Team:Calgary/20 August 2009
From 2009.igem.org
(Difference between revisions)
| (9 intermediate revisions not shown) | |||
| Line 336: | Line 336: | ||
Using a 96 well microplate, AI-2 isolated from <i>Salmonella typhimurium</i> 14028 supernatant will be tested. <i>V. harveyi</i> reporter strain MM32 (luxS- and luxN-) will luminesce in the presence of AI-2. Controls used in the experiment include <i>S. typhimurium</i> SS007, <i>V. harveyi</i> BB120 and <i> E. coli</i> DH5a. Overnight OD600 and luminescence (CPS) readings will be taken by a VICTOR (thank you Surette lab, especially Margot!). <br /><br /> | Using a 96 well microplate, AI-2 isolated from <i>Salmonella typhimurium</i> 14028 supernatant will be tested. <i>V. harveyi</i> reporter strain MM32 (luxS- and luxN-) will luminesce in the presence of AI-2. Controls used in the experiment include <i>S. typhimurium</i> SS007, <i>V. harveyi</i> BB120 and <i> E. coli</i> DH5a. Overnight OD600 and luminescence (CPS) readings will be taken by a VICTOR (thank you Surette lab, especially Margot!). <br /><br /> | ||
| - | Preliminary testing was done on the bacterial transformation and genomic DNA isolation activities in the virtual lab. Additionally, I now have transparent wings which glow | + | Preliminary testing was done on the bacterial transformation and genomic DNA isolation activities in the virtual lab. Additionally, I now have transparent wings which glow. |
</div> | </div> | ||
</td> | </td> | ||
| Line 381: | Line 381: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Fundamental Displays | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | I recently remembered that I will have to start making cleaning scripts for a lot of the equipment in the lab to prevent clutter for easy maintenance. I started to make these and added them to bacterial transformation, the sequencing station and DNA extraction. It is not imperative that these be completed immediately since this type of abuse is not expected for testing. | |
| + | <br> | ||
| + | <br> | ||
| + | Added rho termination, but I am not satisfied with it yet and will continue to change where objects travel too although the pushing part looks satisfactory I need a better place for the mRNA strand. I have no idea how I am going to do the hairpin termination at this time but it will probably mean I have to fiddle with rotation again. There is now a complementary strand of DNA that will change its bases when an avatar makes changes to the template strand (the bottom one strand in this animation) so that the two strands will remain complementary to one another. | ||
| + | <br> | ||
| + | <br> | ||
| + | Also did some more wiki updates. | ||
<html> | <html> | ||
| Line 427: | Line 433: | ||
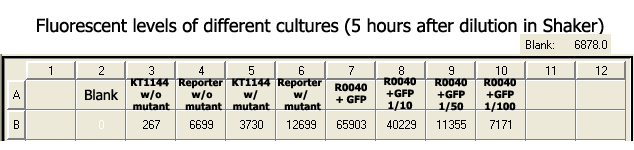
1. Both with and without mutant readings of TOP10 cells with reporter are than the readings of KT1144 cells. I do not know exactly why it is as such, but my guess is that different type of cells absorb different amount of wavelengths. | 1. Both with and without mutant readings of TOP10 cells with reporter are than the readings of KT1144 cells. I do not know exactly why it is as such, but my guess is that different type of cells absorb different amount of wavelengths. | ||
<br> | <br> | ||
| - | 2. The fluorescent levels of the cells that | + | 2. The fluorescent levels of the cells that were grown for an extra 5 hours in the shaker were lower than the cells that grew in the plate reader, except for the cells that were diluted with fresh LB media. As above, I do not know the exact reason. Perhaps it is because the cells have already reached their 16~20 hours of growing overnight, growing for an extra 5 more hours at 37˚C killed the bacteria, decreasing the production of GFP, while the growth of the cells that were stored in the plate reader for an extra 5 hours slowed down due to a relatively colder temperature, thus being able to produce GFP more stably. |
As for the diluted cells with R0040+GFP, because they were transferred to a fresh media before the additional growth of 5 hours, storing them in the shaker at 37˚C led to faster replication, which sped up the production of GFP, while storing them in the colder plate reader slowed down the replication, which slowed the production of GFP. | As for the diluted cells with R0040+GFP, because they were transferred to a fresh media before the additional growth of 5 hours, storing them in the shaker at 37˚C led to faster replication, which sped up the production of GFP, while storing them in the colder plate reader slowed down the replication, which slowed the production of GFP. | ||
