Team:Newcastle/Metalsensing
From 2009.igem.org
Metal Sensing
This sub-project focusses on getting the Bacillus subtilis to sense the cadmium and being able to respond to this detection. Please click on the links below for more information:
- Introduction
- Modelling
- BioBrick Constructs
- Lab Work Strategies
- Other Presentations and Diagrams
- Lab Work done
Introduction
If our project is to process cadmium and not other metals, we need to genetically engineer Bacillus subtilis to carry out a set of cellular processes based on the action of metal sensors. These metal sensors will detect cadmium through a system known as AND Gating. This system of combined metal sensors ensures the Bacillus subtilis cells have a tightly controlled cadmium sensor, as naturally occurring metal sensitive promoters can respond to OTHER metals too
There are two metal sensing repressors, which are known to respond to cadmium: ArsR and CzrA. However, their specificity for cadmium is not unique. ArsR also detects arsenic and CzrA also detects zinc and copper. The table below shows the metals which release both ArsR and CzrA from their DNA binding sites:
| Metal Sensor | Metals Sensed | |||
|---|---|---|---|---|
| ArsR | As(III) | Ag(I) | Cu | Cd |
| CzrA | Zn | Co | Ni | Cd |
By positioning the operator binding sites for these two metal sensing repressors next to each other in a promoter region, the gene regulated by that promoter will be transcribed only when a metal that binds to both sensors is present; in this case Cadmium.
This is a combinatorial approach for gene expression regulation. To read more information on the cadmium sensor, please visit the Cadmium Sensing section of the Project Overview.
To create this AND gate the team made use of the CadA promoter; this promoter, which usually regulates expression of the CadA cadmium efflux channels in the presence of cadmium ions, bears the CzrA binding site. Therefore if this promoter is coupled to the ArsR binding site, expression of a signal protein will only be made in the presence of cadmium.
Why don't we just use the CadA promoter then? The answer is this - cadA can respond to other metals too so a system totally reliant on the cadA promoter as a sensor would not be efficient in achieving the aims of this project. We need a tightly-controlled cadmium sensor
Modelling
To simulate the effects of cadmium intake and sensing, the Metal Sensing Team constructed models using CellML. Please click on the link here: Metal Intake/Efflux Model
BioBrick constructs
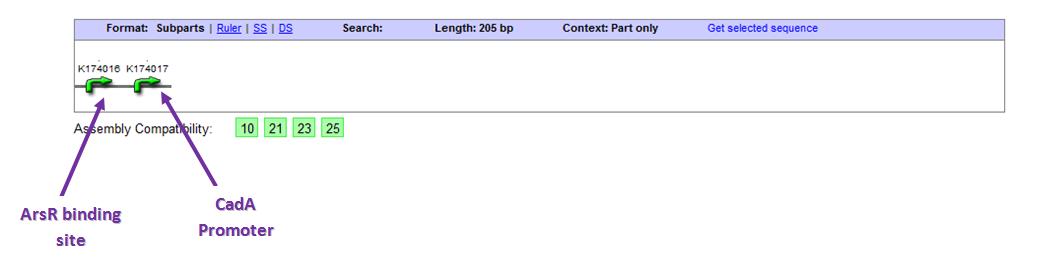
For our cadmium-sensing BioBrick design, we have ligated the ArsR binding site(BBa_K174016) next to the CadA promoter (BBa_K174017). The CadA promoter drives expression of the CadA efflux channel and this expression activity increase in the presence of cadmium. Yet more importantly the CadA promoter region contains the CzrA binding site the team needs for the cadmium-sensor AND gate.
By placing the ArsR binding site next to the CadA promoter, expression of a signal protein will only be achieved if there is cadmium in the cell.
Lab Work Strategies
In carrying out the lab work required for the construction of the cadmium-sensor, we adhered to the following lab strategy. Please click on the link here to review our strategy: Metal Sensor: czrA+arsR.
