Team:Washington/Project/Target
From 2009.igem.org
| Line 1: | Line 1: | ||
| - | + | __NOTOC__ | |
{{Template:Team:Washington/Templates/Header}} | {{Template:Team:Washington/Templates/Header}} | ||
Revision as of 20:32, 18 October 2009
Background
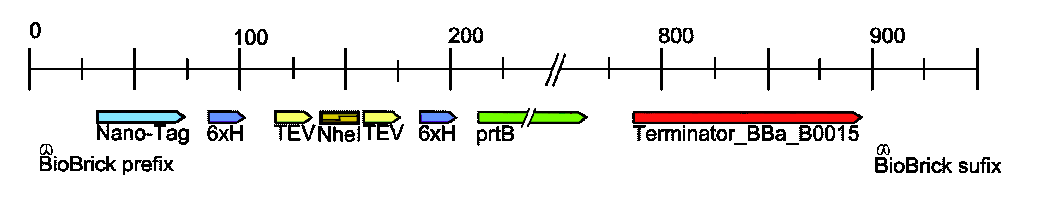
When a favorite protein (afp) is cloned into the target vector ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K215002 BBa_K215002]) two tags are fused onto the N and C terminal of afp. These tags are depicted below:
The first key feature of the target vector is the NheI restriction site, since this is where afp get's inserted. NheI is compatible with XbaI and SpeI, meaning that a biobrick digested at the X and S sites can be ligated into the target vector at the NheI site (for detailed protocol see: NheI Insertion Protocol).
At the N-terminus of the Target is the display (aka Nano) tag, which is a 15 amino acid sequence that binds to streptavidin. Since streptavidin is being displayed on the surface of the cell this allows our protein to stick to the outside of the cell, but can still be released by the addition of biotin. For more details see: Surface Display System .
At the C-terminus of the Target is a secretion tag (prtB) that is recognized by the Type I secretion system, which secretes proteins from the cytosol, through the periplasim, and into the media. For more details go to: Secretion System .
Flanking each side of the NheI site are 6 consecutive hisitidines (6x-His) and TEV protease sites. The histidines allow for traditional immobilized metal affinity chromatography (IMAC) protein purification. The TEV sites allows for the N and C terminal tags to be cleaved off of afp, and due to the strategic placement of the 6x-His tags these tags can then be seperated from afp by simply running the cleaved solution over a column in which the tags stick but afp flows right through.
Experiments
For the target vector our goal was to:
- Construct the target vector
- Insert GFP into the target vector and characterize expression and function
- Biobrick and characterize OpdA, a nerve agent degrading enzyme
- Insert OpdA into the target vector and characterize expression and function
Target Vector Construction
To create the target vector we first synthesized a coding sequence that would produce the protein as described above. To do this we synthesized a gene from oligo's as described in our Gene Synthesis Protocol.
After creating the target construct, we created and BioBricked an expression vector (BBa_K215000) which would express the target protein upon induction with IPTG. We then added the target construct into the expression vector using standard assembly.
