Team:KULeuven/Project/Details
From 2009.igem.org
Bart Bosmans (Talk | contribs) |
(→Vanillin receptor and Antikey) |
||
| (18 intermediate revisions not shown) | |||
| Line 5: | Line 5: | ||
== Blue light receptor == | == Blue light receptor == | ||
| + | [[Image:Missblue-deckchair.png|100px|left]] | ||
[[Image:Biologie_blue_light.png|center]] | [[Image:Biologie_blue_light.png|center]] | ||
The protein YcgF is a known blue-light sensor in certain ''E. coli'' strains. Upon photo-excitation it dimerizes and acts as an anti-repressor for YcgE. YcgE is bound to the promotor-region and inhibits RNA Polymerase. The dimerized YcgF interacts directly with the repressor, releasing it from the DNA and allowing transcription. | The protein YcgF is a known blue-light sensor in certain ''E. coli'' strains. Upon photo-excitation it dimerizes and acts as an anti-repressor for YcgE. YcgE is bound to the promotor-region and inhibits RNA Polymerase. The dimerized YcgF interacts directly with the repressor, releasing it from the DNA and allowing transcription. | ||
| - | We designed the | + | We designed the {{kulpart|BBa_K238002}} part in such way that irradiation with a certain amount of blue light activates transcription of key-mRNA. To achieve this, we purified the promoter-region of ''E. coli'' MC4100. After mutating out possible restriction sites, the blr promoter, {{kulpart|BBa_K238000}} part was added to the registry. |
== Key/Lock == | == Key/Lock == | ||
| - | + | [[Image:Miss_Blue_key_lock.png|150px|right]] | |
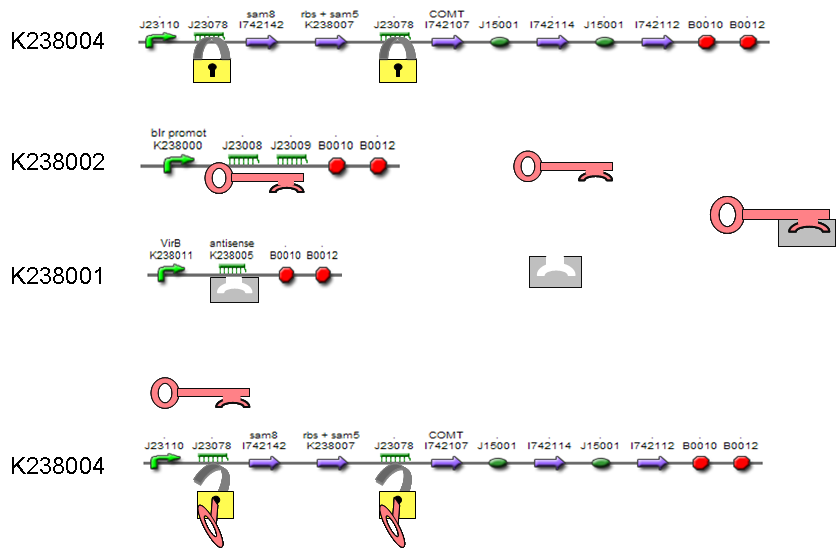
The mRNA ‘key’ and ‘lock’ sequences form a pair of riboregulators. The ‘lock’ DNA is located directly upstream of the controlled gene’s RBS. It is folded into a stem-loop secondary structure which prevents access to the ribosome and inhibits translation. ‘Keys’ are expressed from separate genes (in trans) and code for sequences complementary to the lock. Upon annealing they unlock the ‘closed’ stem-loops, thereby exposing the RBS and permitting expression of the gene(s) downstream. | The mRNA ‘key’ and ‘lock’ sequences form a pair of riboregulators. The ‘lock’ DNA is located directly upstream of the controlled gene’s RBS. It is folded into a stem-loop secondary structure which prevents access to the ribosome and inhibits translation. ‘Keys’ are expressed from separate genes (in trans) and code for sequences complementary to the lock. Upon annealing they unlock the ‘closed’ stem-loops, thereby exposing the RBS and permitting expression of the gene(s) downstream. | ||
| - | The most efficient key is a combination of the | + | The most efficient key is a combination of the {{kulpart|BBa_J23008}} and {{kulpart|BBa_J23009}} keys. They are placed behind the blue light promoter to ensure transcription after blue light irradiation. {{kulpart|BBa_B0015}} is used as terminator. This brick consists of {{kulpart|BBa_B0010}} and {{kulpart|BBa_B0012}} and is most commonly used. One {{kulpart|BBa_J23078}} lock is placed before Sam5 and Sam8 genes and another before COMT to guarantee maximal efficiency of the key/lock system. These genes are part of the vanillin production pathway. Therefore, after a blue light activates transcription of the key, vanillin synthesis starts. |
[[Image:Key_lock_antikey.png|thumb|600px|center]] | [[Image:Key_lock_antikey.png|thumb|600px|center]] | ||
== Vanillin production == | == Vanillin production == | ||
