Team:Groningen/Project/Vesicle
From 2009.igem.org
m (→Planning and requirements:) |
m |
||
| Line 9: | Line 9: | ||
Gas vesicles are hollow proteinous organelles made of gvpA and gvpC (in cyanobacteria) and are permeable to gases and fills by diffusion. It is impermeable to water because of its hydrophobic inside. GvpA is a small 70AA long protein which forms a linear crystalline array of ribs to form the cylindrical shell and conical ends. GvpC is usually on the outside of the gas vesicle to make it stronger and stabilizes the structure (see Figure 1). | Gas vesicles are hollow proteinous organelles made of gvpA and gvpC (in cyanobacteria) and are permeable to gases and fills by diffusion. It is impermeable to water because of its hydrophobic inside. GvpA is a small 70AA long protein which forms a linear crystalline array of ribs to form the cylindrical shell and conical ends. GvpC is usually on the outside of the gas vesicle to make it stronger and stabilizes the structure (see Figure 1). | ||
| - | In ''B. megaterium'' a gas vesicle gene-cluster was found, which contained 14 gvp genes (gas vescicle polycitonic genes) which were functionally expressed in ''E. coli'' by Ning Li and Maura Cannon (Li and Cannon, 1995). The best bouyant fenotype was found when gvpA, gvpP, gvpQ and ORF-1 were excluded (see figure 2). It was suggested that gvpB on the gvp-cluster of B. megaterium is an homolog of gvpA. | + | In ''B. megaterium'' a gas vesicle gene-cluster was found, which contained 14 gvp genes (gas vescicle polycitonic genes) which were functionally expressed in ''<i>E. coli</i>'' by Ning Li and Maura Cannon (Li and Cannon, 1995). The best bouyant fenotype was found when gvpA, gvpP, gvpQ and ORF-1 were excluded (see figure 2). It was suggested that gvpB on the gvp-cluster of B. megaterium is an homolog of gvpA. |
[[Image:Gvp_cluster_b_meg_-_Li_1995.png|900px]] | [[Image:Gvp_cluster_b_meg_-_Li_1995.png|900px]] | ||
| Line 18: | Line 18: | ||
In iGEM 2008 Kyoto had the idea to lift the titanic from the bottom of the sea with the help of bouyant bacteria. | In iGEM 2008 Kyoto had the idea to lift the titanic from the bottom of the sea with the help of bouyant bacteria. | ||
| - | [http://parts.mit.edu/igem07/index.php/Melbourne/Lab_GV_Notebook Melbourne iGEM 2007 team] constructed a bouyant | + | [http://parts.mit.edu/igem07/index.php/Melbourne/Lab_GV_Notebook Melbourne iGEM 2007 team] constructed a bouyant <i>E. coli</i>'' and also they added a <partinfo>BBa_I750016</partinfo> of the short version of the gvp-cluster (without gvpA,P,Q, ORF-1 and AraC). This BioBrick is available and also in the microtiter plate send by HQ to us, so we can use this. Melbourne changed the gvp-cluster by cleaning it from 3 ''Eco''RI sites and 1 ''Pst''I site, this leaded to a accidental addition of a 10x repeat of "TCTGCAAATTA". They mention that they added the BioBrick prefix and suffix to the BioBrick, though the restriction sites of these additions cannot be found by CloneManager in the sequence available on the Partregistry. Hopefully the part is available on a standard vector, which has the pre- and suffix for BioBricks. For cloning this part we can use the [http://parts.mit.edu/igem07/index.php/Melbourne/Lab_Notebook_gv_6 optimized protocol ] (restriction on part with ''Xba''I and ''Spe''I, on the vector with ''Spe''I) for ligation of the gvp-cluster and a {{part|BBa_J61035|vector BBa_J61035}} (3539bp, copy nr??), this unluckily leads to ligation of the gvp-cluster in an unspecific direction. This can of course be tested by restriction, PCR or sequencing, but it takes more time as another step will be introduced. The vector has two selection markers: Ampicillin and Gentamycin. |