Other Presentations and Diagrams
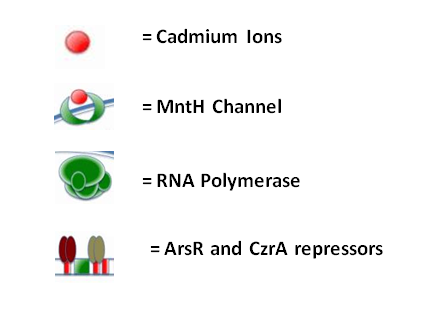
The diagram below illustrates the way in which ArsR and CzrA respond to cadmium ions, which will make their way into the cell via the MntH (manganese transporter) channel:
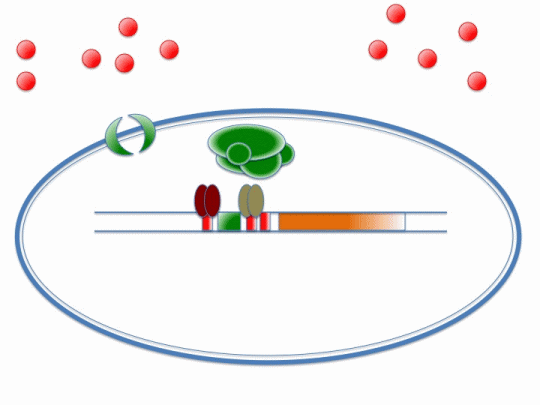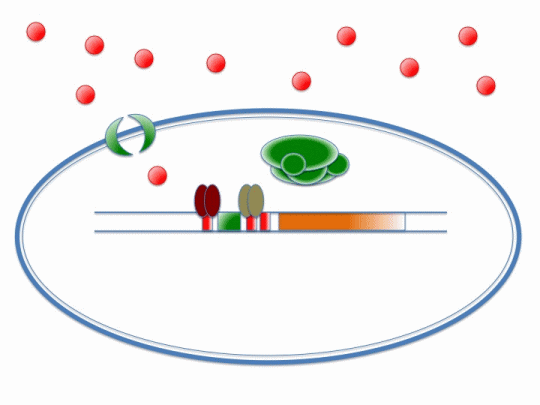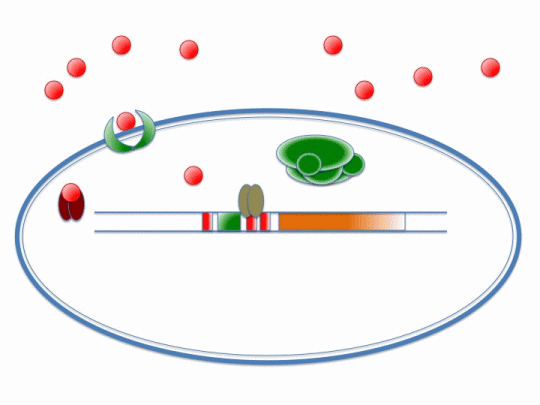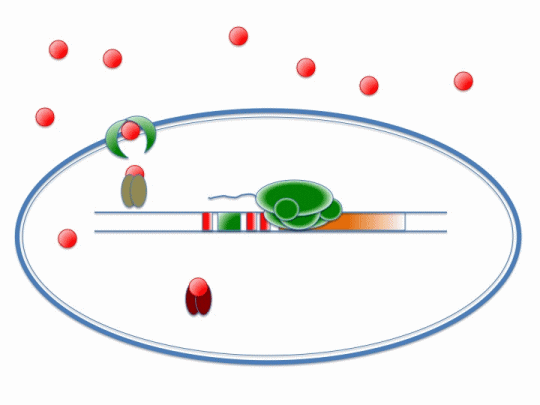
Please refer the key below to better understand this diagram! In short, cadmium ions make their way through the MntH channel and bind to the ArsR and CzrA repressor proteins. Their removal allows the DNA polymerase to make a signal protein which will feed into the stochastic switch.
