Team:TzuChiU Formosa/Result
From 2009.igem.org
(→Result) |
|||
| (3 intermediate revisions not shown) | |||
| Line 12: | Line 12: | ||
==Result== | ==Result== | ||
| - | <gallery> | + | 1. |
| + | <gallery widths=250px heights=250px> | ||
Image:Result_f1a.jpg|a | Image:Result_f1a.jpg|a | ||
Image:Result_f1b.jpg|b | Image:Result_f1b.jpg|b | ||
Image:Result_f1c.jpg|c | Image:Result_f1c.jpg|c | ||
</gallery> | </gallery> | ||
| - | '''Figure.1(a.)pSB1A3 cloning PCR gel image. start with left to right part:(1)100 bps marker,(2) | + | '''Figure.1(a.)pSB1A3 cloning PCR gel image. start with left to right part:(1.)100 bps marker,(2.)OmpC promoter,(3.)OmpC promoter,(4.)Negative control。(b.)the three red circles are single colony that have been transformed OmpC promoter containing plasmid。(c.)OmpC plasmid gel image。start with left to right part:(1.)100 bps marker,(2.)pSB1A3 plasmid,(3.)pSB1A3 plasmid.''' |
| - | Our PCR show that pSBA1B3 contained OmpC promoter, and the size was about 108 base pairs. From Fig.1-a Show that band was placed at 108 base pairs, which confirmed that pSB1A3 contained OmpC promoter. From Fig1-b we can see the growing of single colony. | + | Our PCR show that pSBA1B3 contained OmpC promoter, and the size was about 108 base pairs. From Fig.1-a Show that band was placed at 108 base pairs, which confirmed that pSB1A3 contained OmpC promoter. From Fig1-b we can see the growing of single colony. Fig1-c shows plasmid was at 2.75 Kb, so it was confirmed that pSB1A3 had been successfully transformed into DH5-α competent cell. |
| - | <gallery> | + | 2. |
| + | <gallery widths=250px heights=250px> | ||
Image:Result_f2a.jpg|a | Image:Result_f2a.jpg|a | ||
Image:Result_f2b.jpg|b | Image:Result_f2b.jpg|b | ||
Image:Result_f2c.jpg|c | Image:Result_f2c.jpg|c | ||
</gallery> | </gallery> | ||
| - | '''Figure.2(a.)Aequorin-GFP sequence after purify gel image。Left:(1)100 bps marker,(2)Aequorin-GFP | + | '''Figure.2(a.)Aequorin-GFP sequence after purify gel image。Left:(1.)100 bps marker, (2.)Aequorin-GFP, (b.)After TA cloning transformation into DH5-α competent cell and run the blue white selection.(c.)Aequorin-GFP cloning PCR gel image. Start with left to right:(1.)100 bps marker,(2.3.)Aequorin-GFP. |
''' | ''' | ||
| - | The purify and concentration are | + | The purify and concentration are unknown,so we run the purification to increase. Fig.2-a Show that template was placed at 2000 baes pairs. And took Aequorin-GFP to the TA cloning, let plasmisd could ligate to pGEM-T-easy plasmid. After transformation into DH5-α competent cell, then spirting the plate contain Ampicilin, 100ul IPTG, 50 ul X-gal. Fig2-b: Show that the blue white selection was white single colony. Picked two closing size single colony to amplify, and run colony PCR show that Aequorin-GFP size was about 2000 baes pairs. Fig.2-c Show that the sequence was at 2000 base pairs. |
| - | <gallery> | + | |
| - | Image:Result_f3a.jpg| | + | 3. |
| - | Image:Result_f3b.JPG| | + | <gallery widths=250px heights=250px> |
| + | Image:Result_f3a.jpg|3a | ||
| + | Image:Result_f3b.JPG|3b | ||
</gallery> | </gallery> | ||
| - | '''Figure.3(a.)pSB1A3 digestion gel image.Start with left to right:(1) | + | '''Figure.3(a.)pSB1A3 digestion gel image. Start with left to right:(1.)100bs marker, (2.)Uncut pSB1A3, (2.)Uncut pSB1A3, (3.)Cut pSB1A3; (b.) (1.)100bps marker, (2.)Dephosphorelation of Pst1 cutting site in pSB1A3. |
''' | ''' | ||
| - | When we confirm the PSB1A3 contain OmpC promoter. And we amplify it, used Pst1 to cutting off the sequences and run the gel electrophoresis. Due to the cutting form plasmid will change circular to linear form caused molecular weight increasing so the gel electrophoresis speed will slow down and stop at the upper strata show at the Fig.3-a. | + | When we confirm the PSB1A3 that contain OmpC promoter. And we amplify it, used Pst1 to cutting off the sequences and run the gel electrophoresis. Due to the cutting form plasmid will change circular to linear form caused molecular weight increasing so the gel electrophoresis speed will slow down and stop at the upper strata show at the Fig.3-a. |
In order to improve the successful rate of putting insert into the vector we treated, Pst1 digested pSB1A3 with calf Intestinal Alkaline Phosphatase (CIP) to remove the phosphate group thus reducing the chance of self ligation | In order to improve the successful rate of putting insert into the vector we treated, Pst1 digested pSB1A3 with calf Intestinal Alkaline Phosphatase (CIP) to remove the phosphate group thus reducing the chance of self ligation | ||
Latest revision as of 03:27, 22 October 2009
Result
1.
