|
|
| (5 intermediate revisions not shown) |
| Line 1: |
Line 1: |
| | {{Imperial/09/TemplateTop}} | | {{Imperial/09/TemplateTop}} |
| | | | |
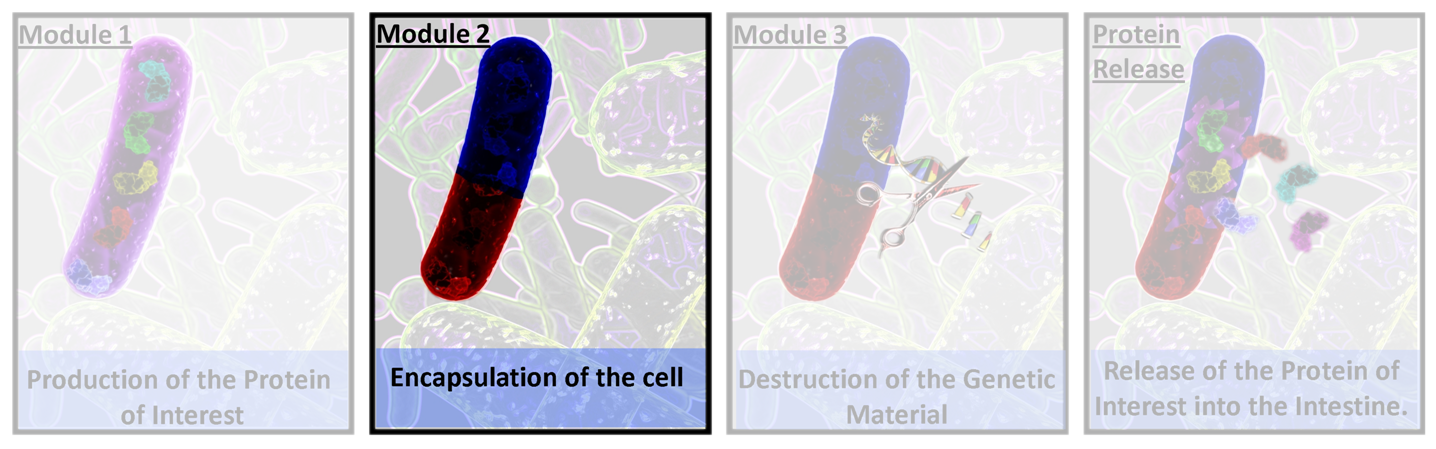
| - | =[[Image:II09_Thumb_m2.png|40px]]<font size='5'><b>Module 2 - Encapsulsation Overview</b></font>= | + | =[[Image:II09_Thumb_m2.png|40px]]<font size='5'><b>Module 2 - Acid Resistance Overview</b></font>= |
| | | | |
| | [[Image:II09_TimelineM2.png |center| 600px]] | | [[Image:II09_TimelineM2.png |center| 600px]] |
| - |
| |
| - | To ferry polypeptides through the stomach, each of our microcapsules must withstand the rigours of a heated protease–rich acid bath. Let there be no illusions, this is no mean feat. The stomach is a highly evolved microbe–mincer that few chassis have the potential to withstand. What is more, since our microcapsules are inanimate, we cannot rely on any of the active acid–resistance strategies that living bacteria are able to deploy.<br><br>
| |
| - |
| |
| - | <html><a href="https://2009.igem.org/Team:Imperial_College_London/Stomach"><img style="vertical-align:bottom;" width=50px align="left" src="http://i691.photobucket.com/albums/vv271/dk806/II09_Learnmore.png"></a></html> <b><i>About proteolysis.</i></b>
| |
| - |
| |
| | | | |
| | {{Imperial/09/Division}} | | {{Imperial/09/Division}} |
| | | | |
| - | ==<b>The need for encapsulation:</b>==
| |
| - |
| |
| - | While <i>E.coli</i> has endogenous acid resistance pathways, colonisation of the gut is based on a "numbers approach". In essence, the majority of <i>E.coli</i> cells in a population do not survive the gut but the few that do are able to regenerate the population once in the intestine.