GFP Insertion and Characterization
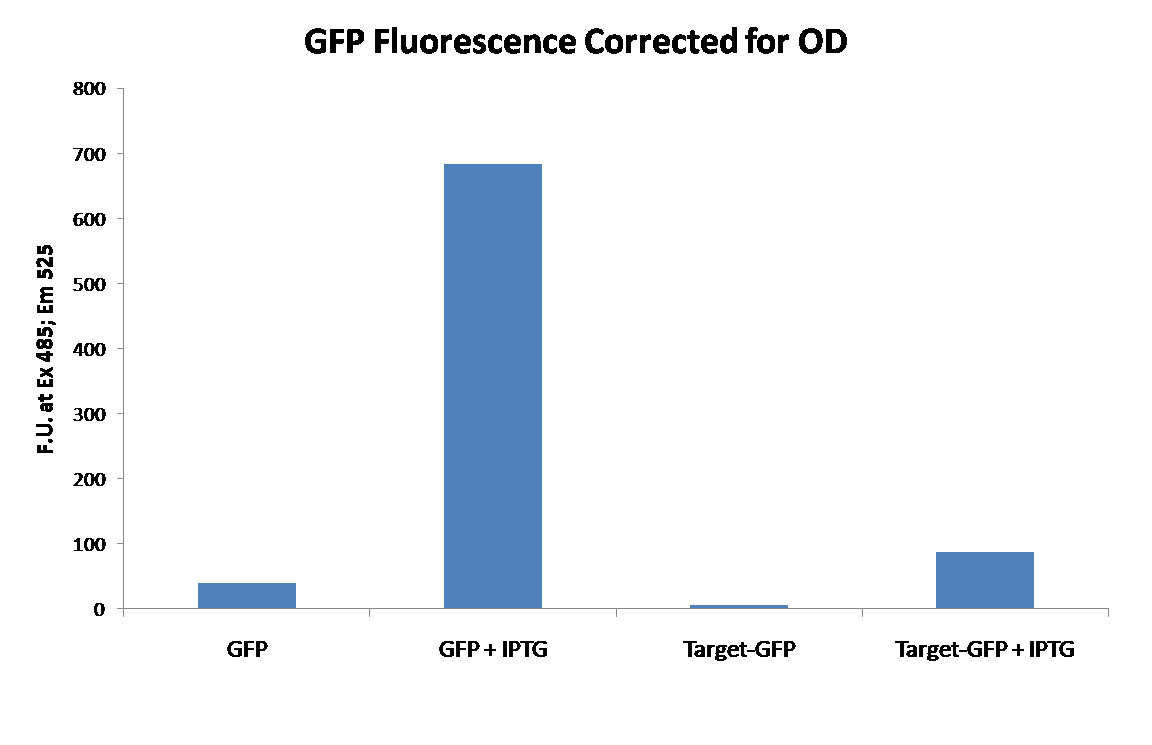
Upon completion of constructing the target vector our first experiment was to determine if GFP was still functional as the fusion protein. To do this GFP (E0040) was inserted into the target construct use the NheI method as described above. After insertion of E0040 into the target construct, the tagged GFP (target-GFP) was transformed into BL21(lacIq) cells, and subsequently grown in the presence or absence of IPTG. As a control an untagged E0040 was cloned into our expression vector and also grown in the presence or absence of IPTG. This would allow us to determine the effects of the tags on fluorescence. The cells were then washed with PBS, normalized to the same cell density, and fluorescence measured using an excitation of 485 and emission of 525 (cutoff at 515) in a XXXXX plate reader. The data is show below:
From this data we were able to conclude that our expression vector was functional, as is evident from the large increase in fluorescence with the addition of IPTG. We were also able to conclude that the Target-GFP is functional, but fluorescence was significantly decreased.
In order to ensure that the Target-GFP had the appropriate 6x-his tags and that fluorescence was a function of protein concentration we purified Target-GFP using a traditional IMAC techniques (ADD TO PROTOCOLS). The protein concentration was measured from its absorbence a 280nm. A serial dilution of the protein was then made and the resulting fluorescence measured as described earlier. The data is shown below:
As expected the fluorescence intensity is linear with respect to protein concentration.
BioBricking and Characterization of OpdA
OpdA Background
OpdA (BBa_K215090) is an organophosphate-degrading enzyme from Agrobacterium radiobacter. It is capable of degrading a wide range of organophosphates, most notably pesticides that are poisonous to humans, such as paraoxon. We chose to biobrick and submit this enzyme to the registry for a number of reasons. First and foremost, this enzyme is easy to assay for since it can hydrolyze substrates very quickly (e.g. paraoxon) and form a bright yellow product. This yellow product would make it easy to see that the OpdA was present and functioning in our system. And secondly, OpdA is a very useful enzyme that could have applications in future iGEM and other synthetic biology projects, so its presence in the Standard Registry of Biological Parts is beneficial.
OpdA Characterization
The Baker lab donated the source plasmid for OpdA (a synthetic gene optimized for E. coli expression). SOEing PCR was used to remove BioBrick sites (ADD PROTOCOL). Upon removal of the unwanted restriction sites the gene was cloned into pSB1A2 (BB#), our expression vector (BB#), and the target vector.
