Team:Minnesota/Project
From 2009.igem.org
| (13 intermediate revisions not shown) | |||
| Line 3: | Line 3: | ||
|} | |} | ||
| + | {| style="color:gold;background-color:#800000;" cellpadding="3" cellspacing="1" border="1" bordercolor="#fff" width="90%" align="center" | ||
{| style="color:gold;background-color:#800000;" cellpadding="3" cellspacing="1" border="1" bordercolor="#fff" width="90%" align="center" | {| style="color:gold;background-color:#800000;" cellpadding="3" cellspacing="1" border="1" bordercolor="#fff" width="90%" align="center" | ||
!align="center"|[[Team:Minnesota|<font color="gold">Home</font>]] | !align="center"|[[Team:Minnesota|<font color="gold">Home</font>]] | ||
!align="center"|[[Team:Minnesota/Team|<font color="gold">The Team</font>]] | !align="center"|[[Team:Minnesota/Team|<font color="gold">The Team</font>]] | ||
!align="center"|[[Team:Minnesota/Project|<font color="gold">The Project</font>]] | !align="center"|[[Team:Minnesota/Project|<font color="gold">The Project</font>]] | ||
| - | |||
| - | |||
!align="center"|[[Team:Minnesota/Designer|<font color="gold">SynBioSS Designer</font>]] | !align="center"|[[Team:Minnesota/Designer|<font color="gold">SynBioSS Designer</font>]] | ||
| - | !align="center"|[[Team:Minnesota/ | + | !align="center"|[[Team:Minnesota/Modeling|<font color="gold">Modeling</font>]] |
| - | !align="center"|[[Team:Minnesota/Notebook|<font color="gold"> | + | !align="center"|[[Team:Minnesota/Notebook|<font color="gold">Experimental</font>]] |
| - | + | !align="center"|[[Team:Minnesota/Parts Characterization|<font color="gold">Competition Requirements</font>]] | |
|} | |} | ||
<h1>The Project</h1> | <h1>The Project</h1> | ||
| + | Although, we will only be judged for the software tools we have developed, we have also experimentally constructed, built and tested BioBricks that function as logical AND gates. The most important aspect of our work is that we combined simulations and experiments. | ||
| + | |||
| + | <h2> SynBioSS Designer (https://2009.igem.org/Team:Minnesota/Designer) </h2> | ||
<h2>AND Gates</h2> | <h2>AND Gates</h2> | ||
| - | <h3> How | + | <h3> How they work</h3> |
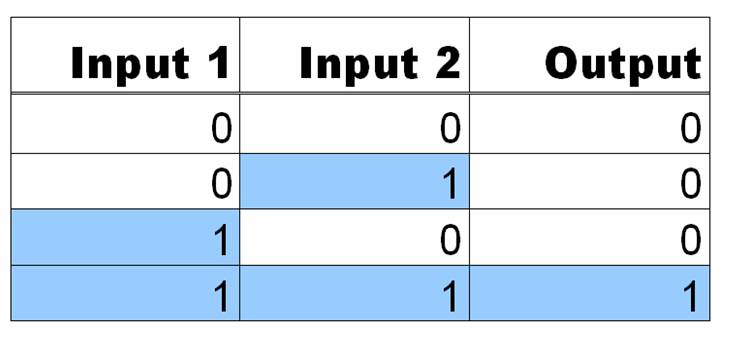
[[Image:ANDgatetable.jpg|300px|right]]AND Gates are logic circuits that produce an output if and only if two inputs are present. For example, our system produces GFP only if two small molecules named aTc and IPTG are present. If one of them is not present, no GFP will be produced. Ideally an AND gate will obey the truth table shown to the right, where 1 means the input is present, and 0 means that it is not. | [[Image:ANDgatetable.jpg|300px|right]]AND Gates are logic circuits that produce an output if and only if two inputs are present. For example, our system produces GFP only if two small molecules named aTc and IPTG are present. If one of them is not present, no GFP will be produced. Ideally an AND gate will obey the truth table shown to the right, where 1 means the input is present, and 0 means that it is not. | ||
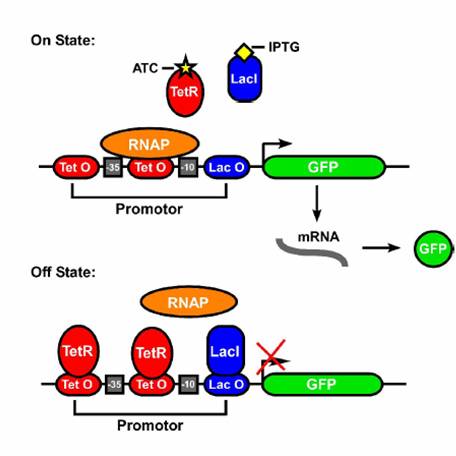
| - | In order to get this behavior, we take advantage of the structure of promoter sequences in prokaryotes. When RNA polymerase(more specifically the sigma factor) binds to the promoter, it binds to region at -35 and -10 relative to the transcriptional start site. Since the entire sequence is not used by the sigma factor, there are three regions of DNA that can be changed to provide AND gate behavior. These are the upstream of -35, between -35 and -10, and downstream of -10. In each of these sites, we can place a DNA sequence that is preferentially bound by proteins in the cell. For this case, the two proteins are TetR and LacI, both of which are constitutively produced by our strain