Our project goal is to maximize the photosynthetic productivity for a series of photobioreactors containing Rhodobacter sphaeroides under both high and low light intensities by synthetically regulating the size of the light-harvesting antenna complex LH2. We chose to undertake such a project in R. sphaeroides due to its well-characterized photosynthetic and genetic system.
The antenna system functions to expand the spectrum of light available for photosynthesis by absorbing different wavelengths than that of the reaction center. Essentially, it is like the dish around the main receiver of any antenna. Marine bacteria, such as R. sphaeroides, evolved very large antenna complexes to absorb light in a natural environment where there is great competition for photons. As a result, the photosynthetic machinery is saturated at low light intensities in a synthetic non-competitive environment, such as a bioreactor. This causes up to 95% of incidental photons to be dissipated as heat or fluorescence by bacteria through a process called Non-Photochemical Quenching (NPQ) (Mussgnug et al., 2007). In essence, these photons are being wasted as NPQ reduces light penetration into and across bioreactors and starves the shaded cells for for photons. One method that has been shown to improve photosynthetic efficiency is the reduction of light-harvesting antenna sizes (Polle et al., 2002, Mussgnug et al., 2007). Though, current approaches to this end are difficult to precisely control from the perspective of biological engineering and synthetic biology.
Our intention is to create a dynamic system to vary antenna size that is dependent on incident light intensity and that can be readily optimized using bioengineering principles. This synthetic regulation of the LH2 complex should result in the bacteria grown under high light intensities expressing fewer LH2 complexes than the cells that are more shaded, leading to a reduction of wasted incident photons while maintaining a high overall absorbance across subsequent bioreactors. Consequently, we expect to see an increase in the total photosynthetic productivity for our mutant Rhodobacter sphaeroides when compared to the wild type grown under similar conditions.
