Team:Groningen/Notebook/1 September 2009
From 2009.igem.org
m (→MBP-ArsR) |
m (→fMT) |
||
| Line 203: | Line 203: | ||
====fMT==== | ====fMT==== | ||
| - | *{{ | + | *{{done|check [https://2009.igem.org/Image:Allignments_fMT.jpg sequence]}} |
| + | |||
====SmtA==== | ====SmtA==== | ||
<center>[[Image:Restriction_analysis_SmtA_pLow_(pSB1AC3)_-_01september2009.jpg| 250px]]</center> | <center>[[Image:Restriction_analysis_SmtA_pLow_(pSB1AC3)_-_01september2009.jpg| 250px]]</center> | ||
Latest revision as of 18:43, 10 September 2009
Wet
GVP Cluster
- → DONE Isolate plasmids from o.n. cultures
- → DONE Determine concentrations (and prepare for ligation of GVP behind LAC/pBAD promoters)
- → DONE Make glycerol stocks of pZntR+RBS, pArsR+RBS, pCueO+RBS
- → TODO Make glycerol stocks of pBad/araC, pBad/araC+RBS+GVP, and pLacI+RBS+GVP (both in pSB1A2 and pSB1AC3 plasmid)
- → TODO Choose colonies from plates for growth of E.coli TOP10 with GVP behind LAC/pBAD (+RBS) promoter in pSB1AC3
- → DONE Transform E.coli TOP10 cells with pSB2K3 plasmid from the registry (plate 1, 7C with RFP Coding Device BBa_J04450, and plate 1, 7K with ccdB cell death gene BBa_P1010)
- → TODO Test control of bouyancy in Saline solution
- → DONE Design synthetic DNA for GVP
- → TODO Order synthetic DNA for GVP
- → TODO Order primer for PstI site removal
- → TODO Test promoter strenght compared to BBa_J23101 promoter (Sven)
- → TODO Enter sequences of constructs to Sandbox
Overnight Precultures
All tubes showed growth of E.coli cells, and must have ampicillin resistance on a plasmid(The tubes differed in cell density though, the ones with 10mL LB were less dense as expected). To be sure of correct ligation products in the plasmids an isolation will be performed, followed by restriction analysis.
Overnight Plates
The plates grown overnight showed single colony growth, the plates were stored in the fridge for preculture inoculation.
Transformation
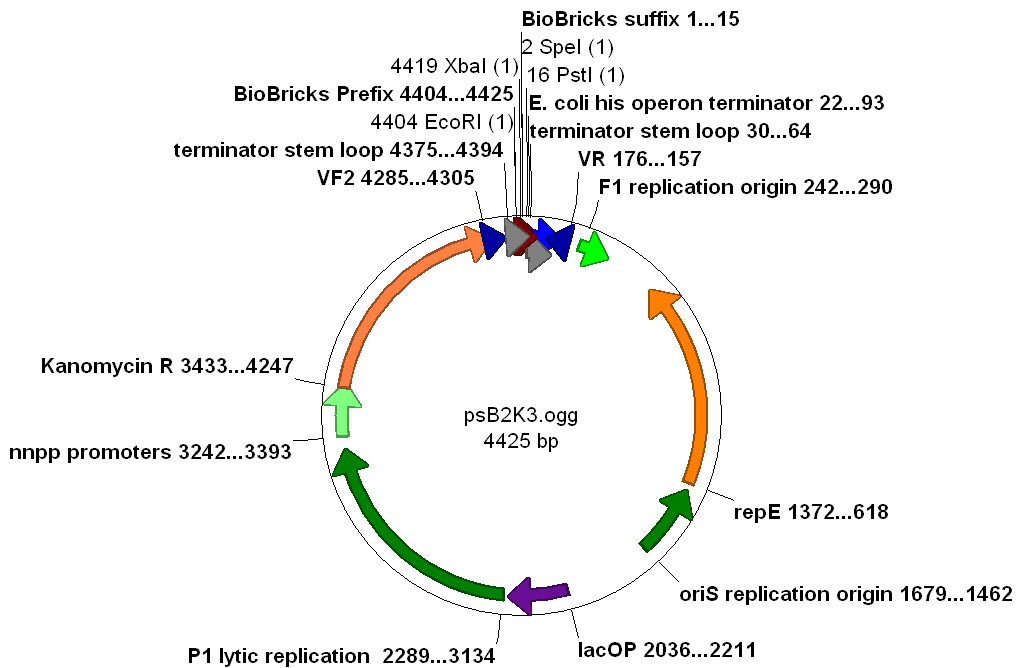
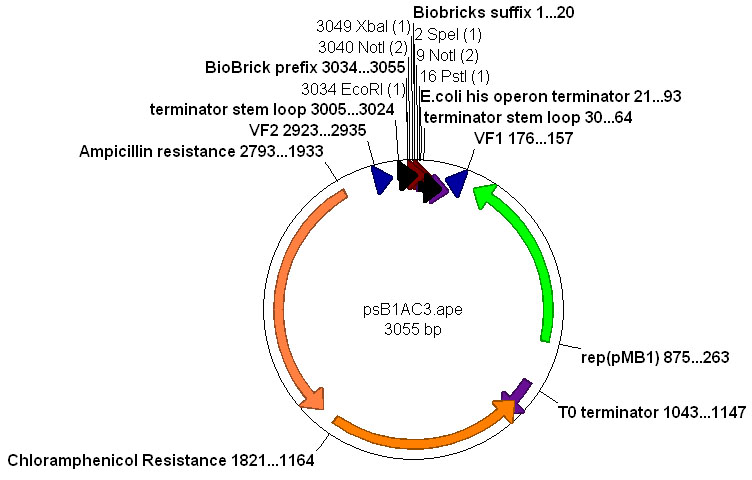
The pSB2K3 vector was chosen to use together with the pSB1AC3 vector to create a two plasmid system. The MEtal promoters with GVP on pSB2K3, and accumulation with transport proteins on pSB1AC3. These two vectors should be compatible with respect to their origins of replication.