of E. coli(DH5-alpha-pro). In each of the three sites, either a | + | In order to get this behavior, we take advantage of the structure of promoter sequences in prokaryotes. When RNA polymerase (more specifically, the sigma factor) binds to the promoter, it binds to region at -35 and -10 relative to the transcriptional start site. Since the entire sequence is not used by the sigma factor, there are three regions of DNA that can be changed to provide AND gate behavior. These are the upstream of -35, between -35 and -10, and downstream of -10. In each of these sites, we can place a DNA sequence that is preferentially bound by proteins in the cell. For this case, the two proteins are TetR and LacI, both of which are constitutively produced by our strain of E. coli(DH5-alpha-pro). In each of the three sites, either a lacO or tetO2 operator can replace the normal DNA. In order to create an AND gate, at least one of these sites must be a tetO2 operator and one must be a lacO operator. |
[[Image:ANDgate.jpg|400px|left]] | [[Image:ANDgate.jpg|400px|left]] | ||
| - | We studied the potential of a synthetic, single promoter AND gate. This helps us understand at a fundamental level what is happening in a network with multiple regulators and opens further research paths. The device consists of parts of the Tet (tetracycline) and Lac (lactose) and responds to commonly used inducers IPTG (Isopropyl β-D-1-thiogalactopyranoside) and aTc. Three or fewer of these operators can be combined to form a promoter and the order and quantity of each operator site result in different constructs, that is, given three blank spots for operators on a promoter and the choice of either -, T or L for each spot, there are a variety of possible promoter designs. We chose to focus on | + | We studied the potential of a synthetic, single promoter AND gate. This helps us understand at a fundamental level what is happening in a network with multiple regulators and opens further research paths. The device consists of parts of the Tet (tetracycline) and Lac (lactose) operons and responds to commonly used inducers IPTG (Isopropyl β-D-1-thiogalactopyranoside) and aTc (anhydortetracycline). Three or fewer of these operators can be combined to form a promoter and the order and quantity of each operator site result in different constructs, that is, given three blank spots for operators on a promoter and the choice of either -, T or L for each spot, there are a variety of possible promoter designs. "-" represents a region of non-coding DNA from λ phage functions as a "null" operator site; no proteins in our E. coli strain should specifically bind it. "T" represents the tetO2 operator site, and "L" represents the lacO operator site. A number of these constructs were built in the Kaznessis lab using wild-type operator sites prior to this summer's iGEM team project. To tune and more thoroughly understand the function of these constructs, we replaced the wild-type tetO2 operator sites with several mutants which we found in the literature. We chose to focus on three constructs: |
<ul> | <ul> | ||
<li>T T L</li> | <li>T T L</li> | ||
<li>T T -</li> | <li>T T -</li> | ||
<li>T - -</li> | <li>T - -</li> | ||
| - | |||
</ul> | </ul> | ||
| - | |||
| - | <h2> | + | |
| - | + | ||
| - | + | ||
| - | + | ||
| + | <h2> tetO2 mutants </h2> | ||
| + | The tetO2 operator site is a 19bp palindromic DNA sequence which is tightly bound by the dimer TetR, repressing the expression of downstream genes in the absence of an inducer. Helbl and Hillen have documented several mutations of tetO2 and the binding affinities of wild-type and mutant TetR for them. Since TetR is a dimer which binds both sides of the palindrome in the same way, each of these mutations were also made at the same location on both sides of the palindrome. | ||
