Team:MoWestern Davidson/project wetlab
From 2009.igem.org
Macampbell (Talk | contribs) (→Completed Constructs - Suppressor tRNAs and FSL-RFP Reporters) |
|||
| (15 intermediate revisions not shown) | |||
| Line 2: | Line 2: | ||
__TOC__ | __TOC__ | ||
== Completed tRNAs == | == Completed tRNAs == | ||
| - | We assembled the genes for our suppressor tRNAs by ligating single-stranded oligos with sticky BioBrick ends. Go [https://2009.igem.org/Team:MoWestern_Davidson/tRNA here] to see a full explanation, or [http://www.bio.davidson.edu/courses/Molbio/Protocols/anneal_oligos.html here] for the protocol. This generated double-stranded DNA ready to ligate into a standard vector. All tRNAs | + | We assembled the genes for our suppressor tRNAs by ligating single-stranded oligos with sticky BioBrick ends. Go [https://2009.igem.org/Team:MoWestern_Davidson/tRNA here] to see a full explanation, or [http://www.bio.davidson.edu/courses/Molbio/Protocols/anneal_oligos.html here] for the protocol. This generated double-stranded DNA ready to ligate into a standard vector. All tRNAs deliver the same amino acid, serine. The expected size of our tRNAs with BioBrick ends is 184 bp. We verified that we assembled the expected tRNAs by checking the insert size of our ligation (seen below), and by sequencing our tRNA plasmids. |
<br> | <br> | ||
| - | <center>[[Image:tRNAgel.png|none|thumb|500px|2.0% agarose gel of plasmids containing | + | <center>[[Image:tRNAgel.png|none|thumb|500px|2.0% agarose gel of plasmids containing the CCAUC-9 tRNA inserts (184 bp), digested with EcoRI and PstI. The insert band size is just below 200 bp, as expected. |
]]</center><br> | ]]</center><br> | ||
| Line 51: | Line 51: | ||
<br/><br/> | <br/><br/> | ||
