Team:Groningen/Notebook/21 July 2009
From 2009.igem.org
Wet
GVP Cluster
Plasmid Isolation from GVP, J23109, 106, 100 cultures
Done with ...
Concentrations of J23109/100/106 and GVP
BBa_J23109 eluted in MQ
- 41.2 ng/μL
- 1.86 (260/280)
- 1.98 (260/230)
BBa_J23100 eluted in MQ
- 191.4 ng/μL
- 1.89 (260/280)
- 2.30 (260/230)
BBa_J23106 eluted in MQ
- 93.5 ng/μL
- 1.89 (260/280)
- 2.13 (260/230)
GVP eluted in MQ
- 121.2 ng/μL
- 1.90 (260/280)
- 2.33 (260/230)
3A assembly of pSB1AC3, GVP, J23109, 106 and 100
This assembly is performed to place one of the promoters in front of the GVP gene cluster in the backbone pSB1AC3.
Digestion
Mixes:
pSB1AC3
- 2 uL fastdigest buffer
- 1 uL EcoRI (fastdigest)
- 1 uL PstI (fstdigest)
- 5 uL DNA
- 11 uL MQ
GVP
- 2 uL fastdigest buffer
- 1 uL XbaI (fastdigest)
- 1 uL PstI (fstdigest)
- 8 uL DNA
- 8 uL MQ
109, 106, 100
- 2 uL fastdigest buffer
- 1 uL EcoRI (fastdigest)
- 1 uL SpeI (fstdigest)
- 16, 10, 5 uL DNA
- 0, 6, 11 uL MQ
-> Incubation: 1 hour, 37°C
Purification
Done with the GenElute PCR clean-up kit (Roche). Only done with the GVP and AC3 fragments because fragments containing the promoters are too small to be purified in this way.
Concentrations of GVP and AC3 restriction fragments after purification
GVP
- 16.1 ng/uL
- 2.10 (260/280)
- 1.86 (260/230)
AC3
- 19.3 ng/uL
- 1.98 (260/280)
- 2.10 (260/230)
Ligation
Mixes (three reactions):
- 1 uL T4 buffer
- 0.5 uL T4 Ligase
- 0.7 uL AC3 restriction fragments
- 1.3 uL GVP restriction fragments
- 109
- 1.8 uL J23109 restriction fragments
- 4.7 uL MQ
- 106
- 1.3 uL J23106 restriction fragments
- 5.2 uL MQ
- 100
- 1.4 uL J23100 restriction fragments
- 5.1 uL MQ
Procedure:
- DNA and MQ added to an eppendorf cup
- Incubation: 2 min, 44°C (to free sticky ends)
- 0.5 uL T4 Ligase and 1uL T4 Ligase buffer added
- Incubation: 1 hour, room temp. (26°C)
- Denaturation of Ligase 75°C, 10 min
Transporters
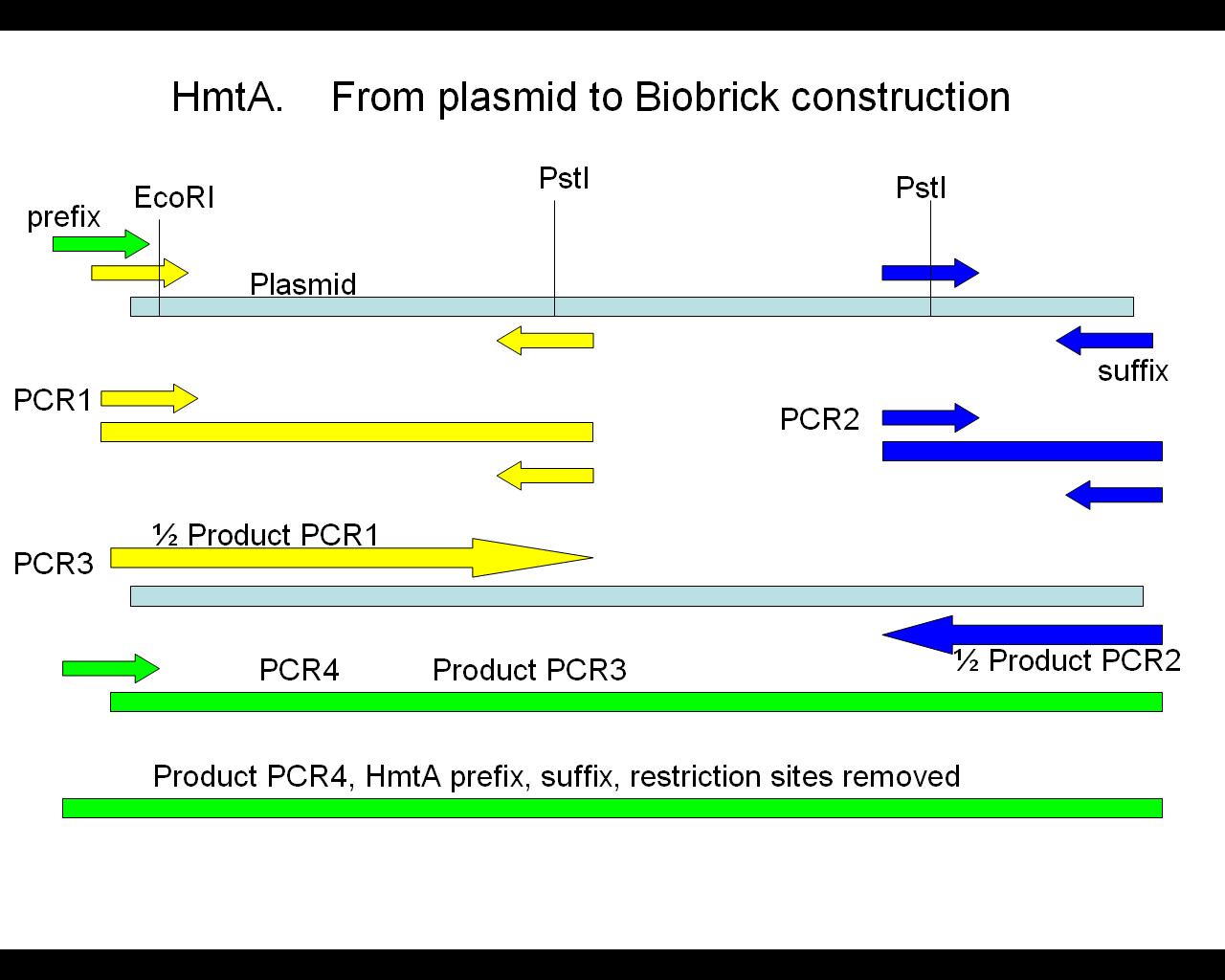
HmtA
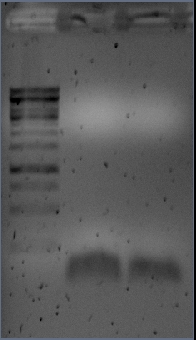
The band at ~1150 kb was cut out of the gel and used for pcr described yesterday gave poor results. No band of expected size and strong primerdimer bands Therefor we will redesign the cloning. The Forward primer will be modified into 2 part. which will require an additional PCR but increases the chance of getting product.