==Alternative cloning strategy== | ==Alternative cloning strategy== | ||
| Line 162: | Line 162: | ||
|} | |} | ||
| - | Now we can consider the buoyant density of E. coli with gas vesicles. We have chosen to approach this problem using densities and volume ratios. According to [[Team:Groningen/Literature#Baldwin1995|Baldwin1995]], [[Team:Groningen/Literature#Bylund1991|Bylund1991]] and [[Team:Groningen/Literature#Poole1977|Poole1977]] the density stays within about 3% of 1100 kg/m<sup>3</sup> under wildly varying conditions. This makes our method easier than trying to directly compute the density of a single cell, due to the fact that the volume can differ wildly (both during the life cycle and from strain to strain) and a lack of concrete data on the number of gas vesicles produced (in E. coli). Note that the computations below assume that the gas vesicles simply add to the existing structures. | + | Now we can consider the buoyant density of <i>E. coli</i> with gas vesicles. We have chosen to approach this problem using densities and volume ratios. According to [[Team:Groningen/Literature#Baldwin1995|Baldwin1995]], [[Team:Groningen/Literature#Bylund1991|Bylund1991]] and [[Team:Groningen/Literature#Poole1977|Poole1977]] the density stays within about 3% of 1100 kg/m<sup>3</sup> under wildly varying conditions. This makes our method easier than trying to directly compute the density of a single cell, due to the fact that the volume can differ wildly (both during the life cycle and from strain to strain) and a lack of concrete data on the number of gas vesicles produced (in <i>E. coli</i>). Note that the computations below assume that the gas vesicles simply add to the existing structures. |
{| | {| | ||
| Line 251: | Line 251: | ||
* Used promotor for expression of the gvp-cluster: | * Used promotor for expression of the gvp-cluster: | ||
**Inducible (may be used for proof-of-principle) | **Inducible (may be used for proof-of-principle) | ||
| - | ** | + | **Constitutive (may be used for proof-of-principle) |
**Metal sensitive | **Metal sensitive | ||
*What kind of vector was used by Li and Cannon (1995) or Melbourne (2007)? Is there a negative effect of high copy number? | *What kind of vector was used by Li and Cannon (1995) or Melbourne (2007)? Is there a negative effect of high copy number? | ||
Revision as of 15:32, 7 August 2009
- Transport
- Accumulation
- Metal-sensitive Promoters
- Gas Vesicles
Introduction
About 150 species of prokaryotes (well studied examples are cyanobacteria and halophilic archaea) in aquatic habitats have been shown to contain gas vesicles. These gas vesicles provide cells with bouyancy, there function is either to positioning the bacterium (in water) in order to get the right amount of oxygen or the right amount of light. Gas vesicles are made exclusively of proteins and contain gas. When gas vesicles are present in a cell, the overall density of that cell is lowered and the cell becomes bouyant.
We want to utilize this bouyancy for an application like for example separation of specific molecules or specific cells.
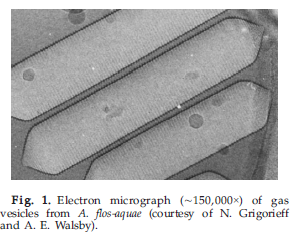
Gas vesicles are hollow proteinous organelles made of gvpA and gvpC (in cyanobacteria) and are permeable to gases and fills by diffusion. It is impermeable to water because of its hydrophobic inside. GvpA is a small 70AA long protein which forms a linear crystalline array of ribs to form the cylindrical shell and conical ends. GvpC is usually on the outside of the gas vesicle to make it stronger and stabilizes the structure (see Figure 1).