| - | == Completed Reporters With Frameshift Suppressor Leaders == | + | == Completed Reporters With Frameshift Suppressor Leaders (FSLs) == |
In order to engineer logical clauses directly upstream of our selected reporter genes, we altered the 5’ end of the reporter using PCR to add frameshift suppressor leaders (FSLs). FSLs were added after the reporter start codon ATG so that the suppressor tRNAs are required to correctly translate the entire reporter. We began with a modular 1-SAT approach - adding a single 5-nt codon as the FSL to the 5' end of the reporter. | In order to engineer logical clauses directly upstream of our selected reporter genes, we altered the 5’ end of the reporter using PCR to add frameshift suppressor leaders (FSLs). FSLs were added after the reporter start codon ATG so that the suppressor tRNAs are required to correctly translate the entire reporter. We began with a modular 1-SAT approach - adding a single 5-nt codon as the FSL to the 5' end of the reporter. | ||
| Line 63: | Line 63: | ||
!width="250"|[[Image:FSLTet.png|center|225px]] | !width="250"|[[Image:FSLTet.png|center|225px]] | ||
|- | |- | ||
| - | | CUAGC || | + | | CUAGC || CUAGC || ----- |
|- | |- | ||
| - | | ----- || | + | | ----- || CCAUC || ----- |
|- | |- | ||
| ----- || CGGUC || CGGUC | | ----- || CGGUC || CGGUC | ||
| Line 74: | Line 74: | ||
|- | |- | ||
| ----- || CCCUC || ----- | | ----- || CCCUC || ----- | ||
| + | |- | ||
| + | | ----- || AGGAC || ----- | ||
| + | |- | ||
| + | | ----- || CUACC || ----- | ||
| + | |- | ||
| + | | ----- || CCAAU || ----- | ||
| + | |- | ||
| + | | ----- || CCACC || ----- | ||
| + | |- | ||
| + | | ----- || CUACU || ----- | ||
|- | |- | ||
|} | |} | ||
| Line 80: | Line 90: | ||
