From 2009.igem.org
Wet
GVP Cluster
Transporters
GlpF
-
| 25uL
| Phusion
|
| 2 uL
| GlpF Fw
|
| 2 uL
| GlpF Rev
|
| 0.5 uL
| GlpF PCR3 (28/07/09) DNA
|
| 21uL
| MQ
|
The resulting PCR product was purified using a high pure PCR product purification kit (Roche).
concentration was measured using Nanodrop;
| 16,0 ng/ul
| GlpFPCR3 30/07
|
| 36.4 ng/ul
| psB1AC3-M digested
|
| 32.0 ng/uL
| GlpF digested
|
Restiction was done on the GlpF PCR3 (30/07) with XbaI and PstI as noted on the notebook of 28/07/09
The resticted GlpF PCR3 (30/07) was ligated with psB1AC3 resticted with EcoRI and SpeI.
-
| 2.0 uL
| T4 Ligase buffer
|
| 0.5 uL
| T4 Ligase
|
| 11.25 uL
| GlpF PCR3
|
Metal Accumulation
Vectors
[http://openwetware.org/wiki/User:Wilfred_J._Poppinga/Notebook/Wilfreds_Project/2009/07/30 Results] were disappointing, no colonies on plate. Hope the fresh ligase and ligase buffer (10x) will help the new ligation that was started. New vectors were cut and purified from gel.
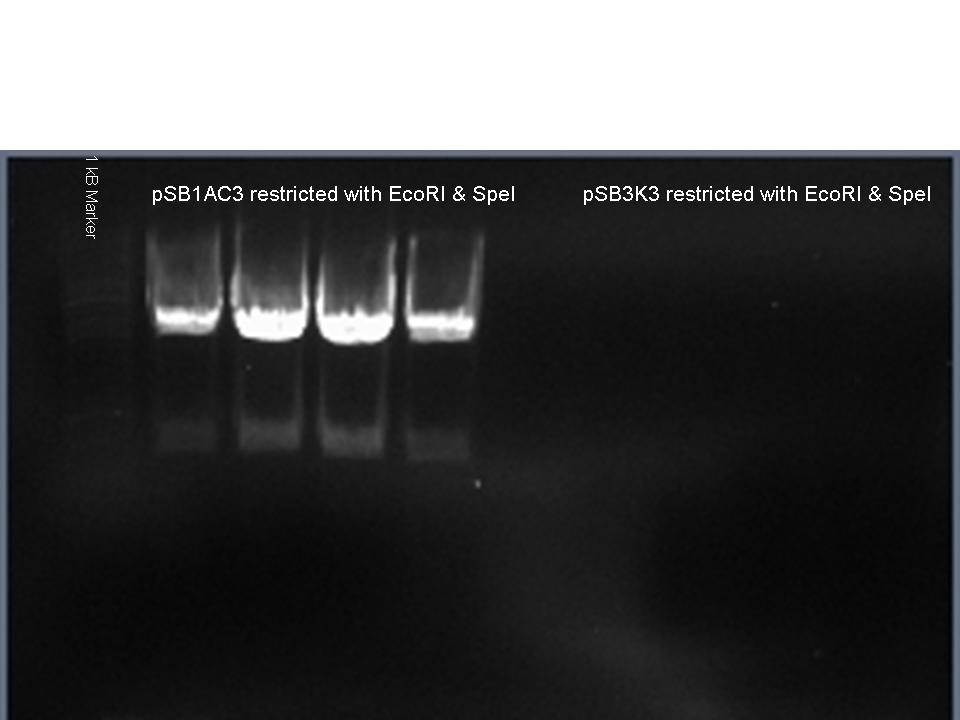
- Ligation (ratio 1:3) of pSB1AC3-HML and pSB3K3-HML + RFP and pSB3K3 and pSB1AC3 + pLac
- Calculated the volumes of vector and insert with Cloning Tools 6.0, using:
- Amount: 200ng
- Excess insert: 3x
In a total volume of 20uL, 2uL 10x Ligase Buffer and 1uL T4 DNA Ligase.
- The fragments were ligated using PCR program AC3 ligation:
| AC3 ligation program
| Temperature
| Time
|
| DNA Denaturing
| 42°
| 2 min
|
| Ligation
| 16°
| 1 hr
|
| Protein inactivation
| 65°
| 10 min
|
- Use 5uL ligation mixture for transformation.