Lab Work done
This is all of the lab work in which the Metal Sensing Team attempted to construct the cadmium sensor - click on the dates to read that day's lab entry:
| Summary of Lab Sessions for Cadmium Sensing | |
|---|---|
| | |
| 18th August 2009 | Transformed DH5-alpha E. coli cells with BBa_J33206 from the Spring Distribution |
| 19th August 2009 | Inoculated 3 tubes of LB with 3 colonies of potential transformant E.coli cells |
| 20th August 2009 | Conduct mini-preps on the three overnight-grown cultures of potential BBa_J33206 transformants and digested them with restriction enzymes. |
| 21st August 2009 | Analysed digested BBa_J33206 mini-prep DNA by DNA gel electrophoresis and prepared midi-preps also. |
| 25th August 2009 | Concentrated (and ethanol precipitated) the BBa_J33206 Midi-prep sample. |
| 26th August 2009 | Digested BBa_J33206 midi-prep DNA with EcoRI and PstI and analysed through DNA gel electrophoresis - digest reaction not successful (a second attempt at digests needed) |
| 27th August 2009 | Digested pSB1A2 (containing BBa_J33206 BioBrick), ran DNA though gel and excised band. Also analysed digested pSB1A2 (BBa_J33206 BioBrick) in gel - bands erroneous |
| 28th August 2009 | Cleaned gel band using Gel extraction kit |
| 1st September 2009 | Attempted to PCR amplify the needed czrA gene from the genome of Bacillus subtilis |
| 2nd September 2009 | PCR amplification of czrA gene failed - conducted B. subtilis genome prep and carried out PCR reaction on this DNA |
| 3rd September 2009 | Second attempt at czrA PCR amplification analysed on gel - also unsuccessful. Reattempted PCR amplification of czrA on B. subtilis genomic DNA at different annealing temperatures. |
| 4th September 2009 | In light of 27/08/09 lab session (i.e. anomalous bands with digested BBa_J33206 BioBrick), we have sent away the BioBrick for sequencing and are now using BBa_J33206 sent to us by Chris French. Used Chris's BBa_J33206 to transform E. coli cells. Also carried out PCR reactions |
| 7th September 2009 | Inoculated LB media with colonies of BBa_J33206 (sent from Chris French) E. coli transformants for mini preps. By afternoon, cultures had grown sufficiently to further inoculate flasks of 50ml LB + amp for midi-preps. Mini-prep attempted but abandoned |
| 8th September 2009 | Skipped mini-prep re-attempt and immediately carried out midi-prep of BBa_J33206 BioBrick sent by Chris French. Digested sample with EcoRI and PstI and analysed through gel. Successful! |
| 10th September 2009 | PCR amplification of the cadA promoter and BBa_J33206 BioBrick (missing promoter) |
| 11th September 2009 | Gel analysis shows PCR reactions worked. Tried to proceed with running the cleaned-up PCR products through gel to excise band but ethanol presence stopped us. Will have to reattempt PCR reactions and other subsequent steps. |
| 14th September 2009 | Both cadA promoter and BBa_J33206 (with promoter missing) PCR reactions worked! Subsequently excised from gel and cleaned up. Both fragments cut with BamHI and NheI and ligated together. Attempted to transform E. coli with ligated cadmium-sensor. |
| 15th September 2009 | No transformant colonies spotted on plates - either ligations or transformations didn't work. Reattempted ligation and transformation in E. coli cells. |
| 16th September 2009 | Transformations failed! Reattempted digesting the BBa_J33206 (with no promoter) and cadA promoter with BamHI and NheI and also reattempted ligations |
| 17th September 2009 | Subsequent work on cadmium sensor put on hold |
News
Events
- 20 – 21 June 2009 - Europe workshop (London)
- 23 – 24 June 2009 - UK iGEM meetup (Edinburgh)
- 23 October Practice Presentation (Newcastle)
- 23 October T-shirts are ready
- 27 October Practice Presentation (Sunderland)
- 27 October Poster is ready
- 30 October – 2 November 2009 - Jamboree (Boston)
Social Net
 "
"