Team:Groningen/Notebook/28 July 2009
From 2009.igem.org
(→Vectors) |
(→Metal Accumulation) |
||
| Line 92: | Line 92: | ||
===Metal Accumulation=== | ===Metal Accumulation=== | ||
| + | *'''Transform ''E.coli'' TOP10 with SmtA''' | ||
| + | **Use normal transformation protocol. | ||
| + | **Transform 2uL pET29a -SmtA and pGEX-3 -SmtA GST to ''E. coli'' TOP10. | ||
| + | **Plate on LBA-Amp and LBA-Kan. | ||
===Vectors=== | ===Vectors=== | ||
Revision as of 10:21, 31 July 2009
Wet
GVP Cluster
Transporters
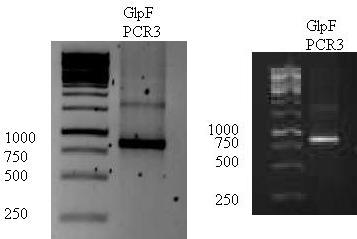
Result of GlpF PCR3 done yesterday.
a band of 903 was seen as expected. The bands were cut out of the gel and gel extraction was done using ..kit. The DNA was used to do another PCR.
| 12,5uL | Phusion |
| 1uL | GlpF FW |
| 1uL | GlpF Rev |
| 0,5 uL | PCR3 DNA |
| 10,5 uL | MQ |
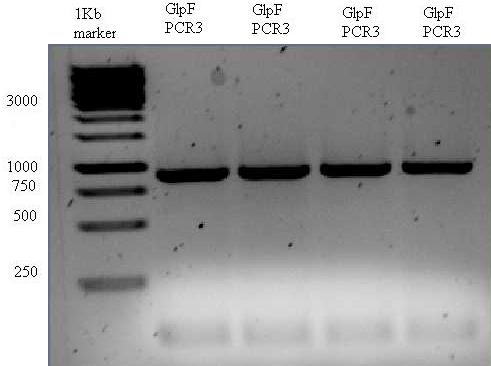
The result can be seen below. The bands were cut out of the gel and purified. The concentration was measured using nanodrop (17ng/uL) and stored at -20.
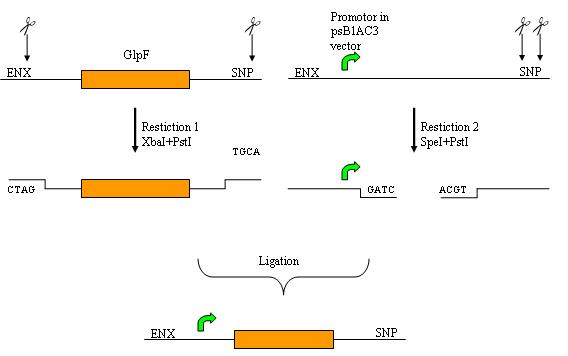
Restriction and ligation was done as can be seen in the image below.
Restiction
|
|
Metal Accumulation
- Transform E.coli TOP10 with SmtA
- Use normal transformation protocol.
- Transform 2uL pET29a -SmtA and pGEX-3 -SmtA GST to E. coli TOP10.
- Plate on LBA-Amp and LBA-Kan.
Vectors
- Ligate metal sensitive promoters with pSB3K3 and pSB1AC3
Oligo's for metal dependent promotors were annealed, 5' phosphorylated using T4 polynucleotide kinase (PNK) and let to cool to room temperature.
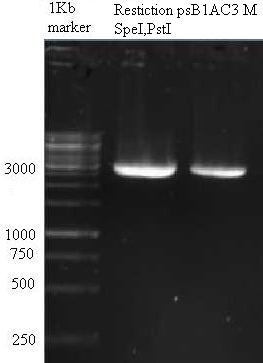
Vectors pSB3K3 & pSB1AC3 were digested using SpeI and EcoRI and put to 1% agarose gel (see images below). Top bands were isolated from gel and purified. The vector (3 μL) and the oligo mix (1 μL) were mixed and heated to 65 °C for 1 minute. After cooling to roomtemperature the ligase buffer (1 μL) and ligase (1 μL) (and 4 μL MilliQ) were added and the ligation was put to 4 °C overnight.
