Team:McGill
From 2009.igem.org
| Line 11: | Line 11: | ||
coords="0,0,720,200" | coords="0,0,720,200" | ||
alt="Home" | alt="Home" | ||
| - | href="https://2009.igem.org/Team:McGill# | + | href="https://2009.igem.org/Team:McGill#Who We are"> |
<area shape="rect" | <area shape="rect" | ||
Revision as of 15:42, 12 August 2009

Project Overview
Activation-Inhibition Coupling
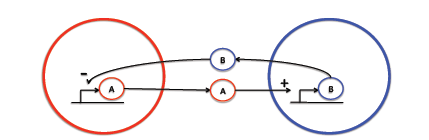
- The biological system to be investigated consists of two spatially separated populations of cells, each expressing a different protein; here named A and B. Protein A diffuses out of the first population of cells and into the second where it stimulates the production of protein B, which is another diffusible protein that can travel back to the first population and inhibit the production of protein A.
Dynamics
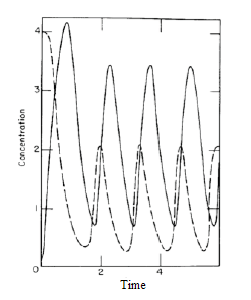
This type of cellular signaling is known as activation-inhibition coupling, and although it can seem fairly mundane,
some interesting dynamics can result. Figure 2 illustrates the mathematical modeling of this system where periodic oscillations were observed. The type ofbehavior observed depends on the parameters of the system, including the rate
of diffusion of the proteins, the distance between the two cell populations, the sensitivity to transcriptional regulation and several others.
Hypothesis, Goals, and Potential Applications
To date, the kinetics of this type of cellular signaling has only been theoretically analyzed using mathematical tools. We aim to provide experimental observations of the system at work. Furthermore, we plan on extending the mathematical framework from a 1 dimensional model to 2 dimensions. In the 2D setup, the two populations of cells will be cultured together with random spatial distribution at varying densities. We expect to see several forms of dynamics depending on the density: at low densities, only local interactions will become significant and pockets of oscillations will be observed; at high densities, a steady state of activation will be achieved; however at intermediate densities, we expect to observe waves of activation propagating through the cell culture. The mathematical model will drive the design of experiments to verify these claims. These investigations could yield the foundations for a novel type of biological sensor. Exposing the cultured cells to a particular compound could alter the qualitative dynamics of the cellular signal: switching from steady state activation to propagating waves, thus alerting the investigator to the presence of the compound.
References
1. Shymko, R., and Glass, L. (1974). Spatial switching in chemical reactions with heterogeneous catalysis. The Journal of Chemical Physics 60, 835.
 "
"