| |
| - |
| |
| - | This point is illustrated by the experiment shown below:
| |
| - |
| |
| - | [[Image:AcidBath.png|left|150px]]
| |
| - |
| |
| - | Constitutive GFP producing <i>E.coli</i> cells were exposed to differing concentrations of acid for 30 minutes. The reduction in fluoresence is indicative of cell lysis and the subsequent acid-induced denaturing of GFP.
| |
| - |
| |
| - |
| |
| - |
| |
| - |
| |
| - |
| |
| - |
| |
| - |
| |
| - |
| |
| - | To tackle this seemingly insurmountable problem we adopted a two phase approach.
| |
| - |
| |
| - | <b>Phase 1:</b> Identify a suitable chassis with the genotypic potential for acid–resistance.
| |
| - |
| |
| - | <b>Phase 2:</b>Manipulate endogenous acid resistance pathways to control the acid resistant phenotype.
| |
| - |
| |
| - | {{Imperial/09/Division}}
| |
| - |
| |
| - | ==<b>Phase 1:</b>==
| |
| - |
| |
| - | Our rationale for looking for natural sources of acid resistance is that it is easier to hack existing pathways than to transfer large numbers of genes into a different chassis with a dissimilar genetic background.
| |
| - |
| |
| - | Based on natural sources of acid resistance, <i>Lactobacillus</i>, <i>E.coli</i> and <i>B.subtilis</i> were shortlisted as potential chassis.
| |
| - |
| |
| - | Of these three organisms, <i>E.coli</i> was chosen as it is safe, easy to work with and possesses a broad range of acid resistance strategies.
| |
| - |
| |
| - |
| |
| - |
| |
| - |
| |
| - | {{Imperial/09/Division}}
| |
| - |
| |
| - | ==<b>Phase 2:</b>==
| |
| - |
| |
| - |
| |
| - | We hacked E.coli’s acid resistance module in three places to achieve the production of a safe acid resistant capsule.
| |
| - |
| |
| - | <b>Acid Resistant Polymer – Colanic acid:</b>
| |
| - | E.coli naturally produces a harmless acid–resistant polymer known as colanic acid. Colanic acid is a polymer of glucose, galactose and glucuronic acid. By tapping into the pathway that initiates colanic acid biosynthesis, we can turn on its production via the modulation of a transcription factor encoded by a gene called RcsB.
| |
| - |
| |
| - | <html><a href="https://2009.igem.org/Team:Imperial_College_London/M2/genes"><img width=50px src="http://i691.photobucket.com/albums/vv271/dk806/II09_Learnmore.png" align="left"></a></html> <b><i>About RcsB</i></b>
| |
| - |
| |
| - |
| |
| - | <b>Safety – Biofilm prevention:</b>
| |
| - | In nature, colanic acid acts as a binding agent between individual cells over which a biofilm can be formed. While colanic acid itself is harmless, biofilm formation is associated with a number of virulence factors. To prevent biofilm formation from occurring, we have tapped into a second pathway such that our cells become locked into colanic acid production. The gene responsible for preventing biofilm formation is a transcription factor encoded by a gene called YgiV.
| |
| - |
| |
| - | <html><a href="https://2009.igem.org/Team:Imperial_College_London/M2/YgiV"><img width=50px src="http://i691.photobucket.com/albums/vv271/dk806/II09_Learnmore.png" align="left"></a></html> <b><i>About YgiV</i></b>
| |
| - |
| |
| - |
| |
| - | <b>Microencapsulation – Colanic acid tethering:</b>
| |
| - | In nature, colanic acid is associated with but not attached to the cell surface. To facilitate whole cell encapsulation, we have modified a third pathway to fix the colanic acid to the surface of the cell. This involves the over–production of an enzyme called Rfal.
| |
| | | | |
| - | <html><a href="https://2009.igem.org/Team:Imperial_College_London/M2/Rfal"><img width=50px src="http://i691.photobucket.com/albums/vv271/dk806/II09_Learnmore.png" align="left"></a></html> <b><i>About Rfal</i></b>
| |
| | | | |
| | | | |

 "
"