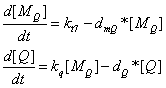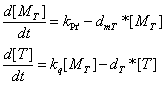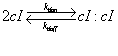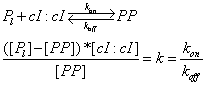Team:Tsinghua/Modeling
From 2009.igem.org
GuoQiangChen (Talk | contribs) |
(→Replication of plasmid) |
||
| (24 intermediate revisions not shown) | |||
| Line 4: | Line 4: | ||
| + | =Introduction= | ||
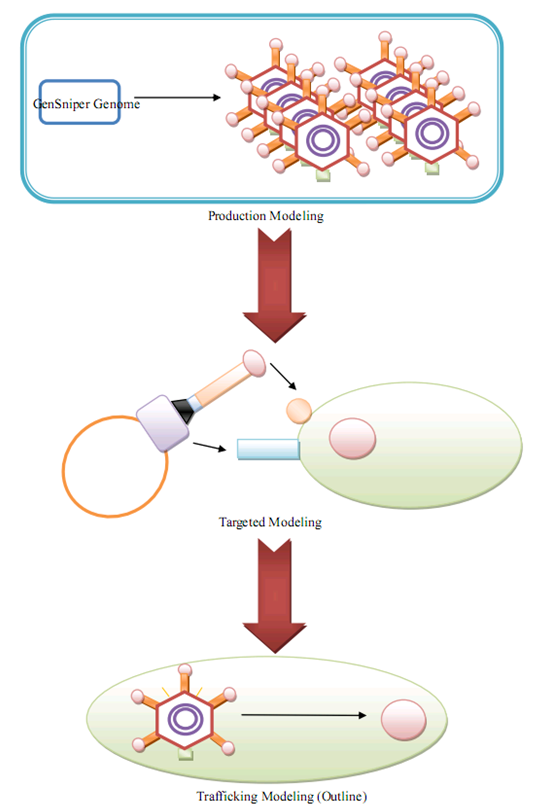
| + | Our wetlab design is subdivided into three interrelated modules ('''in parallel''') according to the BioBrick principle. In contrast, our drylab design is '''in line'''. Three subdividions are proposed, with one of them just outlined due to lack of experimental evidence. | ||
| + | [[Image:dryoutline.png|800px|center|thumb|Modeling Scheme]] | ||
| - | = | + | =SynGenome Based GenSniper Production Modeling= |
In this part, we will use mathematical methods(by solve a set of ODEs with proper parameters) to predict the amount of GenSnipers we can get in a given time(the zero point for time is set as the time when IPTG is added in). This model can inform us whether our system is efficient in industry production. | In this part, we will use mathematical methods(by solve a set of ODEs with proper parameters) to predict the amount of GenSnipers we can get in a given time(the zero point for time is set as the time when IPTG is added in). This model can inform us whether our system is efficient in industry production. | ||
| - | == | + | |
| + | ==Bottom-Up Strategy== | ||
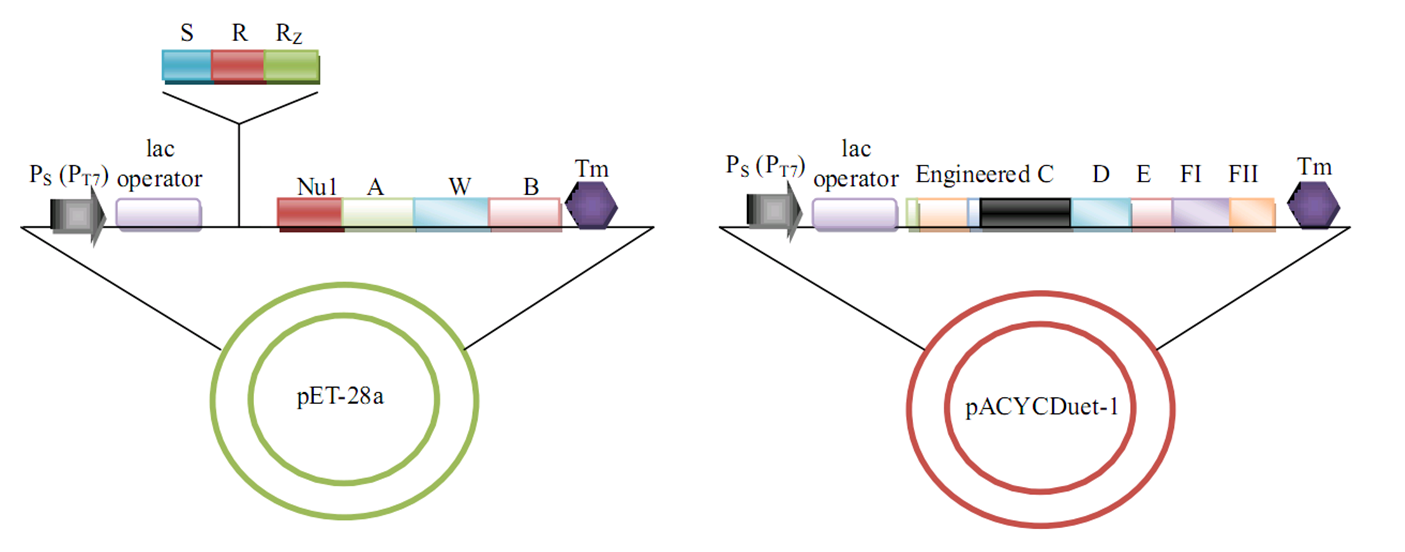
[[Image:BU.png|800px|center|thumb|Bottom-Up approach]] | [[Image:BU.png|800px|center|thumb|Bottom-Up approach]] | ||
| - | === | + | |
| + | ===Diagram of dynamic process=== | ||
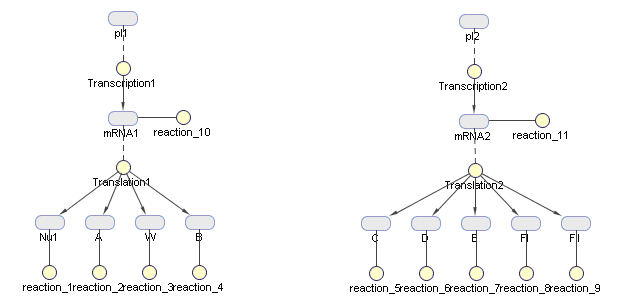
This figure is generated by Sim biology in matlab as a diagram of production modeling in phage vectors production | This figure is generated by Sim biology in matlab as a diagram of production modeling in phage vectors production | ||
