Minnesota/24 June 2009
From 2009.igem.org
| (5 intermediate revisions not shown) | |||
| Line 5: | Line 5: | ||
|'''[[Minnesota/23 June 2009|Go to Previous Day (June 23)]]'''|| width=158|'''[[Minnesota/25 June 2009|Go to Next Day (June 25)]]''' | |'''[[Minnesota/23 June 2009|Go to Previous Day (June 23)]]'''|| width=158|'''[[Minnesota/25 June 2009|Go to Next Day (June 25)]]''' | ||
|} | |} | ||
| + | '''Patrick'''<br> | ||
| + | After discovering the error in the program I wished to submit many of the same programs to try and get back up to speed. First was '''smad_base''' which was the simple base model. '''smad_k2u/d''' changed the kinetic constant for the release of tetO1 by the tetR2:tetO1:2aTc complex (reaction 20, normally k = 100000 1/s). '''smad_k1u/d''' changed the kinetic constant for the release of tetO1 by the tetR2:tetO1:aTc complex (reaction 18, normally k = 1 1/s). '''smad_k12u/d''' did both at the same time. The results are expected tomorrow.<br> | ||
| + | |||
| + | '''Ben'''<br> | ||
Since there are several problems with the first TNN model, we are deciding to focus on them one at a time. The first thing to work on is the leakiness problem. In order to deal with this issue, we are including a reaction that allows for the binding of the RNA polymerase to the operator site even if the site is occupied by a tetR. This will allow for some expression at an aTc concentration of 0, because at this point tetR is almost always bound to the tetO operator. | Since there are several problems with the first TNN model, we are deciding to focus on them one at a time. The first thing to work on is the leakiness problem. In order to deal with this issue, we are including a reaction that allows for the binding of the RNA polymerase to the operator site even if the site is occupied by a tetR. This will allow for some expression at an aTc concentration of 0, because at this point tetR is almost always bound to the tetO operator. | ||
| Line 58: | Line 62: | ||
|Ø -> tetR2||1E-11|| | |Ø -> tetR2||1E-11|| | ||
|- | |- | ||
| - | | | + | |aTc_ext -> aTc||3.3E-04|| |
|- | |- | ||
|gfp_mRNA -> Ø||1.16E-03|| | |gfp_mRNA -> Ø||1.16E-03|| | ||
|- | |- | ||
|gfp -> Ø||3.21E-05|| | |gfp -> Ø||3.21E-05|| | ||
| + | |- | ||
| + | |RNAp + tetO1:tetR2 + lacP -> RNAp:lacP + tetR2||311000|| | ||
| + | |- | ||
| + | |RNAp + tetO1:aTc:tetR2 + lacP -> RNAp:lacP + tetR2||311000|| | ||
| + | |- | ||
| + | ||RNAp + tetO1:aTc2:tetR2 + lacP -> RNAp:lacP + tetR2||311000|| | ||
|} | |} | ||
<br> | <br> | ||
| - | [[Image: | + | [[Image:Tnnatcvariesleaky3.jpg|480px]][[Image:Tnnexperiment.jpg|480px]]<br> |
| + | <br> | ||
| + | As you can see from the above graph, this model shows no response to aTc concentration. The gfp output is the same whether there are 0 or 260 molecules of aTc in the cell. Essentially this system is completely leaky, which is not what is needed. Also, the model still peaks and decreases, instead of plateauing. | ||
Latest revision as of 21:52, 30 July 2009
| Back to Notebook Home | |
| Go to Previous Day (June 23) | Go to Next Day (June 25) |
Patrick
After discovering the error in the program I wished to submit many of the same programs to try and get back up to speed. First was smad_base which was the simple base model. smad_k2u/d changed the kinetic constant for the release of tetO1 by the tetR2:tetO1:2aTc complex (reaction 20, normally k = 100000 1/s). smad_k1u/d changed the kinetic constant for the release of tetO1 by the tetR2:tetO1:aTc complex (reaction 18, normally k = 1 1/s). smad_k12u/d did both at the same time. The results are expected tomorrow.
Ben
Since there are several problems with the first TNN model, we are deciding to focus on them one at a time. The first thing to work on is the leakiness problem. In order to deal with this issue, we are including a reaction that allows for the binding of the RNA polymerase to the operator site even if the site is occupied by a tetR. This will allow for some expression at an aTc concentration of 0, because at this point tetR is almost always bound to the tetO operator.