The PCR template for our FSL additions were reporter constructs with a promoter and RBS. For example, pLac-RBS-RFP was the template plasmid for the FSL-RFPs. This was helpful after PCR and ligation into the template vector, so that we could select potential FSL-RFPs by colour selection. | The PCR template for our FSL additions were reporter constructs with a promoter and RBS. For example, pLac-RBS-RFP was the template plasmid for the FSL-RFPs. This was helpful after PCR and ligation into the template vector, so that we could select potential FSL-RFPs by colour selection. | ||
| - | Below is a plate of colonies transformed with our potential FSL-RFPs. Red colonies clearly have the original construct since they still express RFP in the absence of the appropriate suppressor tRNA. White colonies potentially have our desired construct. | + | Below is a plate of colonies transformed with our potential FSL-RFPs. Red colonies clearly have the original construct since they still express RFP in the absence of the appropriate suppressor tRNA. White colonies potentially have our desired construct containing the FSL upstream of RFP. |
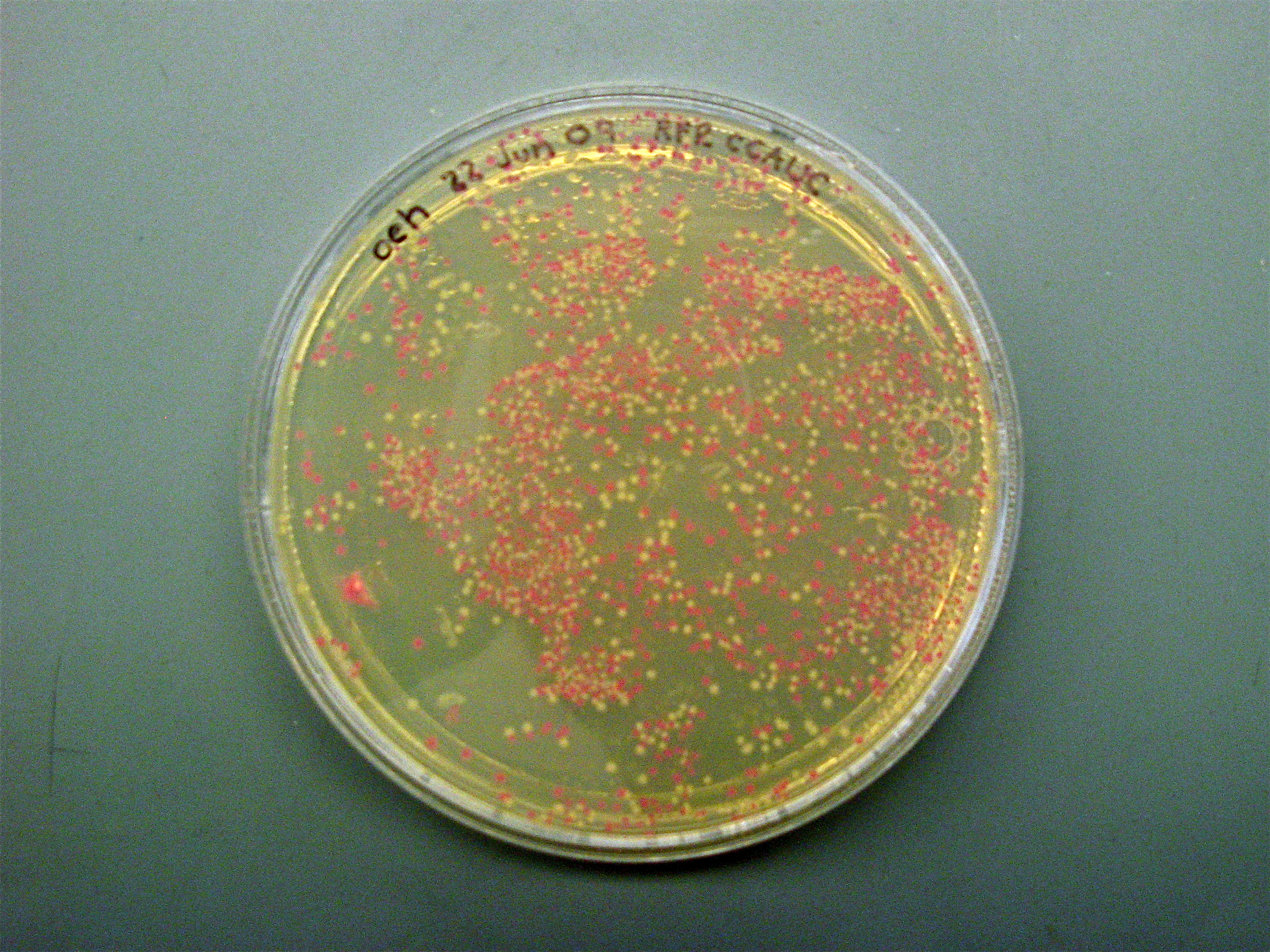
| - | <center>[[Image:RedWhite transform selection.jpg|none|thumb|500px|5-bp FSLs were added by PCR amplification to the 5' end of the RFP gene. FSL-RFP was ligated into a digested RFP vector and transformed. Red colonies indicate an incomplete digestion of the RFP vector, or indicate | + | <center>'''Red/White Selection of New Inserts'''<br> |
| + | [[Image:RedWhite transform selection.jpg|none|thumb|500px|5-bp FSLs were added by PCR amplification to the 5' end of the RFP gene. FSL-RFP was ligated into a digested RFP vector and transformed. Red colonies indicate an incomplete digestion of the RFP vector, or indicate that no FSL has been added to cause a frameshift. White colonies are selected because they no longer contain the RFP expression insert so we know the desired ligation worked.]]</center> | ||