[[Image:diagram phage.jpg|center|600px|thumb|Production modeling in vector production]] | [[Image:diagram phage.jpg|center|600px|thumb|Production modeling in vector production]] | ||
In a simple model, the assembly of proteins are considered negligible compared with the time of transcription and translation of these proteins precursors. The amount of product is the smaller one of the amount of package proteins of phage and the amount of targeted DNA ( to be packaged by phage) per bacterial cell. | In a simple model, the assembly of proteins are considered negligible compared with the time of transcription and translation of these proteins precursors. The amount of product is the smaller one of the amount of package proteins of phage and the amount of targeted DNA ( to be packaged by phage) per bacterial cell. | ||
| - | === | + | ===Results=== |
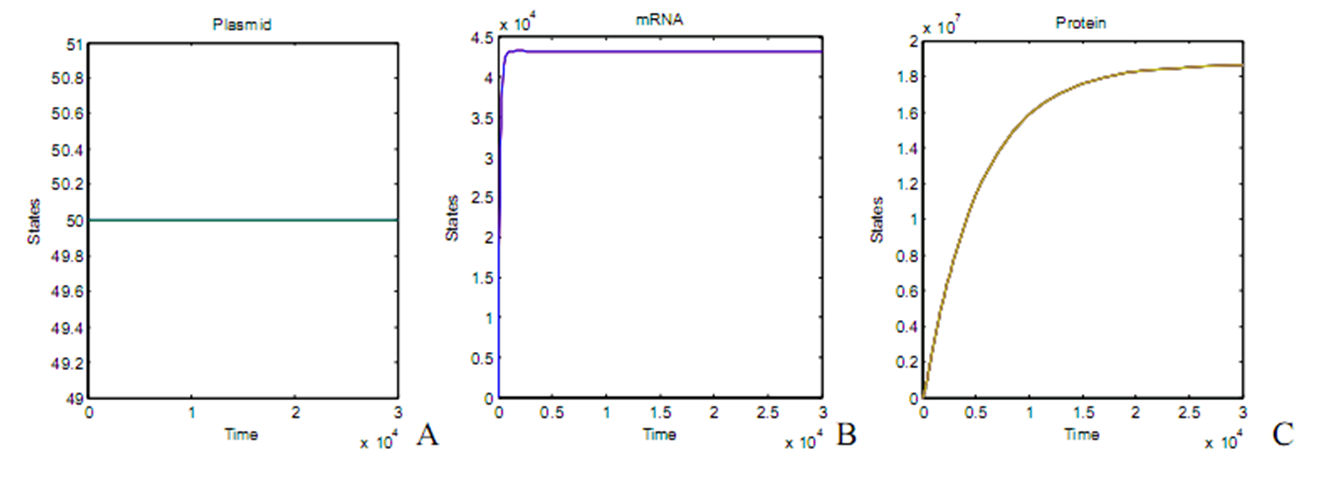
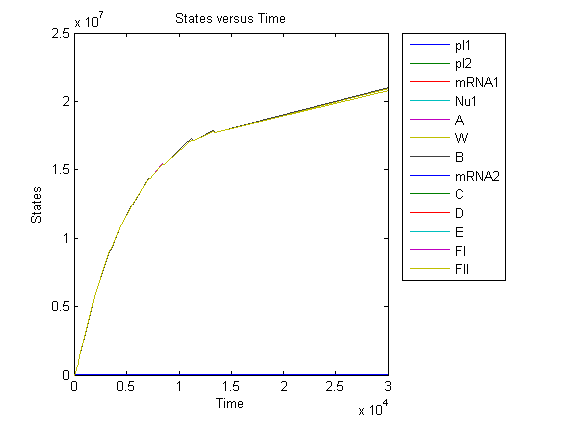
[[Image:result1111.jpg|600px|center]] | [[Image:result1111.jpg|600px|center]] | ||
Beginning with 50 plasmid per cell, the protein amount comes into equilibrium between synthesis and degradation about 10 hours later. The steady state amount of package proteins can reach the amount of about 10 in the orders of 7, given the parameters as showed in the figure, which means it is enough for packaging target DNAs. | Beginning with 50 plasmid per cell, the protein amount comes into equilibrium between synthesis and degradation about 10 hours later. The steady state amount of package proteins can reach the amount of about 10 in the orders of 7, given the parameters as showed in the figure, which means it is enough for packaging target DNAs. | ||
| Line 25: | Line 30: | ||
[[Image:result1112.jpg|600px|center]] | [[Image:result1112.jpg|600px|center]] | ||
| - | === | + | ===Paratmeters=== |
Protein tranaslation rate: r_Translation 0.0833/second | Protein tranaslation rate: r_Translation 0.0833/second | ||
| Line 34: | Line 39: | ||
mRNA degradation rate: d_mRNA 0.00578/second | mRNA degradation rate: d_mRNA 0.00578/second | ||