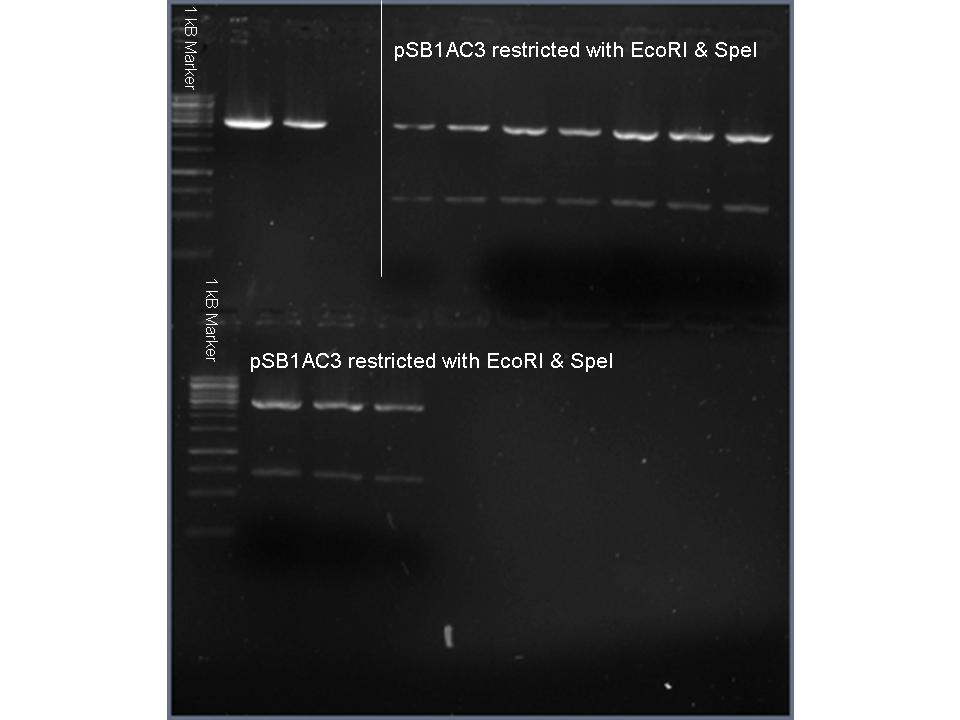
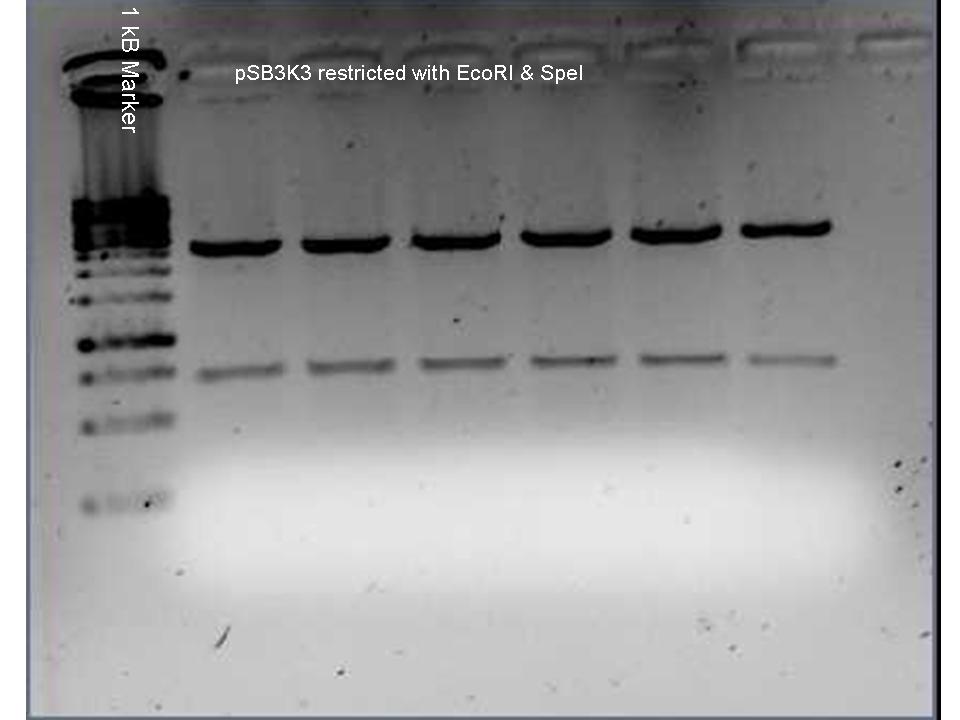
SEE FOR FULL PROTOCOL: Protocol metal dependent promotors
- Restriction digest to ligate vectors+constitutive promoters with the RFP device and pBAD and pLac with pSB3K3/pSB1AC3
- Isolate vector+ pBAD promoter with Sigma Plasmid isolation kit.
- Restriction digest as following:
|
|
|
- Expected sizes of wanted fragments:
- pBAD: 1700bp
- pLac: 200bp
- pSB3K3-const. promoter: 3000
- pSB1AC3-const. promoter: 2700
- RBS-RFP: 1000bp
- Cut bands from gel (pSB1AC3-M showed no bands at all, so nothing to cut out..)
- DNA gel extraction with NucleoSpin Extract II
- Elute with 15uL NE buffer.
- DNA concentration was measured by NanoDrop, they were in the range of 35-5ng/uL.
- Store in the freezer (should be fridge)
- Expected sizes of wanted fragments:
Dry
Jasper started reading about SmtA, but so far no luck in finding any quantitative results that can be used for modelling. At least, it seems that way. The main things that can be found that might be useful are the amount of metal bound to SmtA in Shi1992, a comparison of zinc accumulation with and without smt(A) in Turner1995 (not in E. coli) and PMPS-titration/H+-competition curves in Shi1992 and Blindauer2002. However, it is not yet clear if (and how) this data can be used to derive useful parameters for modelling.
|
|
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"