<br/> | <br/> | ||
Selected colonies were sequenced for verification of the correct FSL addition. The next step is to ligate each tRNA with the corresponding FSL-reporter, so that both genes will be present and expressed within the cell. We are attempting two possible constructs for this: | Selected colonies were sequenced for verification of the correct FSL addition. The next step is to ligate each tRNA with the corresponding FSL-reporter, so that both genes will be present and expressed within the cell. We are attempting two possible constructs for this: | ||
| Line 91: | Line 102: | ||
<center>[[Image:Construct4.png|none|500px]]</center> | <center>[[Image:Construct4.png|none|500px]]</center> | ||
2.The tRNA directly downstream of the reporter. | 2.The tRNA directly downstream of the reporter. | ||
| - | <center>[[Image: | + | <center>[[Image:Construct3.png|none|400px|]]</center> |
<br/><br/> | <br/><br/> | ||
| Line 117: | Line 128: | ||
<br/><br/> | <br/><br/> | ||
| - | == | + | == Completed Constructs - Suppressor tRNAs and FSL-RFP Reporters == |
| - | + | Based on promoter control data, we decided to use pBad promoter upstream of suppressor tRNAs, and to use pLac promoter upstream of the RFP reporter with FSLs. The benefit of these promoter choices is constitutive expression with the possibility of further induction. We chose pLac for RFP expression because it caused the greatest expression in the control construct (promoter-RBS-RFP, no FSL). | |
| - | + | <br> | |