In B. megaterium a gas vesicle gene-cluster was found, which contained 14 gvp genes (gas vescicle polycitonic genes) which were functionally expressed in E. coli by Ning Li and Maura Cannon (Li and Cannon, 1995). The best bouyant fenotype was found when gvpA, gvpP, gvpQ and ORF-1 were excluded (see figure 2). It was suggested that gvpB on the gvp-cluster of B. megaterium is an homolog of gvpA.

- Figure 2: gvp cluster B. megaterium. Taken from Li and Cannon, 1995
Gas vesicles in iGEM
In iGEM 2007, Melbourne created a biobrick for gas vesicle formation. In iGEM 2008 Kyoto had the idea to lift the titanic from the bottom of the sea with the help of bouyant bacteria.
Melbourne iGEM 2007 team constructed a bouyant E. coli and also they added a of the short version of the gvp-cluster (without gvpA,P,Q, ORF-1 and AraC). This BioBrick is available and also in the microtiter plate send by HQ to us, so we can use this. Melbourne changed the gvp-cluster by cleaning it from 3 EcoRI sites and 1 PstI site, this leaded to a accidental addition of a 10x repeat of "TCTGCAAATTA". They mention that they added the BioBrick prefix and suffix to the BioBrick, though the restriction sites of these additions cannot be found by CloneManager in the sequence available on the Partregistry. Hopefully the part is available on a standard vector, which has the pre- and suffix for BioBricks. For cloning this part we can use the optimized protocol (restriction on part with XbaI and SpeI, on the vector with SpeI) for ligation of the gvp-cluster and a vector BBa_J61035 (3539bp, copy nr??), this unluckily leads to ligation of the gvp-cluster in an unspecific direction. This can of course be tested by restriction, PCR or sequencing, but it takes more time as another step will be introduced. The vector has two selection markers: Ampicillin and Gentamycin.
Alternative cloning strategy
Another possibility is to use eg. XbaI and SpeI and also cut a vector with these enzymes, this would lead to ligation of the gvp-cluster in one direction. A possible vector for this strategy on the partsregistry is: BBa_J23018 (2298 bp).
Modelling
Buoyancy
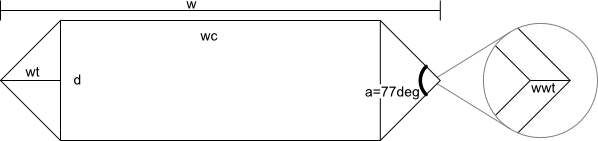
The gas vesicles are shaped roughly like a cylinder with a cone at each end, whose cross-section we model as (based mostly on Walsby1994):
We assume the interior of the wall of the gas vesicle is similarly shaped to the exterior, just slightly smaller (the right-most part of the image above illustrates this situation for the left tip of the gas vesicle).
The different dimensions are related through the equations below. To determine the wall volume, just use them with the given width/diameter (at least for the dimensions given in Walsby1994). To determine the gas volume, use them with w'=w-2*wwt and d'=d-2*tw and then subtract the gas volume (note that the width/diameter of 75nm/50nm comes from Li1998, with the assumption that their "width" should be interpreted as our diameter, as doing it the other way around would leave no room for a cylinder and they specifically mention that the vesicles appear to be shaped like cylinders with conical ends):
| w = total width tw = thickness of wall (1.8-1.95nm) d = diameter a = 77 degrees ρgas = density of gas in vesicle (kg/m^3 = yg/nm^3) ρwall = density of vesicle wall (kg/m^3) wwt = tw/sin(a/2) wt = (1/2)*d/tan(a/2) wc = w - 2*wt Vc = (1/4)*pi*d^2*wc Vt = (1/12)*pi*d^2*wt V = Vc+2*Vt M = ρ*V |
Now we can consider the buoyant density of E. coli with gas vesicles. We have chosen to approach this problem using densities and volume ratios. According to Baldwin1995, Bylund1991 and Poole1977 the density stays within about 3% of 1100 kg/m3 under wildly varying conditions. This makes our method easier than trying to directly compute the density of a single cell, due to the fact that the volume can differ wildly (both during the life cycle and from strain to strain) and a lack of concrete data on the number of gas vesicles produced (in E. coli). Note that the computations below assume that the gas vesicles simply add to the existing structures.