| - | == | + | ==Top-Down Strategy== |
| + | [[Image:TD2132.png|800px|center|thumb|Top-Down approach]] | ||
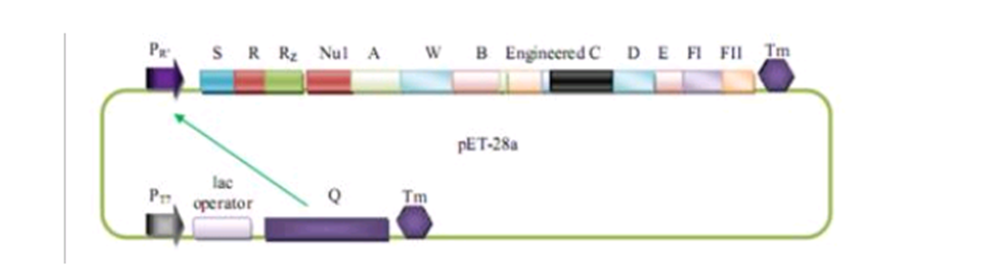
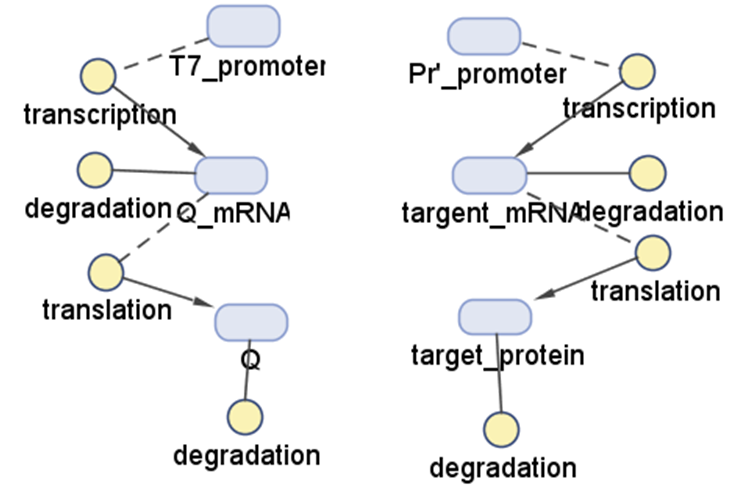
Q protein is a native λ phage protein which regulates λ phage's late gene transcription. In our system, Q protein is transcripted under regulation of T7 promomter then regulates the transcription of λ phage package protein through its anti-termination property to R'promoter. In this modeling section, we'll predict how much package proteins we will get using Pr'-Q regulatioin system. | Q protein is a native λ phage protein which regulates λ phage's late gene transcription. In our system, Q protein is transcripted under regulation of T7 promomter then regulates the transcription of λ phage package protein through its anti-termination property to R'promoter. In this modeling section, we'll predict how much package proteins we will get using Pr'-Q regulatioin system. | ||
| - | [[Image:Q.jpg| | + | [[Image:Q.jpg|500px|center|thumb|Top-Down model outline]] |
| - | === | + | ===Reaction kinetics=== |
| + | |||
the production of Q protein | the production of Q protein | ||
[[Image:Q_protein1.gif]] | [[Image:Q_protein1.gif]] | ||
| Line 53: | Line 60: | ||
As indicated in Ref(13), the Hill coefficient fitted by experiment is 6.4-3.3~6.4+3.3, which means that Q protein is very efficient in antitermination. As we don't know exactly the other parameters in this function, so we can not mimic Q protein's regulation effect for R' promoter dynamically, instead we suppose when 90% percent of Q protein have been reached, the R' promoter is fully activated | As indicated in Ref(13), the Hill coefficient fitted by experiment is 6.4-3.3~6.4+3.3, which means that Q protein is very efficient in antitermination. As we don't know exactly the other parameters in this function, so we can not mimic Q protein's regulation effect for R' promoter dynamically, instead we suppose when 90% percent of Q protein have been reached, the R' promoter is fully activated | ||