Rhodobacter sphaeroides is a purple Alphaproteobacteria. It is flexible metabolically and can grow heterotrophically via aerobic and anaerobic respiration, as well as phototrophically under anaerobic conditions with light.
R. sphaeroides is one of the best understood photosynthetic organisms. The photosystem is located in intracytoplasmic membrane invaginations and the main components are the Light Haresting Complex 2 (LH2), Light Harvesting Complex 1 (LH1), and the Reaction Center (RC). These pigment-protein complexes non-covalently bind bacteriochlorophylls and carotenoids. LH1 and RC make up the core complex and are often found in the ratio 1:1. LH2 is more peripheral and naturally ranges in the ratio to LH1/RC of 3.0:1 to 6.7:1 under varying light conditions (Scheuring et al., 2005). LH2 absorbs photons maximally at the wavelength of 842 nm and funnels its energy to LH1 and the reaction center for photochemistry. (Scheuring et al., 2005).
The two subunits of LH2 are coded for by the pucB/A genes and are naturally promoted by the puc promoter.
Codon Usage Table for R. sphaeroides 2.41
R. sphaeroides 2.41 NCBI Genome Tools
In silico PCR of R. sphaeroides 2.41 genome
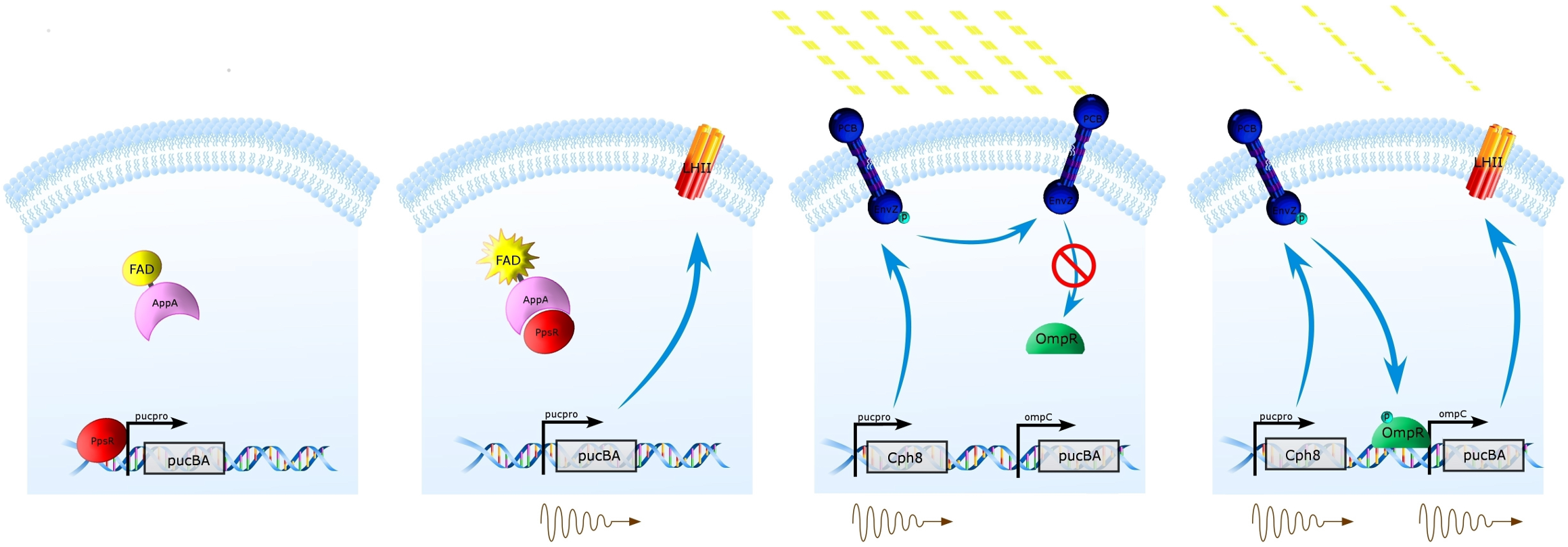 The Natural System (Left) Under high oxygen tension, PpsR prevents transcription from the puc promoter of the pucBA genes, and thus prevents the expression of LH2(1st panel). Under low oxygen tension, this repression is removed by activated AppA, thus allowing expression of LH2 from pucBA(2nd panel).
The Natural System (Left) Under high oxygen tension, PpsR prevents transcription from the puc promoter of the pucBA genes, and thus prevents the expression of LH2(1st panel). Under low oxygen tension, this repression is removed by activated AppA, thus allowing expression of LH2 from pucBA(2nd panel).
