Team:Minnesota/Modeling
From 2009.igem.org
| Home | The Team | The Project | Submitted Parts | Modeling | SynBioSS Designer | Parts Characterization | Experiments and Calendar |
|---|
Contents |
Our Modeling Goals
Create mathematical models of our experimental results. Our computational output will guide future research and is the key to determining what is happening at the molecular level of our logical AND gate.
Mathematical Modeling
Overview
In order to create a model, we must first use a program that will allow us to enter in the contents of the reactions network. The reaction network is formatted as .nc file, which is a special format called NetCDF that works particularly well for entering in the required data. The most basic way to enter in a network is in Linux, as a text file and then converts it to a .nc file. However, this is very tedious and error-prone process. In order to more quickly and accurately create the models, we use one of two programs that have been created for this purpose: SynBioSS or a MATLAB GUI. Each of these programs allows us to graphically see what the model looks like. (See screenshots).<break> Once the model is created, it is sent to a supercomputer where it is run using hybrid stochastic simulation algorithms created for this type of models. This algorithm takes the initial conditions of the model along with the reactions and kinetic constants and determines the state of the system over time. It does this by using the rate constants along with a random number generator to determine the order that reactions happen in the system. After one reaction happens, it changes the state of the system accordingly, and then repeats the algorithm to find the next reaction. This allows the state of the system (the number of molecules of all species) to change over time, just like real cells do. The results of the models are then removed from the supercomputer and analyzed using MATLAB. It is then possible to see if the model ran correctly and if it gave reasonable results. Depending on how the model compares to the experimental data, kinetic constants are changed and then the process of running a model begins again. After many iterations of this process, the model is as accurate as is possible. The accuracy of the model is determined by the difference in reporter gene expression between the model and the experiments. Calculating this value requires several steps to ensure accuracy. First, the models must be rescaled, so that the model data matches with the experimental numerically. In order to do this, the model is compared to another model, called a baseline expression model, which produces the maximum amount of reporter gene the system can produce. This is done by getting rid of any repressors that could slow down this production. Experimentally, a similar procedure is done which results in the baseline reporter gene expression for the actual cells. Using this data, it is possible to rescale the model and the experimental data to the same standard. After this, the error is found by taking the difference between the experiment and model at all points and squaring it. The smaller this number is, the better the model is. Once the best model is found, it can be compared to other models and analysis can be done to determine if there is anything to learn about the AND gate.
It is important to be able to observe the fidelity of the AND gate in silico as well as in vivo. Using our in-house software suite, SynBioSS, we used networks of sixty or more reactions to model the AND gate with different constructs and mutations. Similar to the experimental procedures, we quantified the fidelity of our AND gate with GFP. We also used a MATLAB Graphical User Interface (GUI) to analyze our experimental data. We used our empirical data from the wet lab to perfect our model.
Some Basic Modeling
We are working with four different combinations of lac andtet (L and T, respectively) operators in our promoter design project. These constructs are: T--, TT-, TTL and -T-. Here, we will demonstrate a little basic modeling we have done of the TTL construct using SynBioSS MATLAB and a design GUI.
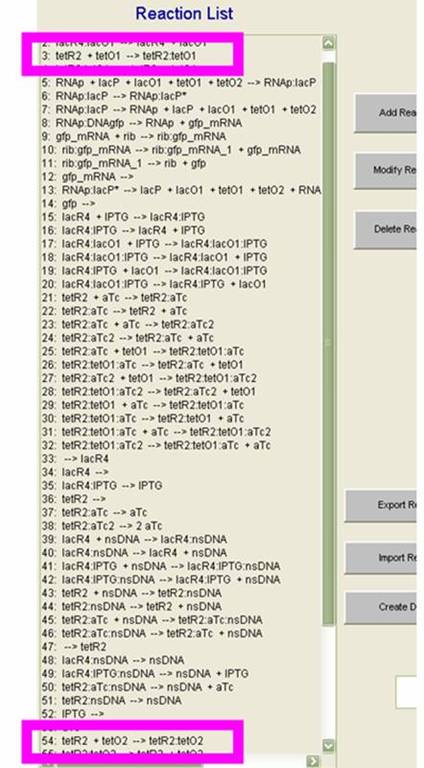
On the SynBioSS wiki, the TTL model is available for download in a .nc format. Upon opening this with the MATLAB GUI, the list of 65 reactions that constitute the repression and induction of transcription. In this model, there are two tet operators TO1 and TO2 and one lac operator LO1. The reactions included in this list account for the repression and induction of transcription in the absence and presence of aTc and IPTG, respectively. In this simple model, we examined reactions 3 and 54, which involve the tet operators being bound by the repressor. (These reactions are shown in the GUI on the left.) We varied the reactions constants from the original 108 as low as 0 and as high as 109 to see if the fidelity of the AND gate could depend on the reaction rates for repression. We quantified our results by counting molecules of GFP. This not only gave us a clearer idea of what was happening at the molecular level at this promoter site, but also served as a preparation for matching the experimental results computationally. "
"