Here is the model we are working on:
| Reaction | Forward Kinetic Constant | Reverse Kinetic Constant |
|---|---|---|
| RNAp + lacP + lacI4:lacO1 -> RNAp:lacP | 6.23E+05 | |
| RNAp + lacP + lacO1 ↔ RNAp:lacP | 1E+07 | 1 |
| RNA:lacP -> RNAp:lacP* | .01 | |
| RNAp:lacP* -> lacP + lacO1 + RNAp:DNAgfp | 30 | |
| RNAp:DNAgfp -> RNAp + gfp_mRNA | 30 | |
| gfp_mRNA + rib -> rib:gfp_mRNA | 100000 | |
| rib:gfp_mRNA -> rib:gfp_mRNA_1 + gfp_mRNA | 33 | |
| rib:gfp_mRNA_1 -> rib + gfp | 33 | |
| tetR2 + aTc ↔ tetR2:aTc | 2E+09 | 4E-04 |
| tetR2:aTc + aTc ↔ tetR2:aTc2 | 1E+08 | 1E-03 |
| tetR2:aTc + tetO1 ↔ tetR2:tetO1:aTc | 1E+08 | 1 |
| tetR2:aTc2 + tetO1 ↔ tetR2:tetO1:aTc2 | 1E+08 | 1E+05 |
| tetR2:tetO1 + aTc ↔ tetR2:tetO1:aTc | 1E+08 | 1E-03 |
| tetR2:tetO1:aTc + aTc ↔ tetR2:tetO1:aTc2 | 1E+08 | 1E-03 |
| tetR2 -> Ø | 2.89E-04 | |
| tetR2:aTc -> aTc | 2.89E-04 | |
| tetR2:aTc2 -> 2 aTc | 2.89E-04 | |
| tetR2 + nsDNA ↔ tetR2:nsDNA | 1000 | 3.2409 |
| tetR2:aTc + nsDNA ↔ tetR2:aTc:nsDNA | 1000 | 3.2409 |
| tetR2:aTc:nsDNA -> aTc + nsDNA | 1.93E-04 | |
| tetR2:nsDNA -> nsDNA | 1.93E-04 | |
| Ø -> tetR2 | 1E-11 | |
| aTc_ext -> aTc | 3.3E-04 | |
| gfp_mRNA -> Ø | 1.16E-03 | |
| gfp -> Ø | 3.21E-05 | |
| RNAp + tetO1:tetR2 + lacP -> RNAp:lacP + tetR2 | 311000 | |
| RNAp + tetO1:aTc:tetR2 + lacP -> RNAp:lacP + tetR2 | 311000 | |
| RNAp + tetO1:aTc2:tetR2 + lacP -> RNAp:lacP + tetR2 | 311000 |
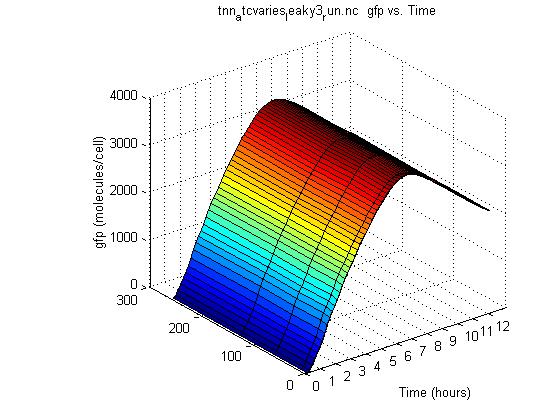
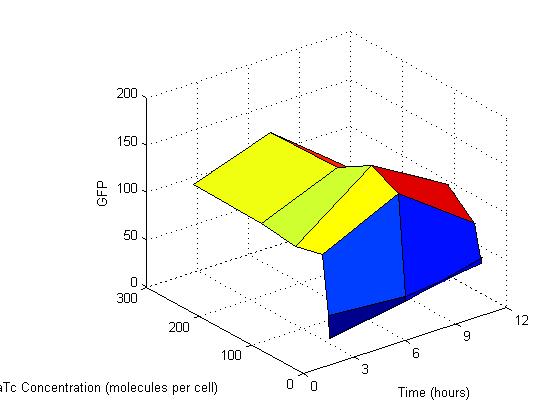
As you can see from the above graph, this model shows no response to aTc concentration. The gfp output is the same whether there are 0 or 260 molecules of aTc in the cell. Essentially this system is completely leaky, which is not what is needed. Also, the model still peaks and decreases, instead of plateauing.
 "
"