- → To the slots 7C (pSB2K3 with RFP reporter) and 7K (pSB2K3 with DEAD gene) on plate 1, 15uL MQ was added to dissolve dried plasmid.
- add 3uL of the dissolved plasmid to 50uL competent E.coli TOP10 cells.
Incubate:
- 30 min @ ice
- 90 sec 42°C
- 2 min @ ice
- add 800uL LB-medium
- incubate for 1 h at 37°C
- plate 100uL on LB-kan100 plates
- plate 100uL of concentrate on LB-kan100 plates
- plates were stored at 37C o.n.
- → Positive control was BBa_J61002-J23101 plasmid, and negative control was MQ.
Plasmid Isolation
Plasmid isolation was performed on the cultures of E.coli TOP10 containing plasmids pSB1A2 with pLacI, andpBad/araC inducible promoters, BBa_J61035 with GVP, and pNL29 with the "Sygma-Aldrich™ GenElute™ Plasmid Miniprep Kit".
- From each tube 8mL of culture was collected in a 2.0mL cup, and the cells were pelleted by centrifugation for 1 min. at max. speed and the supernatant discarded.
- Plasmids were eluted with 40μL MQ and stored in the fridge
Concentration
| Plasmid | Conc. ng/μL | 260/280 | 260/230 | -20 box (michael | Restriction Control |
| GVP | 393.2 | 1.86 | 2.38 | No | See gel |
| pNL29 | 18.2 | 2.11 | 2.01 | No | See gel |
| pSB1A2-pBad/araC | 41.6 | 1.95 | 1.57 | No | See gel |
| pSB1A2-pLacI | 52.9 | 1.97 | 1.83 | No | See gel |
Restriction Control/For Assembly
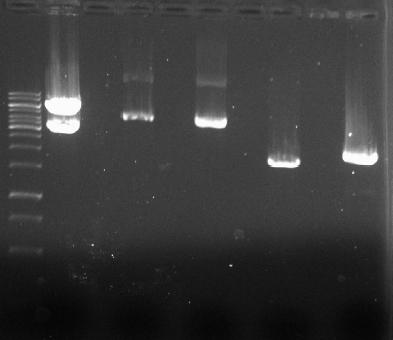
Isolated plasmids were cut with a combination of fast-digest enzymes EcoRI, XbaI, SpeI and PstI to create ligation fragments and control fragments to check the correctness of pSB1A2 pLacI/pBad-araC.
- → From left to right: 1kB ladder, GVP (XbaI,PstI), Empty Slot, pSB1A2-pBad/araC (EcoRI,PstI), Empty Slot, pSB1A2-pBad/araC (SpeI,PstI), Empty Slot, pSB1A2-pLacI (EcoRI,PstI), Empty Slot, pSB1A2-pLacI (SpeI,PstI)
Purification
- → The bands for GVP, pSB1A2-pLacI (SpeI,PstI), and pSB1A2-pBad/araC (SpeI,PstI) were used for purification, even though the pSB1A2-pBad/araC did not seem to be correct.
A "Agarose Gel DNA Extraction Kit" Standard Protocol from Roche Applied Science.
- → In step ten, 30μL MQ was added to the dry pellet and incubated at room temperature.