Metal Accumulation
Vectors
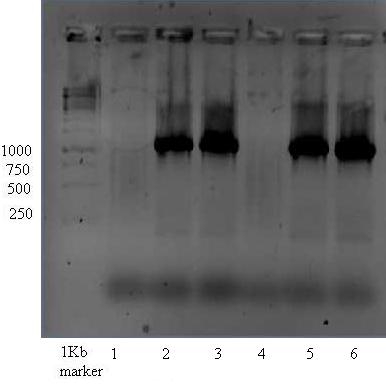
Positive control for the vector with insert done with both new and last years VR VF2 primers resulting in 1107bp bands.
|
|
|
This image shows 4 positive results, just the same as yesterdays but slighty higher bands. This picture indicates that one (VR iGEMGr09) out of 5 primers is not working.
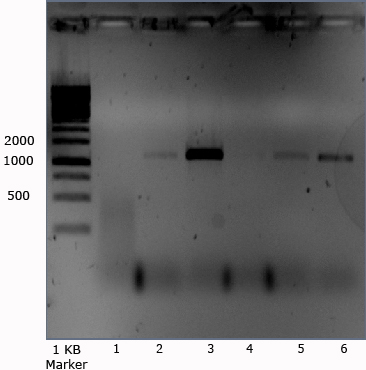
- Check for DNA contimination of PCR mixture contents:
The above noted PCR was made again only this time without DNA, gel shown below.
Since no bands at all are expected, this picture indicates that probably the standard primers of iGEM 2008 are contaminated with DNA (J61002?), or the Mastermix is. The band with most intensity was the following combination: VF2 iGEMGr09 + VR iGEMGr June. Tomorrow a gradient PCR will be done with new mastermix.
- Isolation of pSB1AC3 + promoters and pSB3K3 + promoters to check constructs.
- Isolated the vectors from o/n culture
- Used colonies for o/n culture were:
- AC3-1: colony 3
- AC3-2: colony 10
- AC3-3: colony 11
- K3-1: colony 23
- K3-2: colony 26
- K3-3: colony 37
- Used Sigma plasmid isolation kit, eluted in 50ul MQ
- Used colonies for o/n culture were:
- Isolated the vectors from o/n culture
DNA concentrations were:
| Sample | ng/ul | 260/280 | 260/230 |
| pSB3K3-1 | 6.8 | 1.86 | 2.79 |
| pSB3K3-2 | 7.1 | 1.83 | 3.26 |
| pSB3K3-3 | 46.4 | 1.82 | 2.37 |
| pSB1AC3-1 | 58.0 | 1.81 | 2.27 |
| pSB1AC3-2 | 66.2 | 1.86 | 2.42 |
| pSB1AC3-3 | 76.0 | 1.85 | 2.39 |
--> DONE fill table
- PCR on plasmids to check insert size.
- Used pSB1K3 as negative control
- Compare with J61002 positive control (done also today)
- Used pipeting scheme:
|
|
- Keep in the fridge and check on Agarose gel tomorrow.
- Transform E. coli TOP10 with (pBAD promoter in pSB2K3) and (Lac promoter in pSB1A2)
- Use normal transformation protocol (17 June 2009)
- Also a negative control was taken, MQ.
- Plate the pBAD transformant on LB-Kan and the Lac transformant on TY-Amp.
- O/n @ 37dg
Dry
Jasper finished the GlpF model by fitting a Michaelis-Menten equation to the raw data (using the method of Goudar1999). Mathematica proved to be the best tool for this, as it is one of the few programs that has the Lambert W function built-in (as ProductLog). Furthermore, an interactive model was added to the Wiki demonstrating the results.
There is a specific interaction between the ArsA and ArsB proteins.
The expression of the ArsB gene is required for membrane association of the ArsA protein.
arsB expression is not dependent on gene dosage, so that both arsenical extrusion and resistance are limited by the amount of ArsB protein in the membrane. Tisa1990.
There are two As(III) binding sites on the arsR Summers2009.
|
|
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"