- Transform E. coli TOP10 via normal protocol.
Dry
We worked on migrating graphs from Google spreadsheets to Wiki tables, making sure that at least the raw data used to generate the graphs is "safe".
| April
|
| M | T | W | T | F | S | S
|
|
|
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/1_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 1]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/2_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 2]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/3_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 3]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/4_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 4]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/5_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 5]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/6_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 6]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/7_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 7]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/8_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 8]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/9_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 9]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/10_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 10]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/11_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 11]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/12_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 12]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/13_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 13]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/14_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 14]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/15_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 15]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/16_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 16]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/17_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 17]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/18_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 18]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/19_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 19]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/20_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 20]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/21_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 21]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/22_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 22]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/23_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 23]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/24_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 24]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/25_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 25]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/26_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 26]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/27_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 27]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/28_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 28]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/29_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 29]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/30_April_2009&preload=Team:Groningen/Notebook/Day&action=edit 30]
|
|
| May
|
| M | T | W | T | F | S | S
|
|
|
|
|
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/1_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 1]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/2_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 2]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/3_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 3]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/4_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 4]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/5_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 5]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/6_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 6]
| [http://2009.igem.org/Team:Groningen/Notebook/7_May_2009 7]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/8_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 8]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/9_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 9]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/10_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 10]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/11_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 11]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/12_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 12]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/13_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 13]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/14_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 14]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/15_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 15]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/16_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 16]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/17_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 17]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/18_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 18]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/19_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 19]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/20_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 20]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/21_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 21]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/22_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 22]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/23_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 23]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/24_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 24]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/25_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 25]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/26_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 26]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/27_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 27]