Concentration
| Plasmid | Conc. ng/μL | 260/280 | 260/230 | -20 box (michael | Restriction Control |
| GVP (XbaI,PstI) | 42.0 | 1.83 | 0.86 | In fridge | See gel |
| pSB1A2-pBad/araC (SpeI,PstI) | 8.2 | 1.78 | 0.17 | In fridge | See gel |
| pSB1A2-pLacI (SpeI,PstI) | 23.6 | 1.80 | 0.40 | In fridge | See gel |
Restriction Control
The first control of pSB1A2 with pBad/araC did not confirm the right fragments of ~2000bp and ~1300bp. As a follow up control plasmid pSB1A2 with pBad/araC from myself and Frans were cut with FD EcoRI and PstI to see if the correct fragments show up on gel.
- → From left to right: 1kB ladder, pSB1A2 with pBad/araC (2x, own), pSB1A2 with pBad/araC (2x, Frans)
Observations
- → The restriction control for pBad/araC in pSB1A2 turned out to be negative. The transformation of E.coli TOP10 with plasmid from the registry seemed to have failed (why was it not noticed straight away?). After a three day search, the cup with dissolved plasmid from plate 3, 1M turned up unexpectedly. A new transformation will be undertaken tomorrow morning!
- → All previous work with pBad/araC has to be checked extensivly to be sure it is pBad/araC!!
Designed DNA
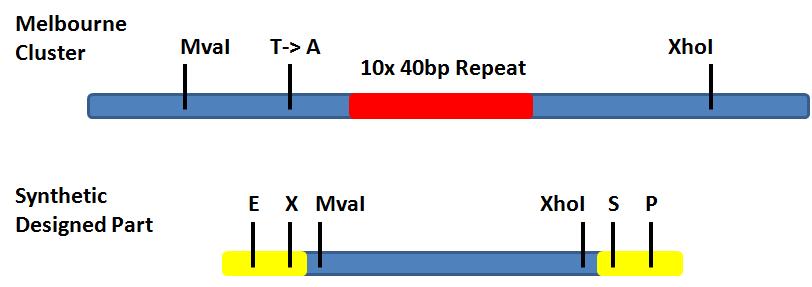
Image representation of our idea:
- → By substituting the part in the GVP biobrick between MvaI and XhoI with our synthetic DNA, the repeat and mutation introduced by Melbourne should be lost.
Transporters
HmtA
PCR1.2 and PCR2 isolated from gel.
PCR3 pcr1.2 and pcr2 used as primers on pBAD-HmtA template in a PCR3 to get HmtA.
PCR1 PCR1.2 used as template in PCR1, F1 + mut1rc
Metal Accumulation
MBP-ArsR
- Check sequence sequence doesn't look all that well (a whole lot of n's, bad quality isolation?
- Put promotors in front --> Work in progress
- Put transporters in back --> Need transporters first!
fMT
- check sequence
SmtA
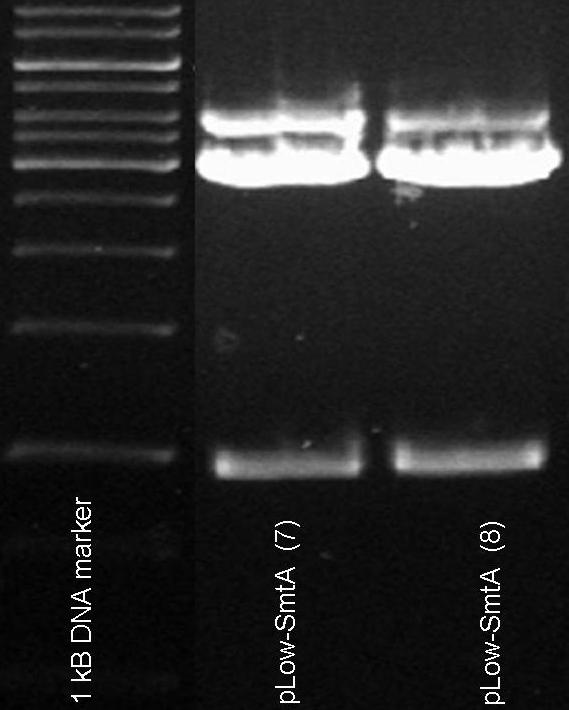
Restriction analysis of the metallothionein SmtA with a low constitutive promotor. Restriction analysis was performed using XbaI & PstI, SmtA+pLow should give a band of ~750 bp, pSB1AC3 ~3000 bp. The lowest band is at a height of ~1000 bp.
Vectors
Dry
|
|
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"