| + | [[Image:Miss_Blue_vanillin_production.png|100px|left]] | ||
[[Image:Biologie_vanillin_synthesis.png|center]] | [[Image:Biologie_vanillin_synthesis.png|center]] | ||
[[image:Vanillin_Biosynthesis_Pathway.jpg|thumb|400px|right|Vanillin production overview starting from Tyrosine.]] | [[image:Vanillin_Biosynthesis_Pathway.jpg|thumb|400px|right|Vanillin production overview starting from Tyrosine.]] | ||
| - | A vanilla odour is created by synthesizing the molecule Vanillin. The starting point is tyrosine, an amino acid produced endogenously in E.coli. The subsequent pathway involves a combination of five enzymes, biobricked with following codes: | + | A vanilla odour is created by synthesizing the molecule Vanillin. The starting point is tyrosine, an amino acid produced endogenously in ''E.coli''. The subsequent pathway involves a combination of five enzymes, biobricked with following codes: |
| - | |||
| - | |||
| - | • | + | • SAM8: {{kulpart|BBa_I742142}}: coding sequence without RBS |
| - | • | + | • SAM5: {{kulpart|BBa_K238007}}: RBS site + PstI restriction site removed |
| - | • | + | • COMT: {{kulpart|BBa_I742107}}: coding sequence without RBS |
| + | |||
| + | • FCS: {{kulpart|BBa_I742115}}: RBS + fcs in pSB1A2 | ||
| + | |||
| + | • ECH: {{kulpart|BBa_I742113}}: RBS + ech in pSB1A2 | ||
| Line 38: | Line 42: | ||
| - | COMT if found in the alfalfa plant and translates to a caffeic acid-O-methyl transferase. After -OH methylation on C4 of the aromatic ring, it produces ferrulic acid from caffeic acid. The fourth gene Fcs is derived from ''Pseudomonas fluorescens'' and encodes feruoyl CoA synthase. This enzyme ligates acetyl-CoA onto ferrulic acid and produces feruloyl CoA. Ech another enzyme from the same species completes the pathway. The gene-product, an enoyl CoA hydratase, cleaves the CoA group from feruloyl CoA thereby converting it to vanillin. | + | COMT if found in the alfalfa plant and translates to a caffeic acid-O-methyl transferase. After -OH methylation on C4 of the aromatic ring, it produces ferrulic acid from caffeic acid. The fourth gene Fcs is derived from ''Pseudomonas fluorescens'' and encodes feruoyl CoA synthase. This enzyme ligates acetyl-CoA onto ferrulic acid and produces feruloyl CoA. Ech, another enzyme from the same species, completes the pathway. The gene-product, an enoyl CoA hydratase, cleaves the CoA group from feruloyl CoA thereby converting it to vanillin. |
| - | The entire | + | The entire {{kulpart|BBa_K238004}} biobrick is placed under a constitutive promoter {{kulpart|BBa_J23110}} and {{kulpart|BBa_B0015}}. |
== Vanillin receptor and Antikey == | == Vanillin receptor and Antikey == | ||
| + | [[Image:Miss_Blue_vanillin_sensor.png|160px|left]] | ||
[[image:Biologie_vanillin_receptor.png|center]] | [[image:Biologie_vanillin_receptor.png|center]] | ||
[[image:Biologie_antikey.png|center]] | [[image:Biologie_antikey.png|center]] | ||
| - | |||
