of the black boxes is as follows:
- Sensor: Our project was output-focused, so we concentrated on the second two parts. For our proof of concept we used an arabinose-sensitive promoter as the sensor. Future extensions of our project would include using our device with other input-sensitive promoters in the registry.
- Threshold Device: This construct is based on Cambridge 2007's amplifiers. It is composed of a gene coding for a phage activator protein and a phage activator-sensitive promoter. With PoPS input, the phage activator gene is transcribed. After translation to produce the phage activator protein, the protein binds to the phage activator-sensitive promoter to activate transcription. Thus it is a device that generates a distinct PoPS out given a PoPS in. With an input-sensitive promoter alone, output generally varies linearly with input--this, at least, is the case with the arabinose-sensitive promoter pBad/Arac (BBa_I0500). However, when a threshold device is placed downstream of the input-sensitive promoter, the output versus input behavior is altered such that output increases dramatically at a certain input, resulting in sigmoidal behavior with a distinctive "threshold." Because there are 3 different activator genes and 5 different activator-sensitive promoters in the registry, 15 combinations are possible, and each has a distinct "threshold" and peak rate of output. This is the basis for our catalogue of threshold devices.
- Colour: Though E. coli does not naturally produce pigment, several other bacterial species secrete antibiotics which happen to be pigmented. We have used pigment-producing operons from other bacterial species and transformed them into E. coli. In particular, we have devoted our summer to 3 different pigment systems:
- Carotenoids: The enzymes required for carotenoid production originally come from Pantoea ananatis, and were available in the registry. We used them to produce orange and red.
- Melanin: The tyrosinase required for melanin production originally comes from Rhizobium etli and produces brown.
- Violacein: The enzymes required for voilacein production originally come from Chromobacterium violacein. The operon can be manipulated to produce voilet, green, and blue.
Experiments
Results
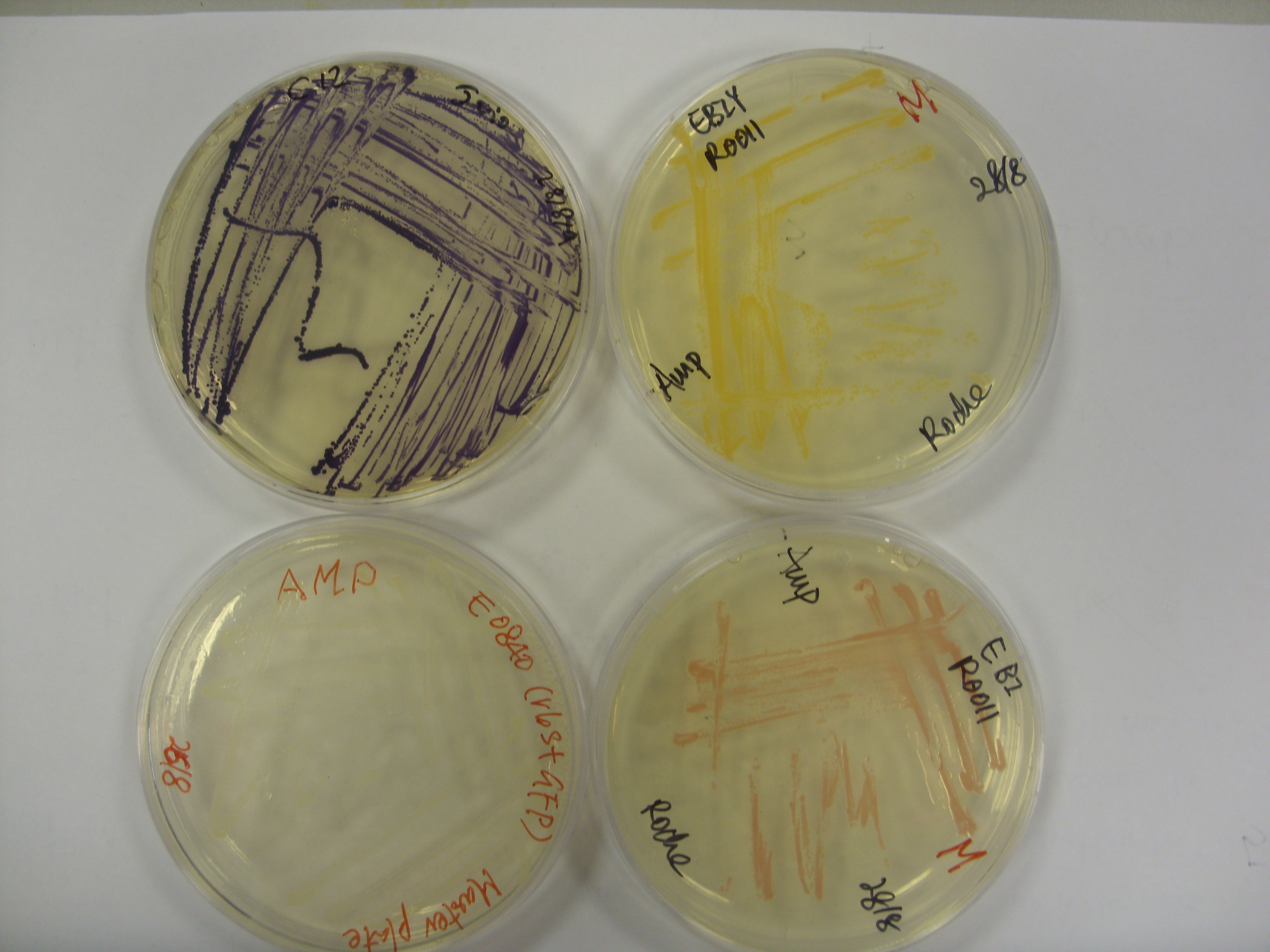
(Preliminary): A few of the pigments we have available:
 "
"