| - | === | + | ===Parameters=== |
T7 promoter transcription rate '''k'''<sub>t7</sub> = 5 molecule/second | T7 promoter transcription rate '''k'''<sub>t7</sub> = 5 molecule/second | ||
| Line 66: | Line 73: | ||
| - | === | + | ===Results=== |
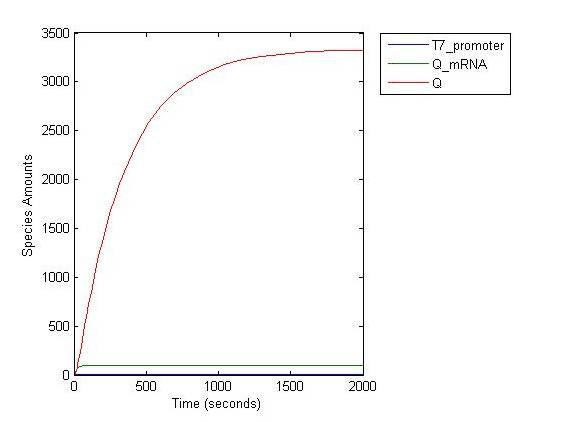
This is the Q protein level changing with time. When time= 1000s , 90% Q protein versus maximum amount haven been produced. At this time point, we assume that R' promoter is fully activated. | This is the Q protein level changing with time. When time= 1000s , 90% Q protein versus maximum amount haven been produced. At this time point, we assume that R' promoter is fully activated. | ||
[[Image:Q-result1.jpg|center]] | [[Image:Q-result1.jpg|center]] | ||
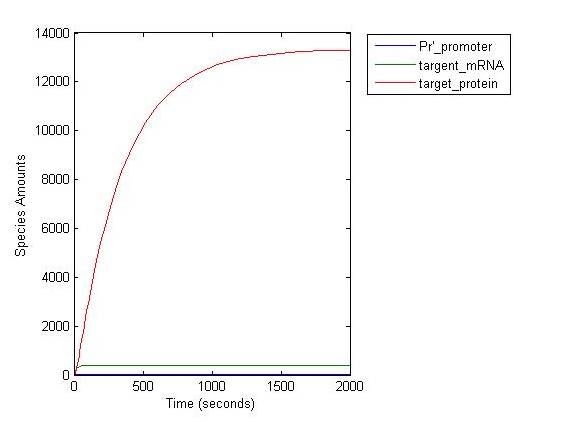
| Line 77: | Line 84: | ||
At the steady state, the concentration of the structral proteins reach at 1.3×10^4. Assuming that there are 420 D or E major structral protein on the surface of one GenSniper viroin, the packaged viroin number is 3.0×10^1 per cell, which is near to the wildtype bacteriophage lambda packaging. When the density of the cells is 10 in the orders of 9, the titer of synthetic GenSniper will be 3.0×10^10 Therapeutic DNA/ml, which also meets the standard for industrialization. | At the steady state, the concentration of the structral proteins reach at 1.3×10^4. Assuming that there are 420 D or E major structral protein on the surface of one GenSniper viroin, the packaged viroin number is 3.0×10^1 per cell, which is near to the wildtype bacteriophage lambda packaging. When the density of the cells is 10 in the orders of 9, the titer of synthetic GenSniper will be 3.0×10^10 Therapeutic DNA/ml, which also meets the standard for industrialization. | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
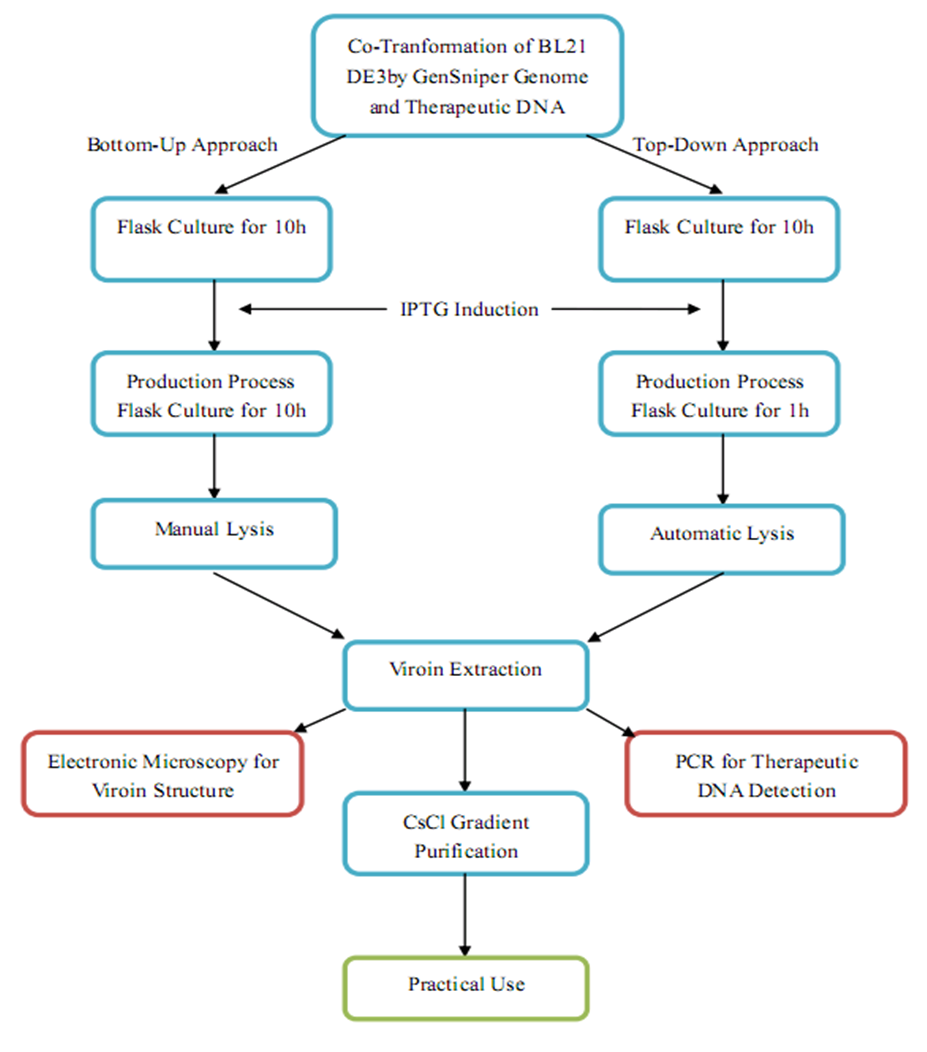
| + | Coupled with our wetlab observation, we outlined a production procedure regarding both Bottome-Up and Top-Down GenSniper Genome. | ||
