Team:McGill
From 2009.igem.org
(Prototype team page) |
(→Project Description (General)) |
||
| (24 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| - | |||
| - | |||
<html> | <html> | ||
| - | < | + | <body> |
| - | < | + | <center> |
| - | + | <br/> | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | </ | + | |
| - | < | + | <img src="https://static.igem.org/mediawiki/2009/d/d5/Mcgill09BannerMod.PNG" width="950" height="250" usemap="#bannermap" id="navbar"> |
| + | <map id="bannermap" name="bannermap"> | ||
| + | |||
| + | <area shape="rect" | ||
| + | coords="0,0,720,200" | ||
| + | alt="Home" | ||
| + | href="https://2009.igem.org/Team:McGill"> | ||
| + | |||
| + | <area shape="rect" | ||
| + | coords="720,80,900,120" | ||
| + | alt="Home" | ||
| + | href="http://www.mcgill.ca/"> | ||
| + | |||
| + | <area shape="rect" | ||
| + | coords="0,200,170,250" | ||
| + | alt="Team" | ||
| + | href="https://2009.igem.org/Team:McGill"> | ||
| + | |||
| + | <area shape="rect" | ||
| + | coords="171,200,317,250" | ||
| + | alt="Team" | ||
| + | href="https://2009.igem.org/Team:McGill/Team"> | ||
| + | |||
| + | <area shape="rect" | ||
| + | coords="318,200,468,250" | ||
| + | alt="Project" | ||
| + | href="https://2009.igem.org/Team:McGill/Project"> | ||
| + | |||
| + | <area shape="rect" | ||
| + | coords="469,200,658,250" | ||
| + | alt="Notebook" | ||
| + | href="https://2009.igem.org/Team:McGill/Notebook"> | ||
| + | |||
| + | <area shape="rect" | ||
| + | coords="659,200,807,250" | ||
| + | alt="Results" | ||
| + | href="https://2009.igem.org/Team:McGill/Results"> | ||
| + | |||
| + | <area shape="rect" | ||
| + | coords="808,0,950,250" | ||
| + | alt="Sponsors" | ||
| + | href="https://2009.igem.org/Team:McGill/Sponsors"> | ||
| + | |||
| + | </map> | ||
| + | </center> | ||
| + | |||
| + | </body> | ||
| + | </html> | ||
| + | =='''Welcome to the 2009 McGill iGEM Team homepage!'''== | ||
| - | + | Welcome to Quebec's only iGEM team! Our team is made up of a group of motivated students and faculty from McGill University in Montreal, Quebec, Canada. We have been hard at work preparing for this year's competition. The following webpages should give you a brief oversight into what we have accomplished this year! | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | =='''Project Description (General)'''== | |
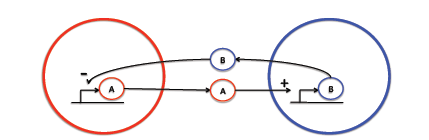
| + | We are interested in exploring the biology of intercellular signaling. The human body is composed of trillions of cells, which have to communicate with each other over long distances. We are interested in investigating the effect of distance on the signaling dynamics. Although there are many cellular communication mechanisms we decided to focus on chemical signaling. Specifically, we are interested in investigating the dynamics of activation-inhibition signaling. This occurs when a cell synthesis and releases a chemical that is capable of diffusing to a second cell and activating production of an inhibitory chemical which diffuses back to the first cell and inactivates synthesis of the former chemical (as illustrated in the following diagram). | ||
| + | [[Image: Mcgill09Projectfig1.png|frame|center|Figure 1 – The diffusible proteins A and B are involved in an opposite feedback mechanism]] | ||
| + | We have explored this system using two approaches: mathematical and microbiological. We began by developing a mathematical model allowing us to perform ''in silico'' experiments of the system. Furthermore, we engineered two strains of bacteria to recreate this system allowing us to begin validating our mathematical model. | ||
| - | + | Aside from simply gaining insight into this type of signaling we also hoped our investigations would lead to a novel type of biological sensor! Explore the following pages for a more detailed explanation of our project! | |
| - | ! | + | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
Latest revision as of 06:22, 21 October 2009

Welcome to the 2009 McGill iGEM Team homepage!
Welcome to Quebec's only iGEM team! Our team is made up of a group of motivated students and faculty from McGill University in Montreal, Quebec, Canada. We have been hard at work preparing for this year's competition. The following webpages should give you a brief oversight into what we have accomplished this year!
Project Description (General)
We are interested in exploring the biology of intercellular signaling. The human body is composed of trillions of cells, which have to communicate with each other over long distances. We are interested in investigating the effect of distance on the signaling dynamics. Although there are many cellular communication mechanisms we decided to focus on chemical signaling. Specifically, we are interested in investigating the dynamics of activation-inhibition signaling. This occurs when a cell synthesis and releases a chemical that is capable of diffusing to a second cell and activating production of an inhibitory chemical which diffuses back to the first cell and inactivates synthesis of the former chemical (as illustrated in the following diagram).
We have explored this system using two approaches: mathematical and microbiological. We began by developing a mathematical model allowing us to perform in silico experiments of the system. Furthermore, we engineered two strains of bacteria to recreate this system allowing us to begin validating our mathematical model.
Aside from simply gaining insight into this type of signaling we also hoped our investigations would lead to a novel type of biological sensor! Explore the following pages for a more detailed explanation of our project!
 "
"