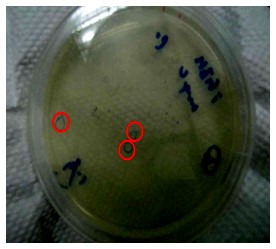
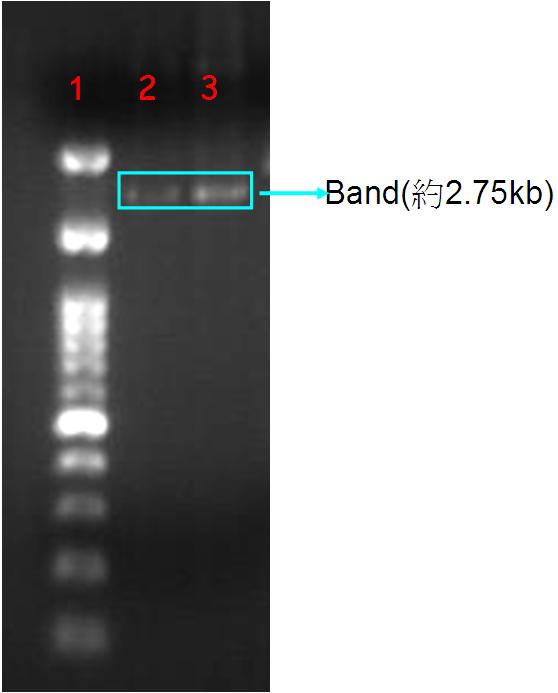
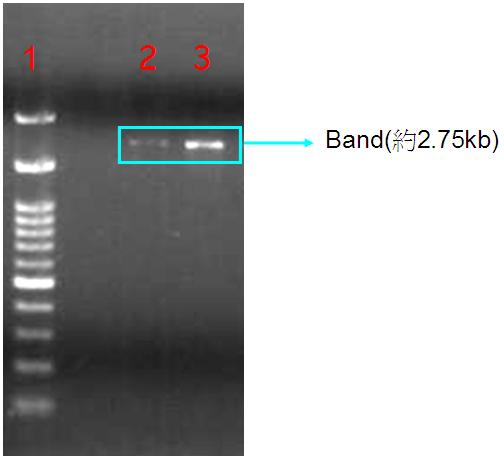
Figure.1(a.)pSB1A3 cloning PCR gel image. start with left to right part:(1.)100 bps marker,(2.)OmpC promoter,(3.)OmpC promoter,(4.)Negative control。(b.)the three red circles are single colony that have been transformed OmpC promoter containing plasmid。(c.)OmpC plasmid gel image。start with left to right part:(1.)100 bps marker,(2.)pSB1A3 plasmid,(3.)pSB1A3 plasmid.
Our PCR show that pSBA1B3 contained OmpC promoter, and the size was about 108 base pairs. From Fig.1-a Show that band was placed at 108 base pairs, which confirmed that pSB1A3 contained OmpC promoter. From Fig1-b we can see the growing of single colony. Fig1-c shows plasmid was at 2.75 Kb, so it was confirmed that pSB1A3 had been successfully transformed into DH5-α competent cell.
2.