| + | |||
| + | [[Image:productionap.jpg|center|thumb|800px|Production Scheme of GenSniper]] | ||
=Plasmid GenSniper Production Modeling= | =Plasmid GenSniper Production Modeling= | ||
| Line 114: | Line 129: | ||
Gibbs free energy. We give all the three unbinding rate of cI dimer from three kinds of binding sites to find out the best site we want to use in the experiment. | Gibbs free energy. We give all the three unbinding rate of cI dimer from three kinds of binding sites to find out the best site we want to use in the experiment. | ||
| - | == | + | ==Replication of plasmid== |
plasmid replication is a complex process, here we use decay rate replication as a simplified model to mimic negative regulation. Three general classes of regulatory mechanisms have been studied in depth, namely those that involve directly repeated sequences (iterons), those that use only antisense RNAs and those that use a mechanism involving an antisense RNA in combination with a protein. For more detailed info about ''Control of plasmid replication and copy-number'': [http://www.sci.sdsu.edu/~smaloy/MicrobialGenetics/topics/plasmids/plasmid-repln.html] . An detailed mathematical model is described in (12) | plasmid replication is a complex process, here we use decay rate replication as a simplified model to mimic negative regulation. Three general classes of regulatory mechanisms have been studied in depth, namely those that involve directly repeated sequences (iterons), those that use only antisense RNAs and those that use a mechanism involving an antisense RNA in combination with a protein. For more detailed info about ''Control of plasmid replication and copy-number'': [http://www.sci.sdsu.edu/~smaloy/MicrobialGenetics/topics/plasmids/plasmid-repln.html] . An detailed mathematical model is described in (12) | ||
<br> | <br> | ||
| Line 128: | Line 143: | ||
| - | *[[Image:plasmid_ori.gif]] | + | *[[Image:plasmid_ori.gif|600px]] |
<br> | <br> | ||
| Line 165: | Line 180: | ||
*[[Image:Eqn7.gif]] | *[[Image:Eqn7.gif]] | ||
