Team:UNICAMP-Brazil/Yeastguard/Killing
From 2009.igem.org
Anezeidler (Talk | contribs) (→Caracterization of the Biobrick) |
Anezeidler (Talk | contribs) (→Caracterization of the Biobrick) |
||
| Line 13: | Line 13: | ||
==Caracterization of the Biobrick== | ==Caracterization of the Biobrick== | ||
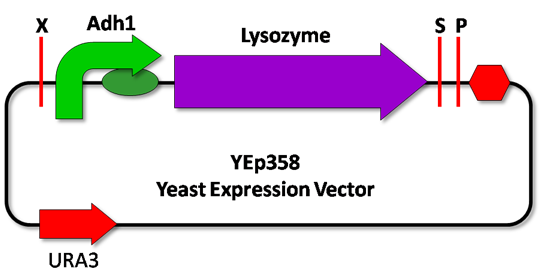
| - | <p style=”text-align:justify;”>The lysozyme biobrick described above is going to be assembled according to the [https://2009.igem.org/Team:UNICAMP-Brazil/Protocols/pGEMStrategy pGEM cloning strategy] proposed by the team. And in order to caracterize the biobrick, its functionality must be seen in yeasts. Thus, the part is going to be transferred to an [https://2009.igem.org/Team:UNICAMP-Brazil/Protocols/YEP yeast expression vector] (YEp358 ura+) and then be transformed into the ''Saccharomyces cerevisiae'' FY23 | + | <p style=”text-align:justify;”>The lysozyme biobrick described above is going to be assembled according to the [https://2009.igem.org/Team:UNICAMP-Brazil/Protocols/pGEMStrategy pGEM cloning strategy] proposed by the team. And in order to caracterize the biobrick, its functionality must be seen in yeasts. Thus, the part is going to be transferred to an [https://2009.igem.org/Team:UNICAMP-Brazil/Protocols/YEP yeast expression vector] (YEp358 ura+) and then be transformed into the ''Saccharomyces cerevisiae'' FY23 URA- strain.</p> |
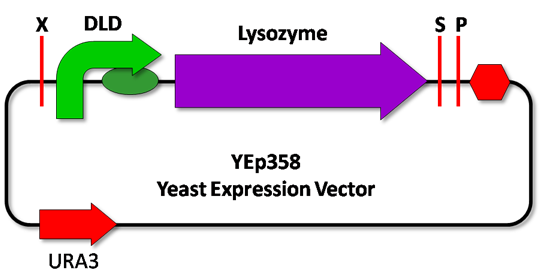
<p style=”text-align:justify;”>The experiments will consists of lysozyme being regulated by a constitutive promoter ADH1(BBa_J63005), and by the two promoters designed for the recognition mechanism: JEN1 and DLD (Figure 2).</p> | <p style=”text-align:justify;”>The experiments will consists of lysozyme being regulated by a constitutive promoter ADH1(BBa_J63005), and by the two promoters designed for the recognition mechanism: JEN1 and DLD (Figure 2).</p> | ||
Revision as of 03:35, 22 October 2009
| ||||||||||||||||||||||||||||||||||
 "
"