| [http://2009.igem.org/Team:Groningen/Notebook/28_May_2009 28]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/29_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 29]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/30_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 30]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/31_May_2009&preload=Team:Groningen/Notebook/Day&action=edit 31]
|
|
| June
|
| M | T | W | T | F | S | S
|
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/1_June_2009&preload=Team:Groningen/Notebook/Day&action=edit 1]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/2_June_2009&preload=Team:Groningen/Notebook/Day&action=edit 2]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/3_June_2009&preload=Team:Groningen/Notebook/Day&action=edit 3]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/4_June_2009&preload=Team:Groningen/Notebook/Day&action=edit 4]
| [http://2009.igem.org/Team:Groningen/Notebook/5_June_2009 5]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/6_June_2009&preload=Team:Groningen/Notebook/Day&action=edit 6]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/7_June_2009&preload=Team:Groningen/Notebook/Day&action=edit 7]
|
| [http://2009.igem.org/Team:Groningen/Notebook/8_June_2009 8]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/9_June_2009&preload=Team:Groningen/Notebook/Day&action=edit 9]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/10_June_2009&preload=Team:Groningen/Notebook/Day&action=edit 10]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/11_June_2009&preload=Team:Groningen/Notebook/Day&action=edit 11]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/12_June_2009&preload=Team:Groningen/Notebook/Day&action=edit 12]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/13_June_2009&preload=Team:Groningen/Notebook/Day&action=edit 13]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/14_June_2009&preload=Team:Groningen/Notebook/Day&action=edit 14]
|
| [http://2009.igem.org/Team:Groningen/Notebook/15_June_2009 15]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/16_June_2009&preload=Team:Groningen/Notebook/Day&action=edit 16]
| [http://2009.igem.org/Team:Groningen/Notebook/17_June_2009 17]
| [http://2009.igem.org/Team:Groningen/Notebook/18_June_2009 18]
| [http://2009.igem.org/Team:Groningen/Notebook/19_June_2009 19]
| [http://2009.igem.org/Team:Groningen/Notebook/20_June_2009 20]
| [http://2009.igem.org/Team:Groningen/Notebook/21_June_2009 21]
|
| [http://2009.igem.org/Team:Groningen/Notebook/22_June_2009 22]
| [http://2009.igem.org/Team:Groningen/Notebook/23_June_2009 23]
| [http://2009.igem.org/Team:Groningen/Notebook/24_June_2009 24]
| [http://2009.igem.org/Team:Groningen/Notebook/25_June_2009 25]
| [http://2009.igem.org/Team:Groningen/Notebook/26_June_2009 26]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/27_June_2009&preload=Team:Groningen/Notebook/Day&action=edit 27]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/28_June_2009&preload=Team:Groningen/Notebook/Day&action=edit 28]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/29_June_2009&preload=Team:Groningen/Notebook/Day&action=edit 29]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/30_June_2009&preload=Team:Groningen/Notebook/Day&action=edit 30]
|
|
| July
|
| M | T | W | T | F | S | S
|
|
|
| [http://2009.igem.org/Team:Groningen/Notebook/1_July_2009 1]
| [http://2009.igem.org/Team:Groningen/Notebook/2_July_2009 2]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/3_July_2009&preload=Team:Groningen/Notebook/Day&action=edit 3]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/4_July_2009&preload=Team:Groningen/Notebook/Day&action=edit 4]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/5_July_2009&preload=Team:Groningen/Notebook/Day&action=edit 5]
|
| [http://2009.igem.org/Team:Groningen/Notebook/6_July_2009 6]
| [http://2009.igem.org/Team:Groningen/Notebook/7_July_2009 7]
| [http://2009.igem.org/Team:Groningen/Notebook/8_July_2009 8]
| [http://2009.igem.org/Team:Groningen/Notebook/9_July_2009 9]
| [http://2009.igem.org/Team:Groningen/Notebook/10_July_2009 10]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/11_July_2009&preload=Team:Groningen/Notebook/Day&action=edit 11]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/12_July_2009&preload=Team:Groningen/Notebook/Day&action=edit 12]
|
| [http://2009.igem.org/Team:Groningen/Notebook/13_July_2009 13]
| [http://2009.igem.org/Team:Groningen/Notebook/14_July_2009 14]
| [http://2009.igem.org/Team:Groningen/Notebook/15_July_2009 15]
| [http://2009.igem.org/Team:Groningen/Notebook/16_July_2009 16]
| [http://2009.igem.org/Team:Groningen/Notebook/17_July_2009 17]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/18_July_2009&preload=Team:Groningen/Notebook/Day&action=edit 18]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/19_July_2009&preload=Team:Groningen/Notebook/Day&action=edit 19]
|
| [http://2009.igem.org/Team:Groningen/Notebook/20_July_2009 20]
| [http://2009.igem.org/Team:Groningen/Notebook/21_July_2009 21]
| [http://2009.igem.org/Team:Groningen/Notebook/22_July_2009 22]
| [http://2009.igem.org/Team:Groningen/Notebook/23_July_2009 23]
| [http://2009.igem.org/Team:Groningen/Notebook/24_July_2009 24]
| [http://2009.igem.org/Team:Groningen/Notebook/25_July_2009 25]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/26_July_2009&preload=Team:Groningen/Notebook/Day&action=edit 26]