| + | <center>[[Image:Construct4.png|thumb|350px|none|Construct 1. See "Completed Reporters With Frameshift Suppressor Leaders (FSLs)" for the discussion of constructs.]]</center> | ||
| + | <br> | ||
| + | We completed successfully Construct 1: pLac-RBS-FSL-RFPs were ligated downstream of the corresponding pBad-tRNA (i.e. The tRNA upstream of pLac-RBS-CGGUC-RFP is the suppressor of the FSL CGGUC). This construct was transformed into E. coli, and transformation was verified by digestion and electrophoresis, and by sequencing isolated plasmid. Cells with the verified construct were grown in LB media with inducers 2.5% L-arabinose and 0.06% IPTG. Fluorescence readings were measured after 20 hours of incubation. The figure below shows the measurements, normalized to the negative control, of all FSL constructs that showed any fluorescence. We observed red fluorescence most notably in the construct containing the tRNA CCAUC 10-bp anticodon (below). | ||
| - | + | <br> | |
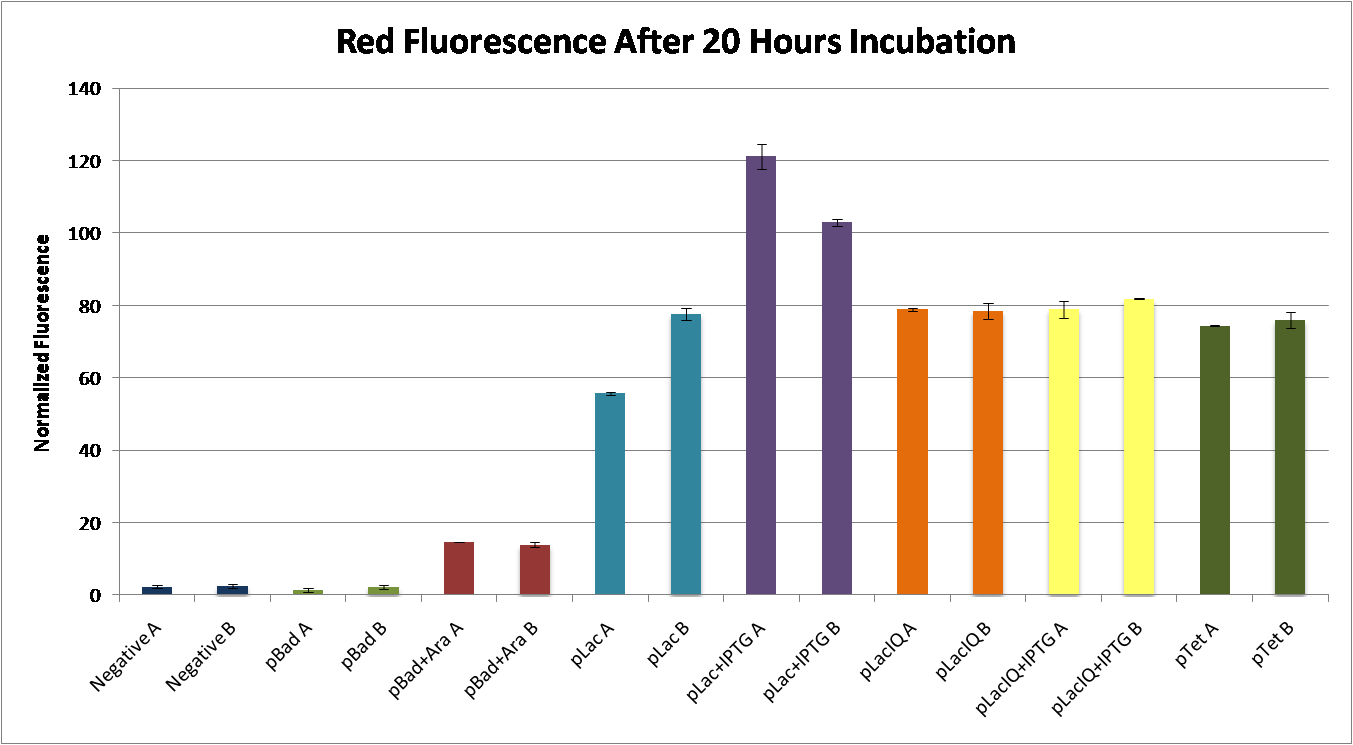
| - | + | <center>[[Image:TRNARFPtest.png|thumb|650px|none|Fluorescence readings were measured from 2 mL dense culture of each tRNA/FSL construct. Fluorescence values were normalized to the cell density, and divided by the negative control fluorescence. All values on the y-axis greater than 1 indicate greater fluorescence than the negative control (no RFP). Thin bars indicate no inducers were added. Medium bars indicate 0.06% IPTG added, and Thick bars indicate IPTG and 2.5% L-arabinose added.]]</center> | |
| - | < | + | |
| - | <center>[[Image: | + | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
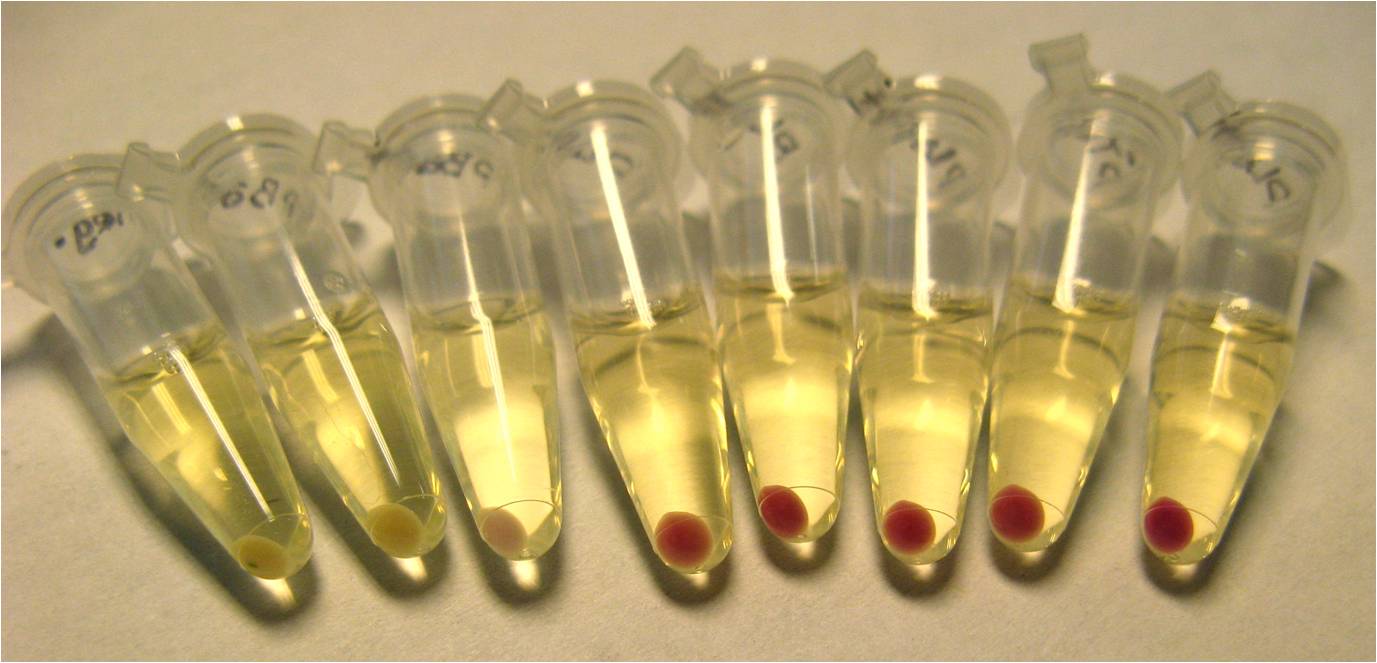
