Team:IIT Madras/Circuit
From 2009.igem.org
One of the plasmids will have the gene of interest which will be expressed only when the plasmids apart from this particular one are lost in a particular order. This particular order or “code” will consist of a sequence of antibiotic treatments given to the transformed cells. The correct code triggers the loss of plasmids in the cells in a particular order. Each plasmid will be linked to the plasmids which are supposed to be lost before and after it in a highly regulated fashion. The plasmid loss can be regulated very tightly using a “suicide gene” (gene coding for the bacterial gyrase poison). These genes will be triggered when the culture is subjected to the wrong antibiotic (Out of sequence). To ensure that the suicide genes don’t fire randomly, they are under the control of repressors which are on the plasmids that are supposed to be lost after it and are expressed constitutively. Thus, they fire only when the plasmid containing the repressor is lost. When all the plasmids other than the plasmid containing the gene of interest are lost, the repressors that have been blocking the required gene are lost, thus allowing its expression, which can manifest as a phenotype or function in the cell. This will be the “unlocking” of the lock.
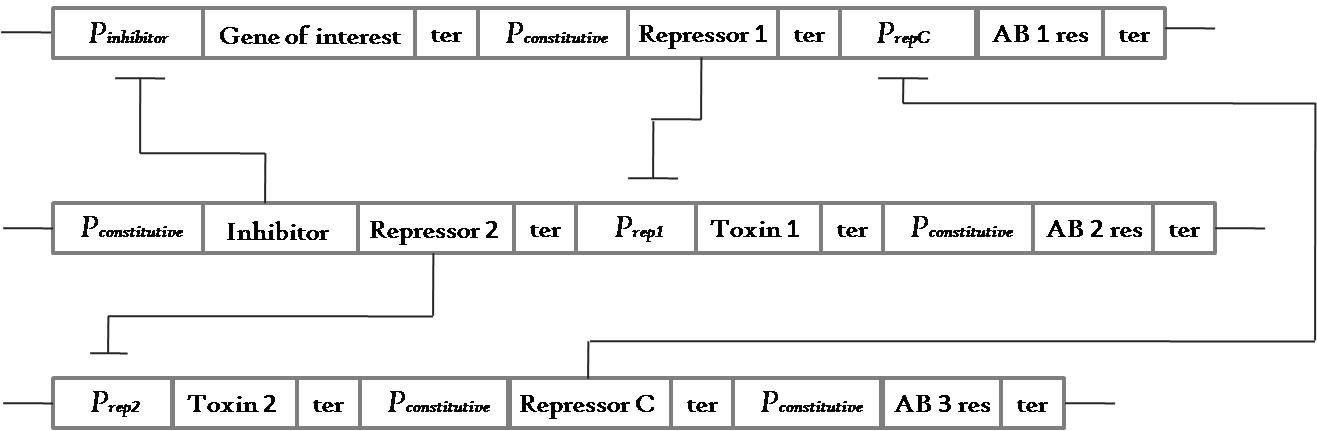
----The grand idea of a 3 plasmid locking system revolved around constructing something like the below:
Fig 3.1: This is a general circuit for a 3 plasmid system which can show locking property with a code length of 2 units - Ab2->Ab1. This means that the cells need to be grown first in antibiotic medium 2 and then transferred to a medium containing antibiotic 1. It is easy to see that to maintain the whole system of 3 plasmids, the cells need to be grown in medium containing antibiotic 3.
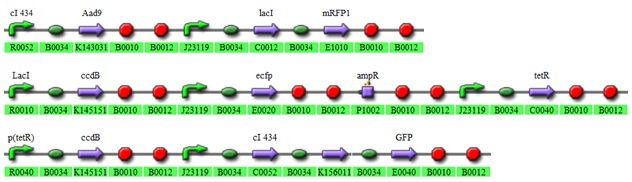
To which we thought we could fit in the following parts (the gene of interest is not included. It could be anything):
Fig 3.2:The 3 plasmid system.
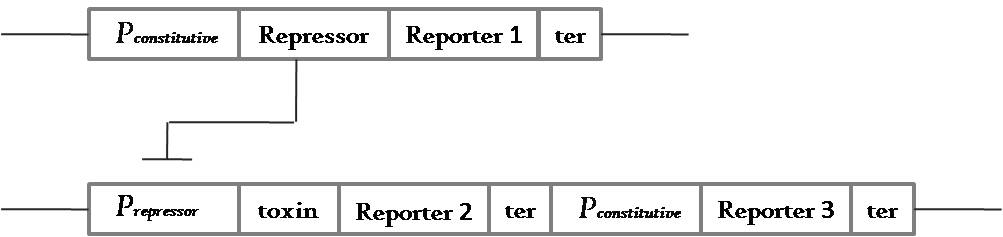
Then we scaled it down and began to build constructs for a 2 plasmid system which works like this:
Fig 3.3:This is a general circuit for a 2 plasmid system that can show locking property with a code length of 1 unit - which means the system is unlocked when it is grown in a medium containing antibiotic 1.
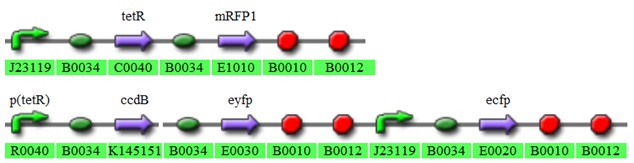
To which we fit in parts from the registry to make it look like this:
Fig 3.4:The 2 plasmid system.
For the proof of concept experiments, we built the following constructs:
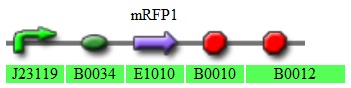
[http://partsregistry.org/Part:BBa_K272001 K272001-constitutive RFP expressor]
Fig 3.5:This plasmid confers constitutive expression of RFP in the transformed cells.
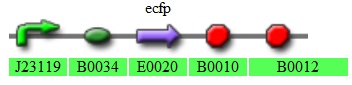
[http://partsregistry.org/Part:BBa_K272002 K272002-constitutive CFP expressor]
Fig 3.5:This plasmid confers constitutive expression of CFP in the transformed cells.
 "
"