| - | == | + | == Numerical simulation == |
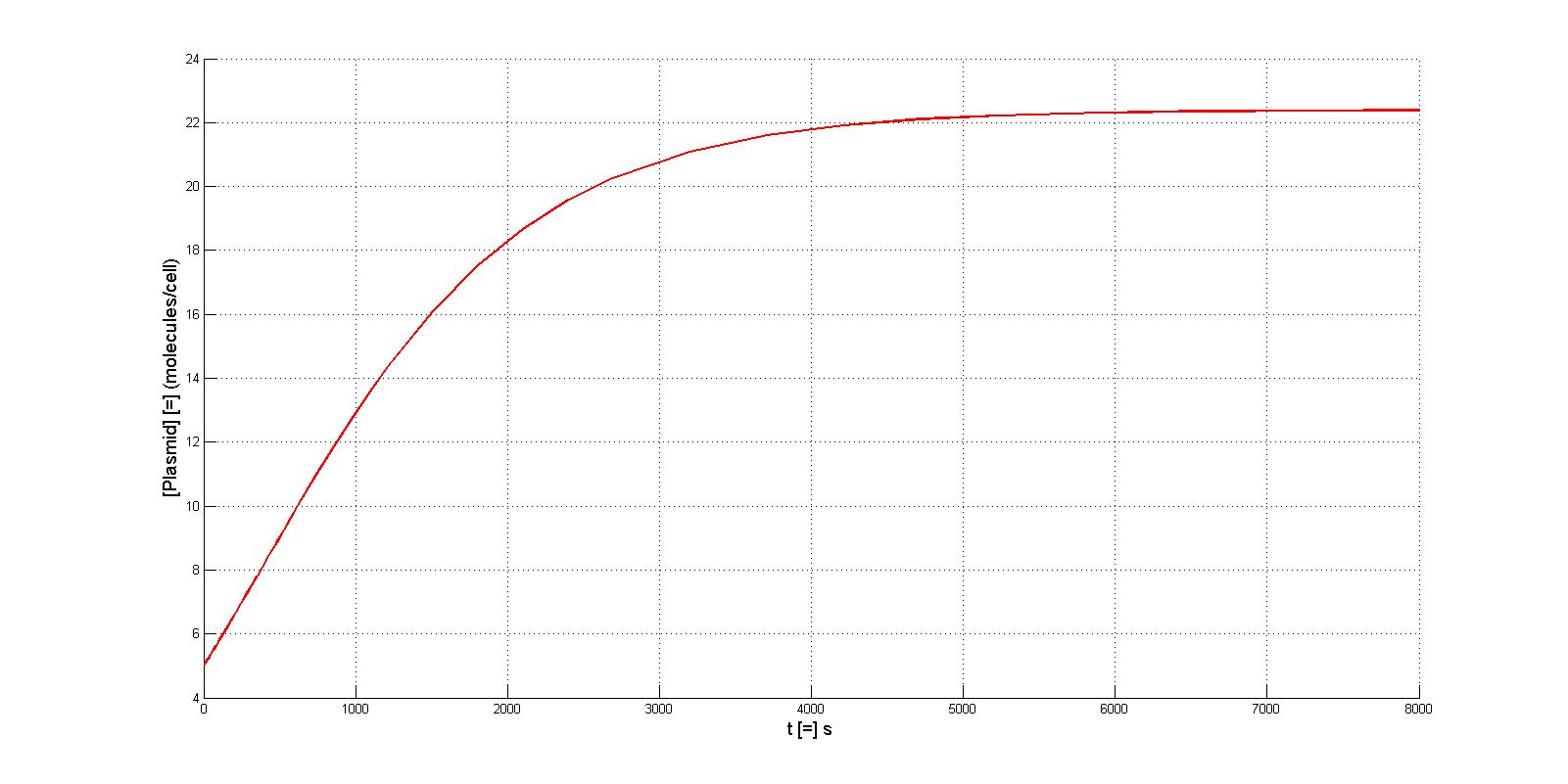
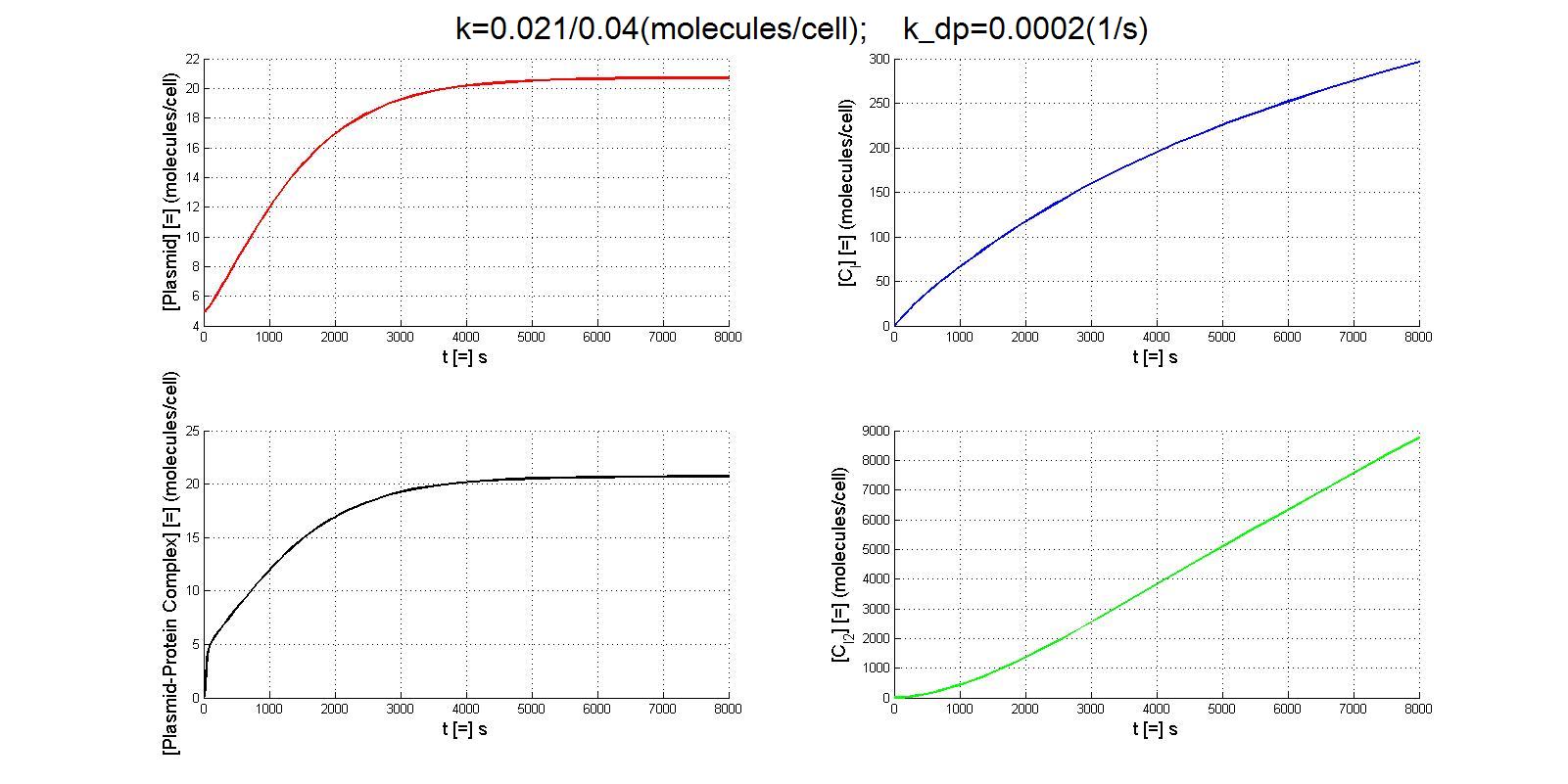
| - | According to the previous analysis, the parameters which we can manipulate in the experiment is protein binding affinity(by choosing different binding sites) and protein degradation rate(by adding degradation tag). Here is the simulation results with | + | According to the previous analysis, the parameters which we can manipulate in the experiment is protein binding affinity(by choosing different binding sites) and protein degradation rate(by adding degradation tag). Here is the simulation results with a parameter of k and kdp |
| - | *[[Image:pp_result1.gif | + | *[[Image:pp_result1.gif|center|900px]] |
| - | + | ||
| - | + | ||
== Parameter dependency == | == Parameter dependency == | ||
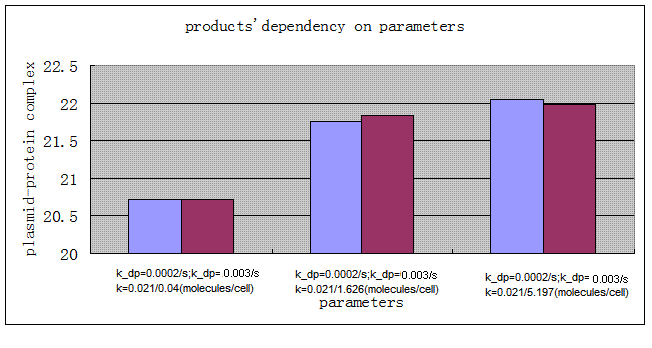
| - | final product plasmid_protein complex we got by using different protein_binding sites: | + | final product plasmid_protein complex we got by using different protein_binding sites and protein degradation rates: |
| - | *[[Image:pp_3.gif]] | + | *[[Image:pp_3.gif|center]] |
| - | + | We can see that higher binding affinity favors the final production of plasmid vectors(plasmid-protein complex). And the influence of protein degration rate is a little ambiguous: fast degradation may either favor or adverse to the production of plasmid vectors (compare when k= 0.021/1.626 and 0.021/5.197)according to the protein binding affinity. After all, both of the influence of protein binding affinity and protein degradation rate is minor and the binding of protein doesn't change the nature replication process much. | |
| - | + | == Conclusions == | |
| - | + | According to the analysis in the modeling section, we find that different parameters such as protein binding affinity and protein degradation rate won't affect the amount of the target protein significantly in the range of experimental values. However, it does give us the clue that protein with higher affinity and moderate degradation rate is suitable for large scale industry production. | |