| - | Together, ''VirA'' and ''VirG'' form a two component system that recognizes phenol derivatives like Vanillin and other signals like low pH, certain aldose monosaccharides and limited phosphate. ''VirA'' dimers function as a sensor kinase. They process all the input signals and phosphorylate ''VirG''. ''VirG'' then binds to a Vir box sequence located in the ''VirB'' promoter region | + | Certain virulent bacteria like ''Agrobacterium Tumefacens'' contain a large tumor-inducing plasmid that carries different virulence (''vir'') genes. The Vanillin receptor is composed of two of those ''vir'' genes: ''VirA'' K238008 {{kulpart|BBa_K238008}} and ''VirG'' {{kulpart|BBa_K238009}}. RpoA ({{kulpart|BBa_K238010}}) is an alfa subunit polymerase that helps ''VirG'' work in'' E. coli''. Transcription is controlled by the constitutional promoter {{kulpart|BBa_J23110}} and terminator {{kulpart|BBa_B0015}}. {{kulpart|BBa_B0032}} is chosen as RBS. |
| + | |||
| + | |||
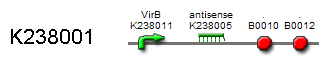
| + | Together, ''VirA'' and ''VirG'' form a two component system that recognizes phenol derivatives like Vanillin and other signals like low pH, certain aldose monosaccharides and limited phosphate. ''VirA'' dimers function as a sensor kinase. They process all the input signals and phosphorylate ''VirG''. ''VirG'' then binds to a Vir box sequence located in the ''VirB'' promoter region {{kulpart|BBa_K238011}}. As both ''VirG'' and ''rpoA'' are necessary for transcription, ''rpoA'' is constitutively transcribed. This action mediates the desired response: transcription of the ‘antikey’ {{kulpart|BBa_K238005}}. | ||
By means of this antisense ‘key’ mRNA, vanillin synthesis can be regulated. Upon transcription, the ‘antikey’ binds the complementary ‘key’-mRNA away from the lock. As a consequence, the pathway to vanillin synthesis remains locked. The amount of ‘antikey’ depends ultimately on the set intensity of irradiated blue light, because this determines the concentration of produced Vanillin that activates the receptor. Therefore, any desired Vanillin concentration can be adjusted just by altering irradiation intensity. | By means of this antisense ‘key’ mRNA, vanillin synthesis can be regulated. Upon transcription, the ‘antikey’ binds the complementary ‘key’-mRNA away from the lock. As a consequence, the pathway to vanillin synthesis remains locked. The amount of ‘antikey’ depends ultimately on the set intensity of irradiated blue light, because this determines the concentration of produced Vanillin that activates the receptor. Therefore, any desired Vanillin concentration can be adjusted just by altering irradiation intensity. | ||
Latest revision as of 14:29, 10 September 2009
Blue light receptor
The protein YcgF is a known blue-light sensor in certain E. coli strains. Upon photo-excitation it dimerizes and acts as an anti-repressor for YcgE. YcgE is bound to the promotor-region and inhibits RNA Polymerase. The dimerized YcgF interacts directly with the repressor, releasing it from the DNA and allowing transcription. We designed the part in such way that irradiation with a certain amount of blue light activates transcription of key-mRNA. To achieve this, we purified the promoter-region of E. coli MC4100. After mutating out possible restriction sites, the blr promoter, part was added to the registry.
Key/Lock
The mRNA ‘key’ and ‘lock’ sequences form a pair of riboregulators. The ‘lock’ DNA is located directly upstream of the controlled gene’s RBS. It is folded into a stem-loop secondary structure which prevents access to the ribosome and inhibits translation. ‘Keys’ are expressed from separate genes (in trans) and code for sequences complementary to the lock. Upon annealing they unlock the ‘closed’ stem-loops, thereby exposing the RBS and permitting expression of the gene(s) downstream.