|
| [http://2009.igem.org/Team:Groningen/Notebook/27_July_2009 27]
| [http://2009.igem.org/Team:Groningen/Notebook/28_July_2009 28]
| [http://2009.igem.org/Team:Groningen/Notebook/29_July_2009 29]
| [http://2009.igem.org/Team:Groningen/Notebook/30_July_2009 30]
| [http://2009.igem.org/Team:Groningen/Notebook/31_July_2009 31]
|
|
| August
|
| M | T | W | T | F | S | S
|
|
|
|
|
|
| [http://2009.igem.org/Team:Groningen/Notebook/1_August_2009 1]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/2_August_2009&preload=Team:Groningen/Notebook/Day&action=edit 2]
|
| [http://2009.igem.org/Team:Groningen/Notebook/3_August_2009 3]
| [http://2009.igem.org/Team:Groningen/Notebook/4_August_2009 4]
| [http://2009.igem.org/Team:Groningen/Notebook/5_August_2009 5]
| [http://2009.igem.org/Team:Groningen/Notebook/6_August_2009 6]
| [http://2009.igem.org/Team:Groningen/Notebook/7_August_2009 7]
| [http://2009.igem.org/Team:Groningen/Notebook/8_August_2009 8]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/9_August_2009&preload=Team:Groningen/Notebook/Day&action=edit 9]
|
| [http://2009.igem.org/Team:Groningen/Notebook/10_August_2009 10]
| [http://2009.igem.org/Team:Groningen/Notebook/11_August_2009 11]
| [http://2009.igem.org/Team:Groningen/Notebook/12_August_2009 12]
| [http://2009.igem.org/Team:Groningen/Notebook/13_August_2009 13]
| [http://2009.igem.org/Team:Groningen/Notebook/14_August_2009 14]
| [http://2009.igem.org/Team:Groningen/Notebook/15_August_2009 15]
| [http://2009.igem.org/Team:Groningen/Notebook/16_August_2009 16]
|
| [http://2009.igem.org/Team:Groningen/Notebook/17_August_2009 17]
| [http://2009.igem.org/Team:Groningen/Notebook/18_August_2009 18]
| [http://2009.igem.org/Team:Groningen/Notebook/19_August_2009 19]
| [http://2009.igem.org/Team:Groningen/Notebook/20_August_2009 20]
| [http://2009.igem.org/Team:Groningen/Notebook/21_August_2009 21]
| [http://2009.igem.org/Team:Groningen/Notebook/22_August_2009 22]
| [http://2009.igem.org/Team:Groningen/Notebook/23_August_2009 23]
|
| [http://2009.igem.org/Team:Groningen/Notebook/24_August_2009 24]
| [http://2009.igem.org/Team:Groningen/Notebook/25_August_2009 25]
| [http://2009.igem.org/Team:Groningen/Notebook/26_August_2009 26]
| [http://2009.igem.org/Team:Groningen/Notebook/27_August_2009 27]
| [http://2009.igem.org/Team:Groningen/Notebook/28_August_2009 28]
| [http://2009.igem.org/Team:Groningen/Notebook/29_August_2009 29]
| [http://2009.igem.org/Team:Groningen/Notebook/30_August_2009 30]
|
| [http://2009.igem.org/Team:Groningen/Notebook/31_August_2009 31]
|
|
| September
|
| M | T | W | T | F | S | S
|
|
| [http://2009.igem.org/Team:Groningen/Notebook/1_September_2009 1]
| [http://2009.igem.org/Team:Groningen/Notebook/2_September_2009 2]
| [http://2009.igem.org/Team:Groningen/Notebook/3_September_2009 3]
| [http://2009.igem.org/Team:Groningen/Notebook/4_September_2009 4]
| [http://2009.igem.org/Team:Groningen/Notebook/5_September_2009 5]
| [http://2009.igem.org/Team:Groningen/Notebook/6_September_2009 6]
|
| [http://2009.igem.org/Team:Groningen/Notebook/7_September_2009 7]
| [http://2009.igem.org/Team:Groningen/Notebook/8_September_2009 8]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/9_September_2009&preload=Team:Groningen/Notebook/Day&action=edit 9]
| [http://2009.igem.org/Team:Groningen/Notebook/10_September_2009 10]
| [http://2009.igem.org/Team:Groningen/Notebook/11_September_2009 11]
| [http://2009.igem.org/Team:Groningen/Notebook/12_September_2009 12]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/13_September_2009&preload=Team:Groningen/Notebook/Day&action=edit 13]
|
| [http://2009.igem.org/Team:Groningen/Notebook/14_September_2009 14]
| [http://2009.igem.org/Team:Groningen/Notebook/15_September_2009 15]
| [http://2009.igem.org/Team:Groningen/Notebook/16_September_2009 16]
| [http://2009.igem.org/Team:Groningen/Notebook/17_September_2009 17]
| [http://2009.igem.org/Team:Groningen/Notebook/18_September_2009 18]
| [http://2009.igem.org/Team:Groningen/Notebook/19_September_2009 19]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/20_September_2009&preload=Team:Groningen/Notebook/Day&action=edit 20]
|
| [http://2009.igem.org/Team:Groningen/Notebook/21_September_2009 21]
| [http://2009.igem.org/Team:Groningen/Notebook/22_September_2009 22]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/23_September_2009&preload=Team:Groningen/Notebook/Day&action=edit 23]
| [http://2009.igem.org/Team:Groningen/Notebook/24_September_2009 24]
| [http://2009.igem.org/Team:Groningen/Notebook/25_September_2009 25]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/26_September_2009&preload=Team:Groningen/Notebook/Day&action=edit 26]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/27_September_2009&preload=Team:Groningen/Notebook/Day&action=edit 27]
|
| [http://2009.igem.org/Team:Groningen/Notebook/28_September_2009 28]
| [http://2009.igem.org/Team:Groningen/Notebook/29_September_2009 29]
| [http://2009.igem.org/Team:Groningen/Notebook/30_September_2009 30]
|
|
| October
|
| M | T | W | T | F | S | S
|
|
|
|
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/1_October_2009&preload=Team:Groningen/Notebook/Day&action=edit 1]
| [http://2009.igem.org/Team:Groningen/Notebook/2_October_2009 2]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/3_October_2009&preload=Team:Groningen/Notebook/Day&action=edit 3]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/4_October_2009&preload=Team:Groningen/Notebook/Day&action=edit 4]
|
| [http://2009.igem.org/Team:Groningen/Notebook/5_October_2009 5]