| - | + | ||
| - | + | ||
| - | + | ||
=Molecular Docking for Targeted Biobrick= | =Molecular Docking for Targeted Biobrick= | ||
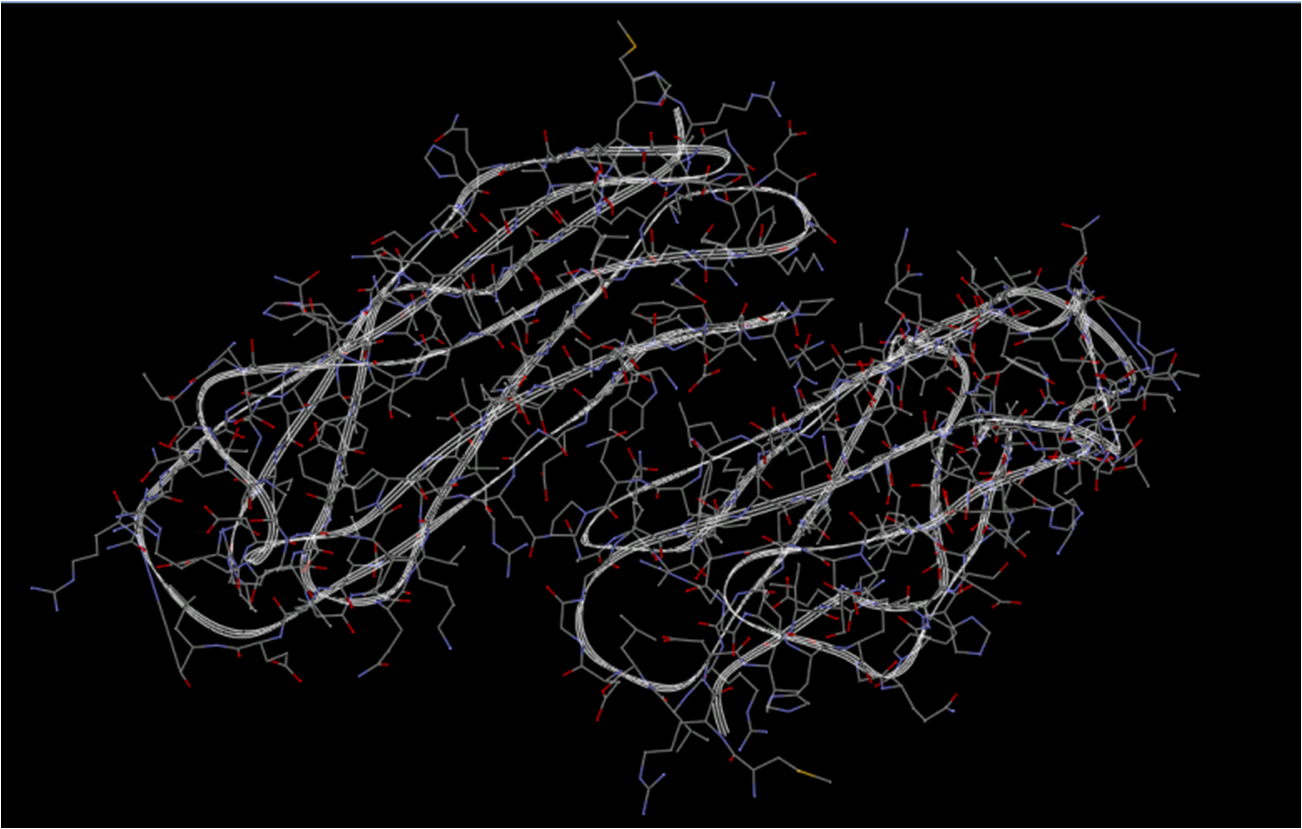
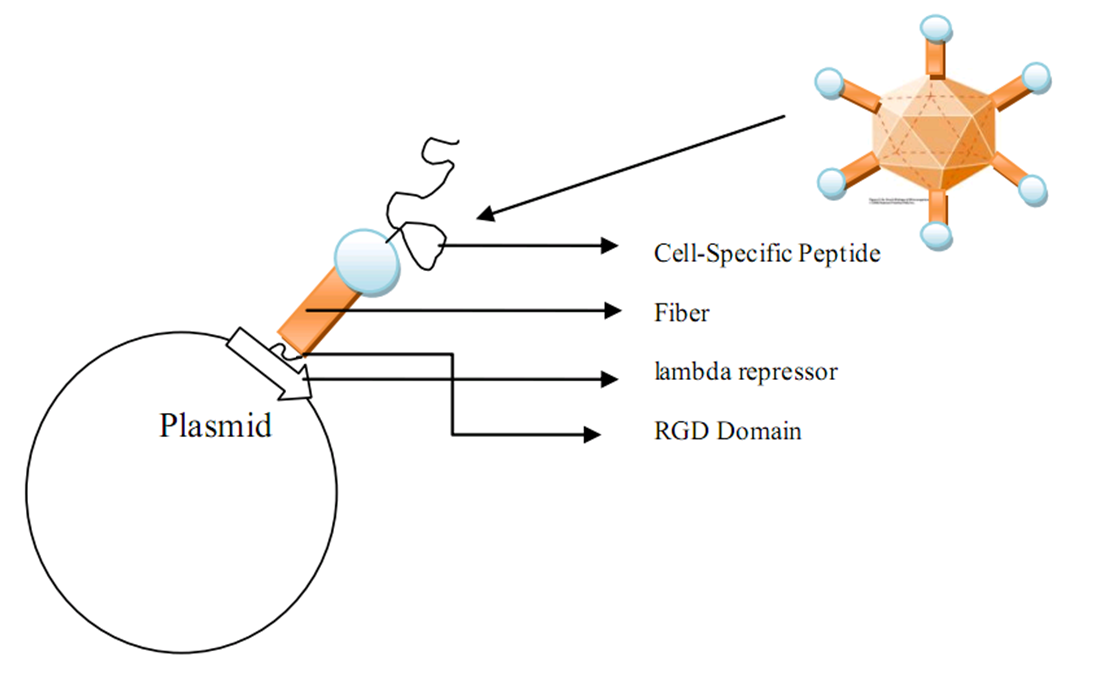
| - | [[Image:receptor.jpg|600px|center|thumb| | + | [[Image:receptor.jpg|600px|center|thumb|Integrin structure]] |
| - | [[Image:fiber.jpg|700px|center|thumb| | + | [[Image:fiber.jpg|700px|center|thumb|Fiber structure]] |
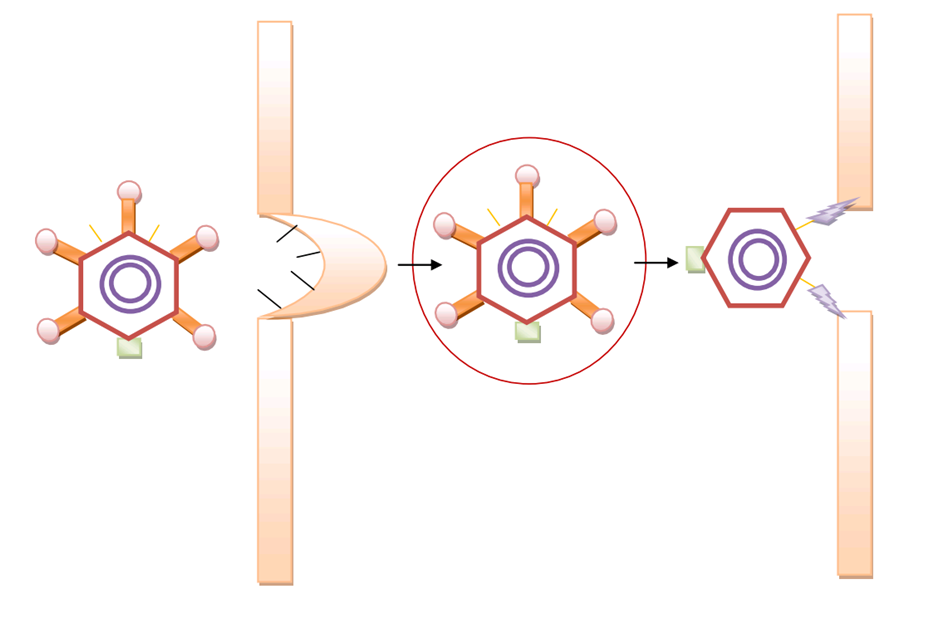
We used ZDOCK software to conduct the molecular docking day experiment. In the generated docking graph, the ligand (we used the crystalized fiber protein to substitute the targeted biobrick) and the receptor (integrin) are both in a stretched confirmation in which the spacial clash is relatively small. Moreover, in the fiber confirmation, the positions for both RGD domain and cell-specific peptide are exposed to the surface to faciliate the specific interaction with the cell specific receptor and integrin. | We used ZDOCK software to conduct the molecular docking day experiment. In the generated docking graph, the ligand (we used the crystalized fiber protein to substitute the targeted biobrick) and the receptor (integrin) are both in a stretched confirmation in which the spacial clash is relatively small. Moreover, in the fiber confirmation, the positions for both RGD domain and cell-specific peptide are exposed to the surface to faciliate the specific interaction with the cell specific receptor and integrin. | ||
| + | =Outline of SynGenome Based GenSniper Intercellular Trafficking Modeling= | ||
| + | Since the intercellular process of GenSniper is quite complicated, we just outline a model for this process according to the established modeling on adenovirus. | ||
| + | |||
| + | [[Image:interce.jpg|900px|center|thumb|Intercellular Trafficking Mechanism of GenSniper]] | ||
| + | |||
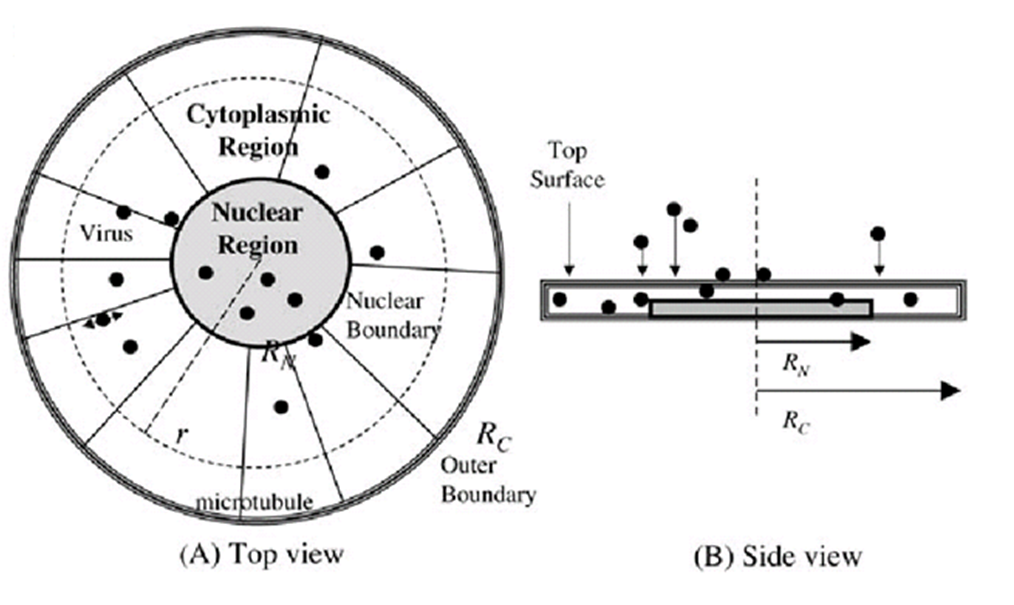