Synthetic Regulation System (Right) Our system takes advantage of constitutive protein expression from the puc promoter at low oxygen tension, and replaces pucBA with Cph8 at this site. Placing pucBA expression under the control of the ompC promoter allows for Cph8/OmpR control of LHII expression under varying light conditions. Under high intensity of incident light, Cph8 will not be autophosphorylated, preventing expression of pucBA (3rd panel). When light intensity is lower, Cph8 will autophosphorylate, thus phosphorylating and activating OmpR and promoting expression of LH2 from the ompC promoter.
Figure 1
a Tissue Flask Position 1 Optical Density of WT and DBComega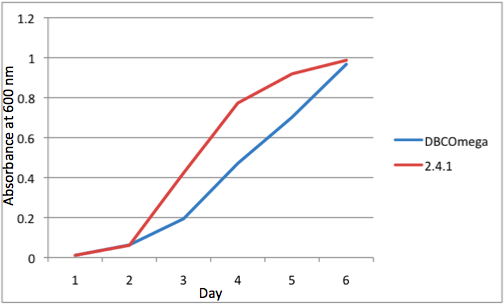 b Tissue Flask Position 2 Optical Density of WT and DBComega
b Tissue Flask Position 2 Optical Density of WT and DBComega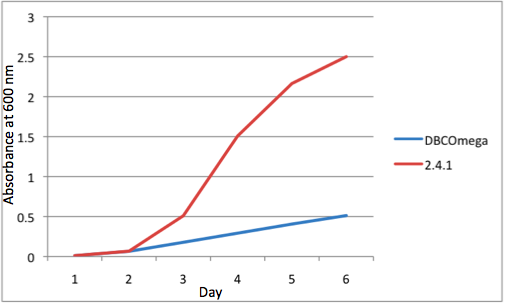
c Tissue Flask Position 3 Optical Density of WT and DBComega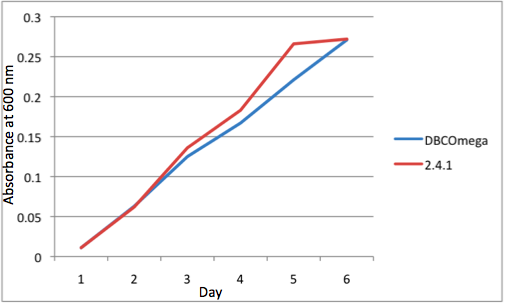 d Tissue Flask Position 4 Optical Density of WT and DBComega
d Tissue Flask Position 4 Optical Density of WT and DBComega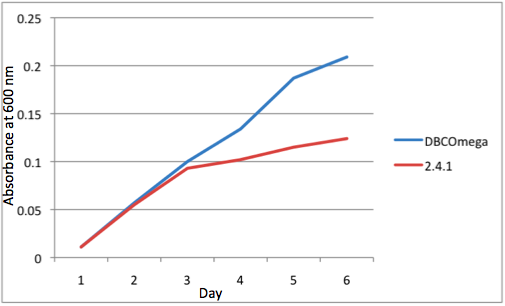
e Tissue Flask Position 5 Optical Density of WT and DBComega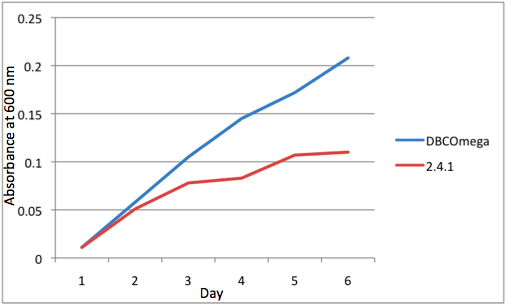 Figure 1a,b,c,d,e are the optical density of a tissue flask, as assessed at 600 nm, over the course of the 6 days of the experiment.
Figure 1a,b,c,d,e are the optical density of a tissue flask, as assessed at 600 nm, over the course of the 6 days of the experiment.
Figure 2
a The Change in the Optical Density of the 5 Different Tissue Flasks Over the Course of 6 Days for the WT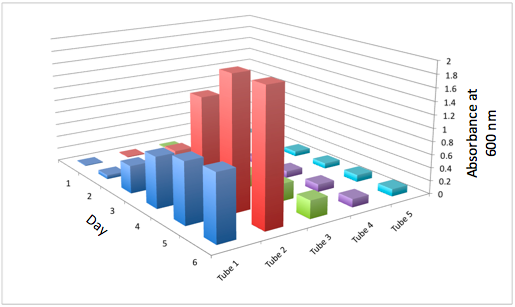 b The Change in the Optical Density of the 5 Different Tissue Flasks Over the Course of 6 Days for DBComega
b The Change in the Optical Density of the 5 Different Tissue Flasks Over the Course of 6 Days for DBComega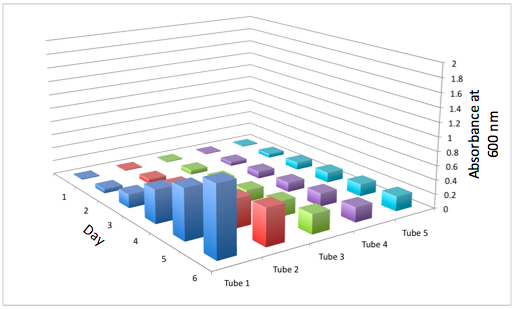
Figure 2a and 2b display the growth of the tissue flasks as measured by OD assessed as absorbance at 600 nm. The graph is organized such that those closest to the "days" axis were closest to the light source (i.e. flask 1 is closer than flask 2).