Loading graph...
| Vc = volume of a cell without gas vesicles Vv = volume of gas vesicles in cell Vcv = volume of a cell with gas vesicles (assumed to be Vc+Vv) ρc = density of a cell without gas vesicles ρv = density of gas vesicles ρm = density of medium The following has to be true if the cell floats: Vc*ρc + Vv*ρv < Vcv*ρm (Vcv-Vv)*ρc + Vv*ρv < Vcv*ρm ρc + (Vv/Vcv)*(ρv-ρc) < ρm Assume (ρv - ρc)<0 Vv/Vcv > (ρm - ρc)/(ρv - ρc) Vv/Vcv > 1 - (ρm - ρv)/(ρc - ρv) Explanation of the graph In the graph on the left we assume the cell and its gas vesicles have a density equal to the medium it lives in. Therefore it is able to float. The X-axis depicts the volume fraction of gas vesicles of the cell. The Y-axis depicts the maximum cell density. This is the maximum cell density of the cell WITHOUT gas vesicles. If the cell without gas vesicles has a density higher than the maximum cell density it will sink, because the cell with gas vesicles will have a density higher than the medium it lives in. Therefore if it has a lower density it will float to the surface. |
Missing information
- Used promotor for expression of the gvp-cluster:
- Inducible (may be used for proof-of-principle)
- Constitutive (may be used for proof-of-principle)
- Metal sensitive
- What kind of vector was used by Li and Cannon (1995) or Melbourne (2007)? Is there a negative effect of high copy number?
- What is the maximum amount of pressure gas vesicles can handle? At which depth would this be, how can one put this kind of pressure on a water column?
- What is the density of gas vesicles in cells (normally or in case of over-expression)
- Modelling parameters (to be measured):
- What is the density of the cell without gas vesicles/metal (largely known, but would be good to check), and how is this affected by letting the cell make gas vesicles and/or metal transporters/accumulators and so on?
- How many gas vesicles are produced? (As volume percentage?)
- (How fast are they produced?)
-Cyanobacteria (Bowen and Jensen, 1965): gas vacuoles made up of gas vesicles (75 nm in diameter and up to 1.0 ,um in length, single wall layer only 2 nm thick) 0,7MPa gives irreversible loss of buoyancy fenotype, but found in next generation.
you can use a nephelometer, to measurement the light scattering of the gas vesicles (walsby1994)
Planning and requirements:
- Modelling:
- Buoyancy
- Permeability
- Number of gasvesicles
- Where do the gasvesciles end, in hight.
- Mass of the bacteria (e.coli)
- How long it takes before it floates
- How long it take untill it is being expressed
- How long it will take untill there are enough gasvesicles
- How does it stay floating
- Buoyancy
- Lab:
- Gvp
- Cluster of biobricks
- Vector ordered from article
- First expression with constutatieve promotor, later with metal sensitive promotor
- Measurements
- Where are the bacteria, what are bac concentrations on a certain hight (not necessary???)
- Making pictures on certain hight and compare with picture of known concentration
- Do we have gas vesicles?
- Measurement vesicle volume using a nephelometer Holland2009 and Walsby1994
- Measurement of cell density by centrifuging in density (Percoll) gradients Walsby1994
- Measurement the volume of gass using a compression tube Walsby1979, Walsby1994
- Where are the bacteria, what are bac concentrations on a certain hight (not necessary???)
- Gvp
Literature
Astrid C. Sivertsen et al. 2009 Solid-State NMR Evidence for Inequivalent GvpA Subunits in Gas Vesicles. Journal of Molecular Biology 387, p1032–1039.
 "
"