| + | According to the adenovirus trafficking model, GenSniper enters the cell from the top. In the nuclear region (r , RN), it can only diffuse. In the cytoplasmic region (RN , r , RC), GenSniper intermittently switches between diffusion and motor-assisted transport. As an approximation, we can assume uniform concentration of endosomal and cytosolic GenSnipers along the vertical direction. | ||
| + | |||
| + | [[Image:interce2.jpg|600px|center|thumb|Schematic representation of a model cell for GenSniper trafficking]] | ||
| + | |||
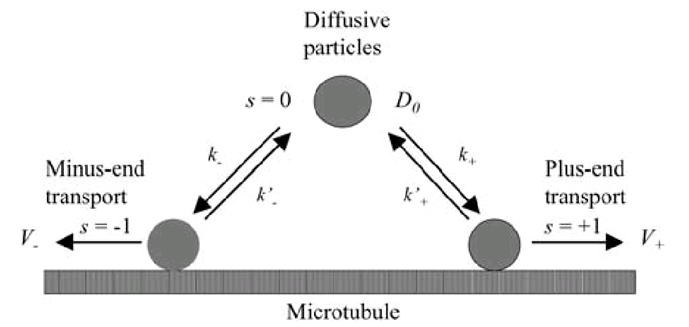
| + | [[Image:interce3.jpg|600px|center|thumb|Transition map of GenSniper trafficking]] | ||
| + | |||
| + | As for this model, it is of note that the nuclear localization signal has not been incorporated into GenSniper. Thus, the trafficking process is hard to predict. Since naked plasmid can also work even without such signals and protein capsid, the design of GenSniper without such signals seems to be OK. In addition, as for a newly proposed approach, this design is recommended for a well-defined wetlab system and a simplified modeling. As for practical use, the NLS is highly recommended to be added by phage display on E of lambda head. | ||
| + | [[Image:interce4.jpg|700px|center|thumb|GenSniper Intracellular trafficking model can be ensured by display NLS sequence on E]] | ||
=References= | =References= | ||
| Line 220: | Line 243: | ||
[13] Kobiler, O., et al., ''Quantitative kinetic analysis of the bacteriophage lambda genetic network.'' Proc Natl Acad Sci U S A, 2005. 102(12): p. 4470-5. | [13] Kobiler, O., et al., ''Quantitative kinetic analysis of the bacteriophage lambda genetic network.'' Proc Natl Acad Sci U S A, 2005. 102(12): p. 4470-5. | ||
| + | [14] Anh-Tuan Dinh, Theo Theofanous, and Samir Mitragotri. A Model for Intracellular Trafficking of Adenoviral Vectors. Biophysical Journal, 2005. 89: 1574–1588. | ||
{{Tsinghua/Pic}} | {{Tsinghua/Pic}} | ||
Latest revision as of 23:49, 21 October 2009
| |||||||||||
 "
"