Team:Minnesota/Project
From 2009.igem.org
| Home | The Team | The Project | Parts Submitted to the Registry | Modeling | Parts Characterization | Notebook |
|---|
Contents |
The Project
AND Gates
How it works
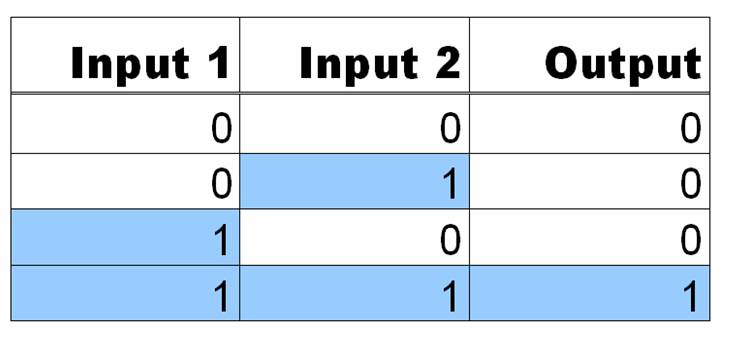
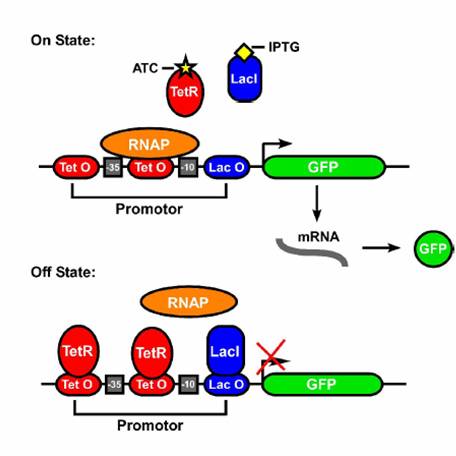
AND Gates are logic circuits that produce an output if and only if two inputs are present. For example, our system produces GFP only if two small molecules named aTc and IPTG are present. If one of them is not present, no GFP will be produced. Ideally an AND gate will obey the truth table shown to the right, where 1 means the input is present, and 0 means that it is not.We studied the potential of a synthetic, single promoter AND gate. This helps us understand at a fundamental level what is happening in a network with multiple regulators and opens further research paths. The device consists of parts of the Tet (tetracycline) and Lac (lactose) and responds to commonly used inducers IPTG (Isopropyl β-D-1-thiogalactopyranoside) and aTc. Three or fewer of these operators can be combined to form a promoter and the order and quantity of each operator site result in different constructs, that is, given three blank spots for operators on a promoter and the choice of either -, T or L for each spot, there are a variety of possible promoter designs. We chose to focus on four constructs:
- T T L
- T T -
- T - -
- - T -
We also examined 4 mutations in the Tet operator site for each construct.
The Experiments
Results
Protocols: Standard techniques that we used in the wet lab
Bacterial Culture
Sterile Technique
- Always work around a flame or in the hood
- Flame the mouth and cap of any bottle, flask or tube upon uncapping and recapping
- Sterilize metal instruments between uses by dipping in 100% ethanol and flaming
Bacterial Culture Maintenance
Culture cells:
- At 36 degrees Celsius
- Shaking at 220 rpm
- At 10% total flask/tube volume
- In mid-log phase(0.1 < OD600 <= 0.4) (with OD600 = 1 ->8.8x108cell/ml)
Bacterial Culture For Gene Expression Experiments
- Pick and individual colony from a plate and inoculate 2ml LB + amp media
- Incubate overnight at 37 C, shaking at 220 rpm
- Inoculate fresh media with overnight culture such that new culture has 2.5% inoculum; this is the secondary culture
- Incubate at 37 C shaking at 220 rpm until OD600 = 0.4 (~2 hrs)
- Inoculate 4 ml LB + amp + inducer (aTc or IPTG) with 100ul secondary culture
- Continue cultures as described above in "bacterial culture maintenance" for 9 hrs
- Isolate cell samples from cultures at 3, 6, and 9 hour time points
- Remove 100ul sample aliquots from cultures
- Pellet samples at 5K rpm for 5 minutes
- Remove supernatant
- Wash cells with 1 ml chilled 1xPBS, pH 7.6
- Resuspend cells by vortexing
- Re-pellet cells at 5K rpm for 5 minutes
- Remove supernatant
- Fix cells; resuspend cells in 1 ml 4% PFA (in PBS)
- Incubate at RT for 30 minutes
- Pellet cells at 5K rpm for 5 minutes
- Remove supernatant
- Resuspend cells in 1 ml 1xPBS
- Store samples at 4 C until analysis by flow cytometry
Transformation of Chemically Competent Cells
- Thaw cells and incubate transformant DNA in ice(~15 minutes)
- Combine 50 ul cells with ~3uL DNA and mix gently
- Incubate samples on ice for 15 minutes
- Heat shock cells in 42C water bath of 50 seconds
- Incubate samples on ice for 5 minutes
- Recover cells in 0.5 ml SOC media, shaking at 37C for 1 hour at 220 rpm
- Transfer cells to a 2 ml microfuge tube
- Spin cells down at 6K rpm for 2 minutes
- Remove all but ~100uL supernatant media
- Resuspend cells gently in remaining media
- Plate cells on LB + ab plates
- Incubate plates overnight at 37C
DNA Work
Plasmid Prep from cultures (using QIAprep Spin Miniprep Kit)
- Pick and individual colony from a plate and inoculate 2ml LB + ab media
- Incubate culture overnight at 37C
- Transfer culture to 2 ml microfuge tube
- Spin cells down at 13K rpm for 2 min at RT and remove supernatant
- Resuspend cells in 250 ul Buffer P1 (stored at 4 C)
- Add 250 ul Buffer P2 and mix thoroughly by inverting-- the solution should turn blue
- Add 350 ul Buffer N3 and mix immediately and thoroughly by inverting-- the solution should turn colorless
- Centrifuge sample at 13K rpm for 10 minutes
- Transfer supernatant to a fresh QIAprep spin column, leaving cell debris pellet behind
- Centrifugre supernatant into column at 13K rpm for 1 minute
- Remove the flowthrough
- Wash column with 0.5 ml Buffer PB; apply to column and spin through at 13K rpm for 1 minute
- Remove the flowthrough
- Wash column with 0.75 ml Buffer PE; apply to column and spin through at 13K rpm for 1 minute
- Remove the flowthrough
- Spin out residual liquid at 13K rpm for 1 minute
- Place column in a fresh 1.5 ml microfuge tube
- Elute DNA; apply 40 ul Buffer EB to column, incubate at room temperature for 2 minutes and spin out of column at 13K rpm for one minute
DNA quantification
- Diluite DNA as appropriate in water (1<= DF <=1/100) to a total volume of 50 ul
- Similarly dilute blank DNA buffer solution with water to a total volume of 50 ul
- Read absobance of blank and DNA sample at lambda = 260 and 280
- Calculate [DNA]; [DNA](ng/ul) = DF*A260*50
- Determine sample purity; pure DNA A260/A280 = 1.8
 "
"