The most efficient key is a combination of the and keys. They are placed behind the blue light promoter to ensure transcription after blue light irradiation. is used as terminator. This brick consists of and and is most commonly used. One lock is placed before Sam5 and Sam8 genes and another before COMT to guarantee maximal efficiency of the key/lock system. These genes are part of the vanillin production pathway. Therefore, after a blue light activates transcription of the key, vanillin synthesis starts.
Vanillin production
A vanilla odour is created by synthesizing the molecule Vanillin. The starting point is tyrosine, an amino acid produced endogenously in E.coli. The subsequent pathway involves a combination of five enzymes, biobricked with following codes:
• SAM8: : coding sequence without RBS
• SAM5: : RBS site + PstI restriction site removed
• COMT: : coding sequence without RBS
• FCS: : RBS + fcs in pSB1A2
• ECH: : RBS + ech in pSB1A2
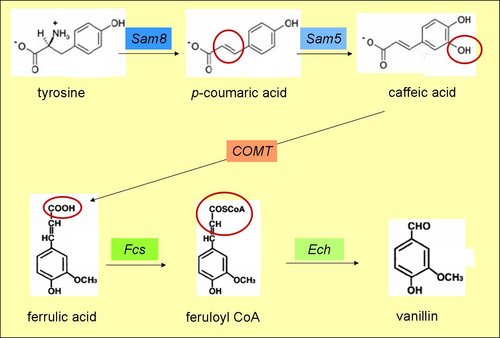
The first gene Sam8 originates from Saccharothrix espeanensis and encodes a tyrosine ammonia lyase. This enzyme catalyses the deamination of tyrosine's amine group and converts tyrosine to p-coumaric acid. Sam5 is derived from the same species, but encodes a 4-coumarate 3-hydroxylase which hydroxylates C4 in the aromatic ring of p-coumaric acid. p-Coumaric acid is converted to caffeic acid.
COMT if found in the alfalfa plant and translates to a caffeic acid-O-methyl transferase. After -OH methylation on C4 of the aromatic ring, it produces ferrulic acid from caffeic acid. The fourth gene Fcs is derived from Pseudomonas fluorescens and encodes feruoyl CoA synthase. This enzyme ligates acetyl-CoA onto ferrulic acid and produces feruloyl CoA. Ech, another enzyme from the same species, completes the pathway. The gene-product, an enoyl CoA hydratase, cleaves the CoA group from feruloyl CoA thereby converting it to vanillin.
The entire biobrick is placed under a constitutive promoter and .
Vanillin receptor and Antikey
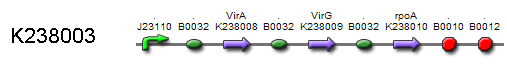
Certain virulent bacteria like Agrobacterium Tumefacens contain a large tumor-inducing plasmid that carries different virulence (vir) genes. The Vanillin receptor is composed of two of those vir genes: VirA K238008 and VirG . RpoA () is an alfa subunit polymerase that helps VirG work in E. coli. Transcription is controlled by the constitutional promoter and terminator . is chosen as RBS.
Together, VirA and VirG form a two component system that recognizes phenol derivatives like Vanillin and other signals like low pH, certain aldose monosaccharides and limited phosphate. VirA dimers function as a sensor kinase. They process all the input signals and phosphorylate VirG. VirG then binds to a Vir box sequence located in the VirB promoter region . As both VirG and rpoA are necessary for transcription, rpoA is constitutively transcribed. This action mediates the desired response: transcription of the ‘antikey’ .
By means of this antisense ‘key’ mRNA, vanillin synthesis can be regulated. Upon transcription, the ‘antikey’ binds the complementary ‘key’-mRNA away from the lock. As a consequence, the pathway to vanillin synthesis remains locked. The amount of ‘antikey’ depends ultimately on the set intensity of irradiated blue light, because this determines the concentration of produced Vanillin that activates the receptor. Therefore, any desired Vanillin concentration can be adjusted just by altering irradiation intensity.
 "
"