| + | [[Image:TetO2.tiff|frame]] | ||
Latest revision as of 18:33, 20 October 2009
| Home | The Team | The Project | SynBioSS Designer | Modeling | Experimental | Competition Requirements |
|---|
Contents |
The Project
Although, we will only be judged for the software tools we have developed, we have also experimentally constructed, built and tested BioBricks that function as logical AND gates. The most important aspect of our work is that we combined simulations and experiments.
SynBioSS Designer (https://2009.igem.org/Team:Minnesota/Designer)
AND Gates
How they work
AND Gates are logic circuits that produce an output if and only if two inputs are present. For example, our system produces GFP only if two small molecules named aTc and IPTG are present. If one of them is not present, no GFP will be produced. Ideally an AND gate will obey the truth table shown to the right, where 1 means the input is present, and 0 means that it is not.In order to get this behavior, we take advantage of the structure of promoter sequences in prokaryotes. When RNA polymerase (more specifically, the sigma factor) binds to the promoter, it binds to region at -35 and -10 relative to the transcriptional start site. Since the entire sequence is not used by the sigma factor, there are three regions of DNA that can be changed to provide AND gate behavior. These are the upstream of -35, between -35 and -10, and downstream of -10. In each of these sites, we can place a DNA sequence that is preferentially bound by proteins in the cell. For this case, the two proteins are TetR and LacI, both of which are constitutively produced by our strain of E. coli(DH5-alpha-pro). In each of the three sites, either a lacO or tetO2 operator can replace the normal DNA. In order to create an AND gate, at least one of these sites must be a tetO2 operator and one must be a lacO operator.
We studied the potential of a synthetic, single promoter AND gate. This helps us understand at a fundamental level what is happening in a network with multiple regulators and opens further research paths. The device consists of parts of the Tet (tetracycline) and Lac (lactose) operons and responds to commonly used inducers IPTG (Isopropyl β-D-1-thiogalactopyranoside) and aTc (anhydortetracycline). Three or fewer of these operators can be combined to form a promoter and the order and quantity of each operator site result in different constructs, that is, given three blank spots for operators on a promoter and the choice of either -, T or L for each spot, there are a variety of possible promoter designs. "-" represents a region of non-coding DNA from λ phage functions as a "null" operator site; no proteins in our E. coli strain should specifically bind it. "T" represents the tetO2 operator site, and "L" represents the lacO operator site. A number of these constructs were built in the Kaznessis lab using wild-type operator sites prior to this summer's iGEM team project. To tune and more thoroughly understand the function of these constructs, we replaced the wild-type tetO2 operator sites with several mutants which we found in the literature. We chose to focus on three constructs:
- T T L
- T T -
- T - -
tetO2 mutants
The tetO2 operator site is a 19bp palindromic DNA sequence which is tightly bound by the dimer TetR, repressing the expression of downstream genes in the absence of an inducer. Helbl and Hillen have documented several mutations of tetO2 and the binding affinities of wild-type and mutant TetR for them. Since TetR is a dimer which binds both sides of the palindrome in the same way, each of these mutations were also made at the same location on both sides of the palindrome. File:TetO2.tiff
 "
"