| + | The red fluorescence in CCAUC 10-bp construct was also visible in the pelleted cell culture (below). These data indicate a construct containing our suppressor tRNAs can successfully suppress their target 5-bp codon, allowing expression of RFP. Therefore, we have designed and built a mechanism to solve a very simple SAT problem in a biological construct. | ||
| + | <br> | ||
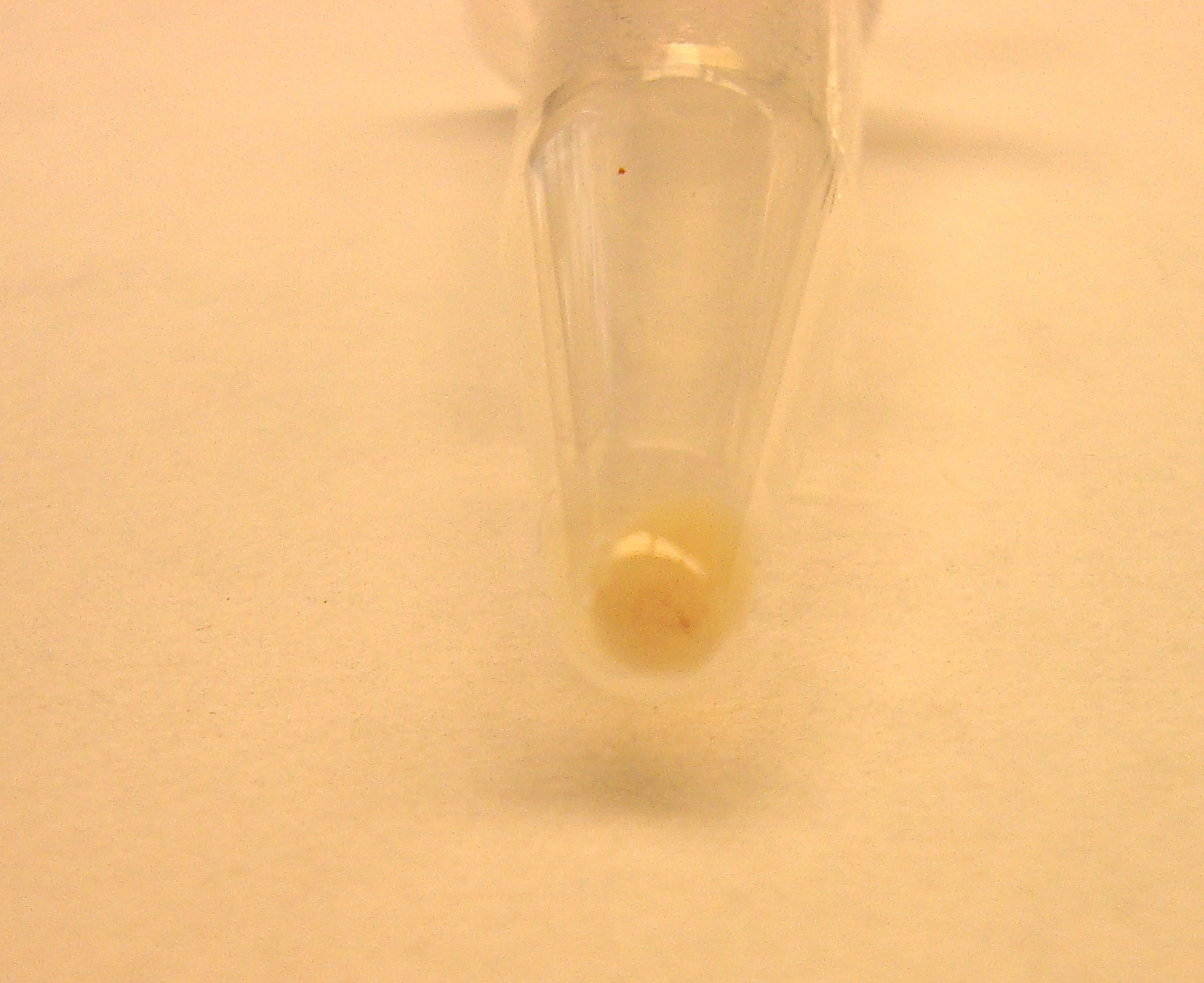
| + | <center>[[Image:CCAUC10RFP.jpg|thumb|500px|none|2 mL of dense culture was pelleted. These cells contain the construct pBad-CCAUC10 tRNA-pLac-RBS-CCAUC FSL-RFP. You can see some red specks of color in the pellet.]]</center> | ||
{{Template:MoWestern_Davidson2009_end}} | {{Template:MoWestern_Davidson2009_end}} | ||
Latest revision as of 20:04, 21 October 2009

Contents |
Completed tRNAs
We assembled the genes for our suppressor tRNAs by ligating single-stranded oligos with sticky BioBrick ends. Go here to see a full explanation, or [http://www.bio.davidson.edu/courses/Molbio/Protocols/anneal_oligos.html here] for the protocol. This generated double-stranded DNA ready to ligate into a standard vector. All tRNAs deliver the same amino acid, serine. The expected size of our tRNAs with BioBrick ends is 184 bp. We verified that we assembled the expected tRNAs by checking the insert size of our ligation (seen below), and by sequencing our tRNA plasmids.
For controlling our suppressor tRNA expression, we wanted a promoter that would transcribe enough tRNA to cause a high probability of suppression but not so much tRNA that cells would become sick. We decided to use a constitutive weak-medium promoter, pBad, to express our suppressor tRNAs.
This promoter choice allows us to induce greater expression with L-arabinose. We found that suppressor tRNAs controlled by pBad is not lethal to E. coli and allows for normal culture growth.