| [http://2009.igem.org/Team:Groningen/Notebook/6_October_2009 6]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/7_October_2009&preload=Team:Groningen/Notebook/Day&action=edit 7]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/8_October_2009&preload=Team:Groningen/Notebook/Day&action=edit 8]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/9_October_2009&preload=Team:Groningen/Notebook/Day&action=edit 9]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/10_October_2009&preload=Team:Groningen/Notebook/Day&action=edit 10]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/11_October_2009&preload=Team:Groningen/Notebook/Day&action=edit 11]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/12_October_2009&preload=Team:Groningen/Notebook/Day&action=edit 12]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/13_October_2009&preload=Team:Groningen/Notebook/Day&action=edit 13]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/14_October_2009&preload=Team:Groningen/Notebook/Day&action=edit 14]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/15_October_2009&preload=Team:Groningen/Notebook/Day&action=edit 15]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/16_October_2009&preload=Team:Groningen/Notebook/Day&action=edit 16]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/17_October_2009&preload=Team:Groningen/Notebook/Day&action=edit 17]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/18_October_2009&preload=Team:Groningen/Notebook/Day&action=edit 18]
|
| [http://2009.igem.org/Team:Groningen/Notebook/19_October_2009 19]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/20_October_2009&preload=Team:Groningen/Notebook/Day&action=edit 20]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/21_October_2009&preload=Team:Groningen/Notebook/Day&action=edit 21]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/22_October_2009&preload=Team:Groningen/Notebook/Day&action=edit 22]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/23_October_2009&preload=Team:Groningen/Notebook/Day&action=edit 23]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/24_October_2009&preload=Team:Groningen/Notebook/Day&action=edit 24]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/25_October_2009&preload=Team:Groningen/Notebook/Day&action=edit 25]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/26_October_2009&preload=Team:Groningen/Notebook/Day&action=edit 26]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/27_October_2009&preload=Team:Groningen/Notebook/Day&action=edit 27]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/28_October_2009&preload=Team:Groningen/Notebook/Day&action=edit 28]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/29_October_2009&preload=Team:Groningen/Notebook/Day&action=edit 29]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/30_October_2009&preload=Team:Groningen/Notebook/Day&action=edit 30]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/31_October_2009&preload=Team:Groningen/Notebook/Day&action=edit 31]
|
|
| November
|
| M | T | W | T | F | S | S
|
|
|
|
|
|
|
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/1_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 1]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/2_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 2]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/3_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 3]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/4_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 4]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/5_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 5]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/6_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 6]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/7_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 7]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/8_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 8]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/9_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 9]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/10_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 10]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/11_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 11]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/12_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 12]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/13_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 13]
| [http://2009.igem.org/Team:Groningen/Notebook/14_November_2009 14]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/15_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 15]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/16_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 16]
| [http://2009.igem.org/Team:Groningen/Notebook/17_November_2009 17]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/18_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 18]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/19_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 19]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/20_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 20]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/21_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 21]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/22_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 22]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/23_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 23]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/24_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 24]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/25_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 25]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/26_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 26]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/27_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 27]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/28_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 28]
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/29_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 29]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Groningen/Notebook/30_November_2009&preload=Team:Groningen/Notebook/Day&action=edit 30]
|
|

 "
"