Figure 3
a Cumulative Growth of the WT Tissue Flasks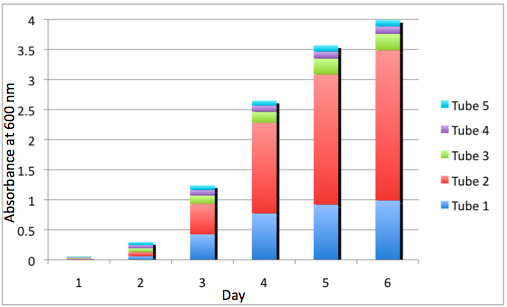 b Cumulative Growth of the DBComega Tissue Flasks
b Cumulative Growth of the DBComega Tissue Flasks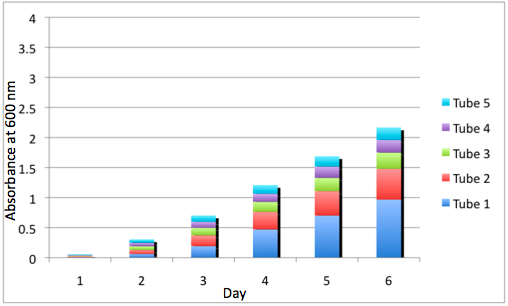
Figure 3a and 3b display the cumulative growth of the tissue flasks as measured by the sum of the optical density of the respective cell type's tissue flasks at a given day. Furthermore, the contribution of a given tissue flask to cumulative growth is displayed.
The tissue flask experiment was an interesting way to observe the effects of a full knockout of the Light Harvesting Antenna Complex 2 (LH2) vs the Wild Type on an array of conseuctive photobioreactors. A couple of conclusions can be drawn from this data regarding the optimization of Light Harvesting Antenna size for photobioreactors.
The most striking difference between the Wild Type Rhodobacter Sphaeroides 2.41 and DBCOmega (LH2 knockout) growth patterns is that the first flask in the WT (closest to the light source) grew less than the second flask, while the converse was true in DBCOmega (figure 2a and 2b). This can be attributed to the photosystem saturation curves for the respective cultures. The Wild Type has an LH2 complex, meaning that their antenna size is inherently larger and that their photosystem will be saturated at lower light intensities than the LH2 deficient mutant. We observed that the incident light intensity used in this experiment led to oversaturation for the first WT tissue flask and resulted in growth-inhibiting photodamage, as is evidenced by its lesser growth relative to the second culture. In contrast, photodamage was not observed in DBCOmega as is evidenced by the first tissue flask that grew at the fastest respective rate. Furthermore, it appears that this photodamage slowed the growth of the first WT tissue flask to the point that the OD of this tissue flask was nearly equal to that of DBCOmega after 5 days of growth(.987 vs. .967) (figure 1a). As such, the contribution of LH2 to the growth of the WT in flask 1 can be assessed to be little- though, this light (at 800 and 850 nm) was still greatly absorbed by flask 1 (see irradiance figure for wt). It is likely that much of this light was wasted through Non-Photochemical Quenching (NPQ), thereby depleting the flasks behind flask 1 of photons at the LH2 absorbtion peaks and decreasing the cumulative productivity of flasks 1-5. This effect of wasting photons though NPQ under high light intensities is one that we sought to minimize in the design of our synthetic regulation for the pucB/A genes.
A second interesting observation from a comparison of the WT and DBCOmega experiments is that the 3rd tissue flask in each had almost identical growth rates (figure 1c), while the DBCOmega 4th and 5th tissue flask outperformed that of the WT (figure 1d and 1e). This can simply be attributed to the higher density of cells in the 2nd flask of the WT that led to an overall lesser quantity of photons passing through to the 4th and 5th flasks in the WT experiment and likely resulting in heterotrophic growth. This can be seen by viewing the absolute irradiance data (incident light intensity at a given wavelength) after the third culture for the WT (point 3, WT Irradiance). This decrease in irradiance is the most pronounced for the LH2 absorption bands at 800 and 850 nm, as was expected.
Thomas, add in some further comments about the sprectroradiometer stuff. Specifically about how it changes in the WT vs. that of DBCOmega using some specific examples on the graphs
Overall, the cumulative culture growth of the WT exceeded that of DBCOmega (figure 3) due to the performance of the second tissue flask of the WT relative to that of DBCOmega (figure 1b) despite the photodamage that occurred in tissue flask 1 for the WT. This cumulative culture growth would be proportional to the amount of energy/desired product obtained from these cultures.
It is an interesting exercise to model how our mutant system would perform under these same conditions: Thomas add in some comments about referencing that diagram and what we took from this tissue flask experiment to construct that (and upload it when you are done with it).
Final conclusion
Future Work
-tissue flask w mutant -characterize puc promoter under light conditions and additional oxygen tensions

|