Successfully Assembled tRNAs
| Codon | Anticodon Loop |
|---|---|
| AGGAC | CUGUCCUAA |
| CGGUC | UUGACCGAC |
| CUACC | GUGGUAGAA |
| CUACU | UUAGUAGAU |
| CUAGC | CUGCUAGAA |
| CUAGU | UUACUAGAC |
| CCAAU | CUAUUGGAC |
| CCACC | UUGGUGGAA |
| CCACU | CUAGUGGAC |
| CCAUC (9-bp anticodon) | GUGAUCCAA |
| CCAUC (10-bp anticodon) | UUUGAUGGAG |
| CCCUC | CUGAGGGUC |
Completed Reporters With Frameshift Suppressor Leaders (FSLs)
In order to engineer logical clauses directly upstream of our selected reporter genes, we altered the 5’ end of the reporter using PCR to add frameshift suppressor leaders (FSLs). FSLs were added after the reporter start codon ATG so that the suppressor tRNAs are required to correctly translate the entire reporter. We began with a modular 1-SAT approach - adding a single 5-nt codon as the FSL to the 5' end of the reporter.
Successful FSL Additions
| CUAGC | CUAGC | ----- |
| ----- | CCAUC | ----- |
| ----- | CGGUC | CGGUC |
| ----- | CUACU | CUACU |
| ----- | CUAGU | CUAGU |
| ----- | CCCUC | ----- |
| ----- | AGGAC | ----- |
| ----- | CUACC | ----- |
| ----- | CCAAU | ----- |
| ----- | CCACC | ----- |
| ----- | CUACU | ----- |
The PCR template for our FSL additions were reporter constructs with a promoter and RBS. For example, pLac-RBS-RFP was the template plasmid for the FSL-RFPs. This was helpful after PCR and ligation into the template vector, so that we could select potential FSL-RFPs by colour selection.
Below is a plate of colonies transformed with our potential FSL-RFPs. Red colonies clearly have the original construct since they still express RFP in the absence of the appropriate suppressor tRNA. White colonies potentially have our desired construct containing the FSL upstream of RFP.
Selected colonies were sequenced for verification of the correct FSL addition. The next step is to ligate each tRNA with the corresponding FSL-reporter, so that both genes will be present and expressed within the cell. We are attempting two possible constructs for this:
1.The tRNA controlled by a separate promoter from the reporter.
2.The tRNA directly downstream of the reporter.
Choosing A Promoter
One important consideration in our project design was choosing the promoter(s) that would induce a notable phenotype from our reporters. For controlling our reporters with a leading logical clause, we wanted a promoter that would have strong enough transcription to allow more chances for suppression, and thereby more reporter expression.
Promoter Control of Fluorescence Reporters
In order to have a visible expression of RFP or GFP, we constructed parts with RBS-RFP under the control of 4 different commonly-used promoters:
- pBad (Potential induction with L-arabinose)
- pLac (Potential induction with IPTG)
- pLacIQ (Potential induction with IPTG)
- pTet
We found that pLac induced with IPTG caused the greatest expression of RFP. This construct, pLac-RBS-RFP, was chosen as our control construct, representing 100% suppression of the engineered frameshift.
Completed Constructs - Suppressor tRNAs and FSL-RFP Reporters
Based on promoter control data, we decided to use pBad promoter upstream of suppressor tRNAs, and to use pLac promoter upstream of the RFP reporter with FSLs. The benefit of these promoter choices is constitutive expression with the possibility of further induction. We chose pLac for RFP expression because it caused the greatest expression in the control construct (promoter-RBS-RFP, no FSL).
We completed successfully Construct 1: pLac-RBS-FSL-RFPs were ligated downstream of the corresponding pBad-tRNA (i.e. The tRNA upstream of pLac-RBS-CGGUC-RFP is the suppressor of the FSL CGGUC). This construct was transformed into E. coli, and transformation was verified by digestion and electrophoresis, and by sequencing isolated plasmid. Cells with the verified construct were grown in LB media with inducers 2.5% L-arabinose and 0.06% IPTG. Fluorescence readings were measured after 20 hours of incubation. The figure below shows the measurements, normalized to the negative control, of all FSL constructs that showed any fluorescence. We observed red fluorescence most notably in the construct containing the tRNA CCAUC 10-bp anticodon (below).

The red fluorescence in CCAUC 10-bp construct was also visible in the pelleted cell culture (below). These data indicate a construct containing our suppressor tRNAs can successfully suppress their target 5-bp codon, allowing expression of RFP. Therefore, we have designed and built a mechanism to solve a very simple SAT problem in a biological construct.
 "
"