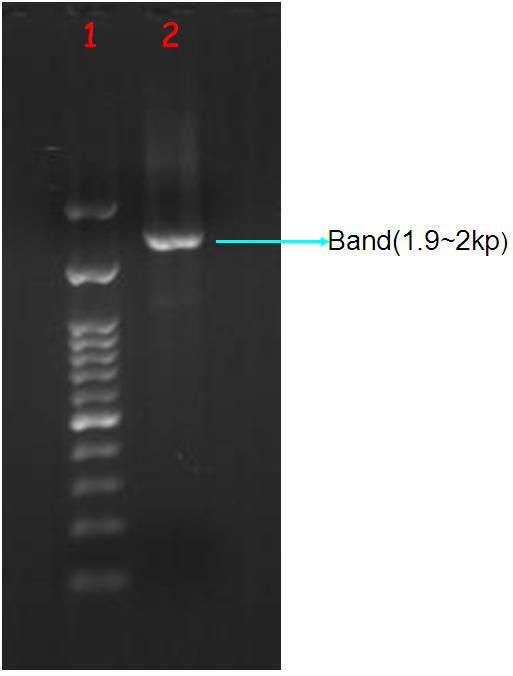
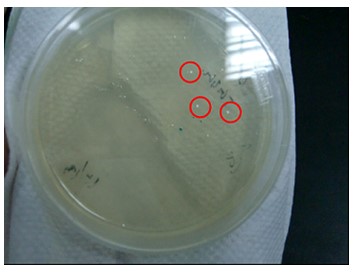
Figure.2(a.)Aequorin-GFP sequence after purify gel image。Left:(1.)100 bps marker, (2.)Aequorin-GFP, (b.)After TA cloning transformation into DH5-α competent cell and run the blue white selection.(c.)Aequorin-GFP cloning PCR gel image. Start with left to right:(1.)100 bps marker,(2.3.)Aequorin-GFP.
The purify and concentration are unknown,so we run the purification to increase. Fig.2-a Show that template was placed at 2000 baes pairs. And took Aequorin-GFP to the TA cloning, let plasmisd could ligate to pGEM-T-easy plasmid. After transformation into DH5-α competent cell, then spirting the plate contain Ampicilin, 100ul IPTG, 50 ul X-gal. Fig2-b: Show that the blue white selection was white single colony. Picked two closing size single colony to amplify, and run colony PCR show that Aequorin-GFP size was about 2000 baes pairs. Fig.2-c Show that the sequence was at 2000 base pairs.
3.
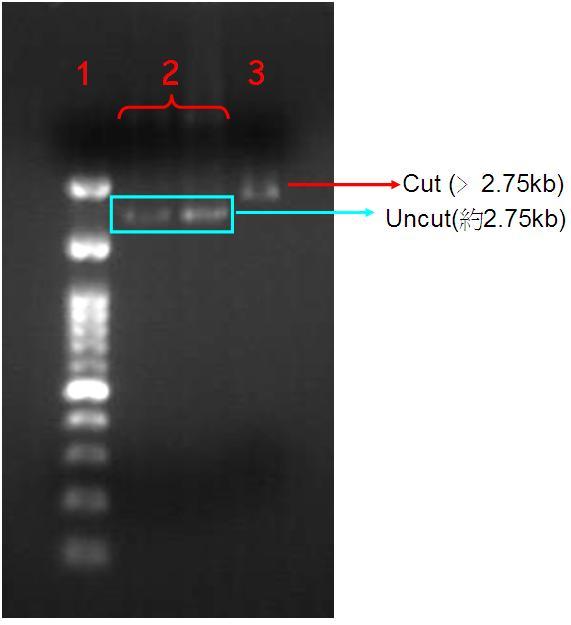
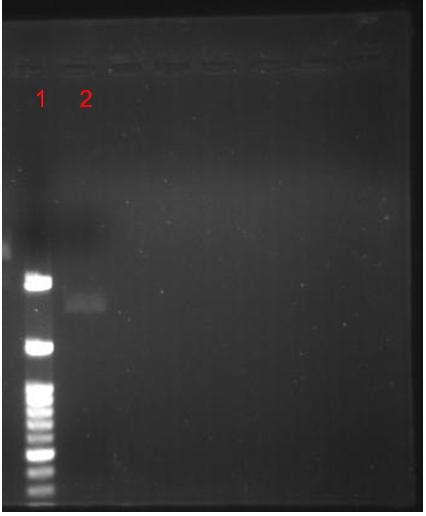
Figure.3(a.)pSB1A3 digestion gel image. Start with left to right:(1.)100bs marker, (2.)Uncut pSB1A3, (2.)Uncut pSB1A3, (3.)Cut pSB1A3; (b.) (1.)100bps marker, (2.)Dephosphorelation of Pst1 cutting site in pSB1A3.
When we confirm the PSB1A3 that contain OmpC promoter. And we amplify it, used Pst1 to cutting off the sequences and run the gel electrophoresis. Due to the cutting form plasmid will change circular to linear form caused molecular weight increasing so the gel electrophoresis speed will slow down and stop at the upper strata show at the Fig.3-a.
In order to improve the successful rate of putting insert into the vector we treated, Pst1 digested pSB1A3 with calf Intestinal Alkaline Phosphatase (CIP) to remove the phosphate group thus reducing the chance of self ligation
 "
"