<html> | <html> | ||
| Line 507: | Line 513: | ||
<b>Sequence of Pqrr4+B0034 (RBS) C1</b> | <b>Sequence of Pqrr4+B0034 (RBS) C1</b> | ||
<Br> | <Br> | ||
| - | + | Legend | |
| + | <Br> | ||
<font color="red">Vector contamination</font> | <font color="red">Vector contamination</font> | ||
<br> | <br> | ||
| Line 536: | Line 543: | ||
<br> | <br> | ||
| - | When I carefully analyzed | + | When I carefully analyzed today's <i>Pqrr4</i>+B0034(C1)+K082003, I could find part of the B0034 embeded between <i>Pqrr4</i> and K082003. The following explains this further: |
<br> | <br> | ||
<br> | <br> | ||
| Line 667: | Line 674: | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
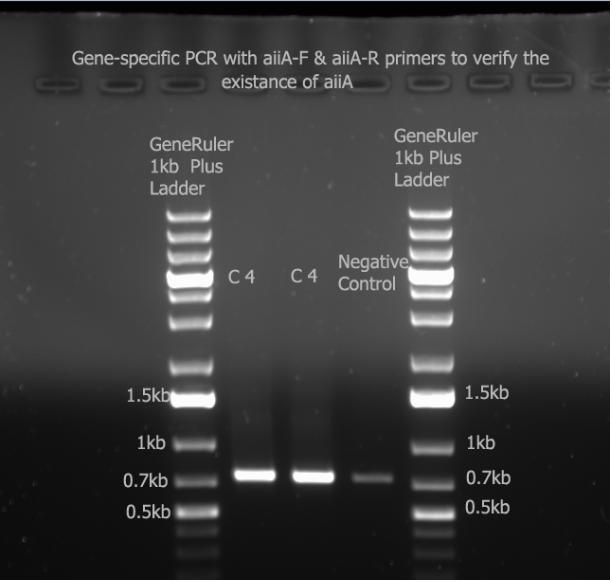
| + | I ran the gene-specifc PCR with aiiA-F/R primers and ran it with a 1% gel at 100V. | ||
| + | <br> | ||
| + | <center> | ||
| + | [[Image:Calgary_2009.08.19.gene.specific.aiiapcr.png|500px]] | ||
| + | </center> | ||
| + | <br> | ||
| + | I had contaminated my negative control (which was just the mastermix) with my aiiA gene. Since all three bands are around 756 bp, we can say that the gene exists. | ||
| + | |||
I followed up with couple pharmaceutical companies. Some of our potential sponsors are currently deciding on sponsoring iGEM Calgary. Again, I couldn't get hold of other companies by phone but I left a message with a call back number. I also emailed those companies with a follow- up email. The good news is that Beckman Coulter has generously donated $200.00 to our iGEM Team this year. Unfortuantely, Alda Phamaceuticals Corporation is unable to sponsor our team this year. | I followed up with couple pharmaceutical companies. Some of our potential sponsors are currently deciding on sponsoring iGEM Calgary. Again, I couldn't get hold of other companies by phone but I left a message with a call back number. I also emailed those companies with a follow- up email. The good news is that Beckman Coulter has generously donated $200.00 to our iGEM Team this year. Unfortuantely, Alda Phamaceuticals Corporation is unable to sponsor our team this year. | ||
| Line 692: | Line 707: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Testing...Testing | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | Today, a generic avatar was created (aptly named Generic Lennie) that was in a separate group that had access to our island. He was used to test each area on our sim. We don't want visitors to be able to move, objects, change script and make new permanent objects. For the synthetic Kingdom everything was in order: | |
| + | <br> | ||
| + | <br> | ||
| + | *All objects were not modifiable | ||
| + | *Generic Lennie was able to move the vitamins, Bad Guy/Champion cells, quorum sensing bacteria and oil clean-up bacteria, which is ok because they have a self-destruct script inside | ||
| + | *GFP station drag and drop worked! | ||
| + | <br> | ||
| + | This means that the Synthetic Kingdom is unofficially done! | ||
<html> | <html> | ||
| Line 718: | Line 740: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | ...And some more work on the modelling front | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | * Chinee and I met with Afshin and Iman to align our models with each other. The SimBiology and Membrane Computing reactions were subsequently updated, so that we are now modelling the same thing where applicable. The only major differences lie in that the membrane computing model accounts for LuxS (which the SimBiology model omits) and the membrane computing model accounts for AI-2 molecules leaving the cell and going to another one (while ours just includes a reverse reaction for AI-2 binding to LuxPQ, as we are only modelling performance in one cell). | |
| + | * I helped Chinee download Latex and will provide a lesson on how to use that platform | ||
| + | * I looked up degradation rates for GFP:LVA mRNA | ||
| + | * Our sensitivity analysis crashes with the addition of more reactions to our model. I worked on fixing that, to little avail. I will try again tomorrow. | ||
<html> | <html> | ||
Latest revision as of 21:40, 10 September 2009
UNIVERSITY OF CALGARY

 "
"