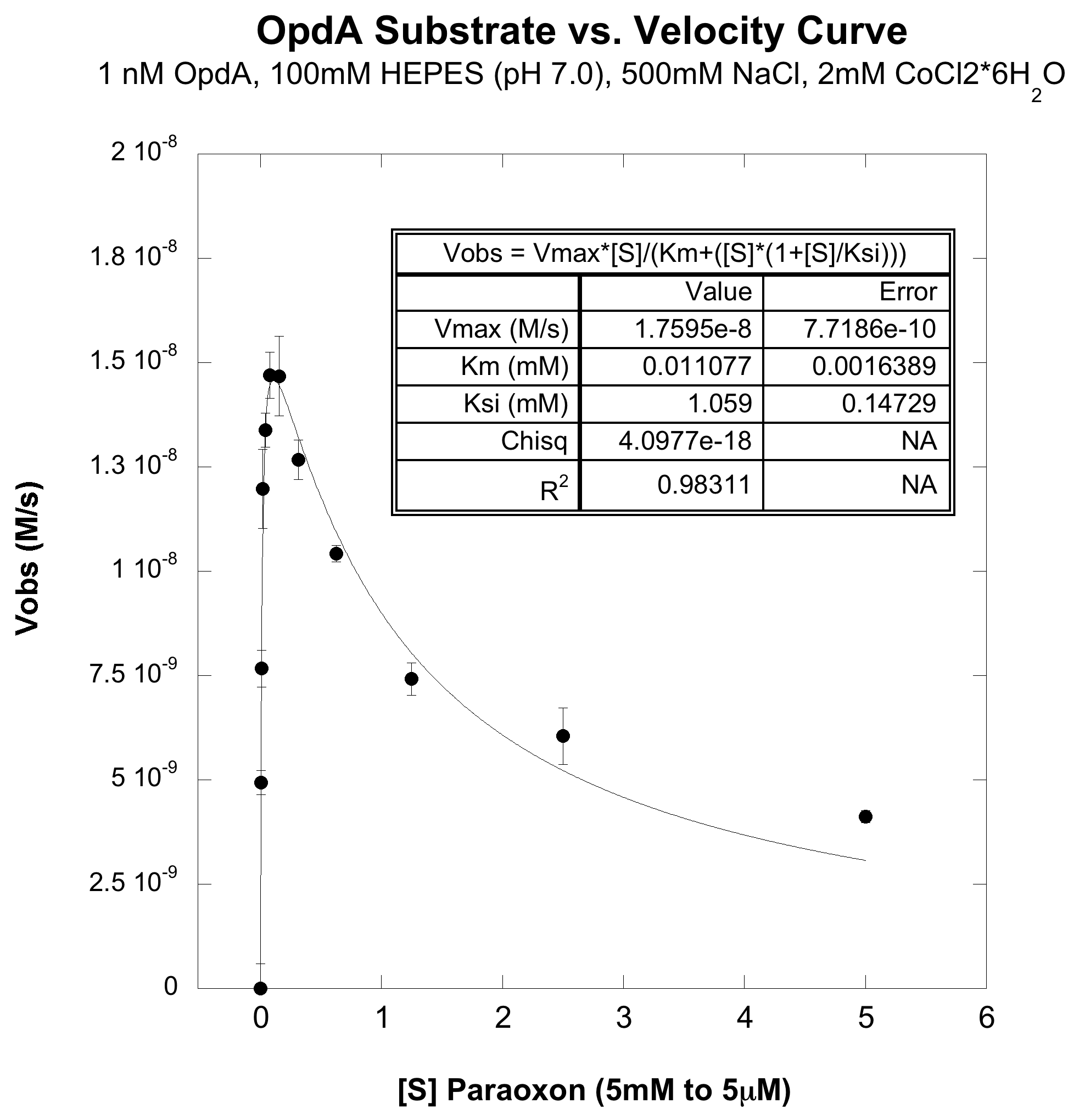
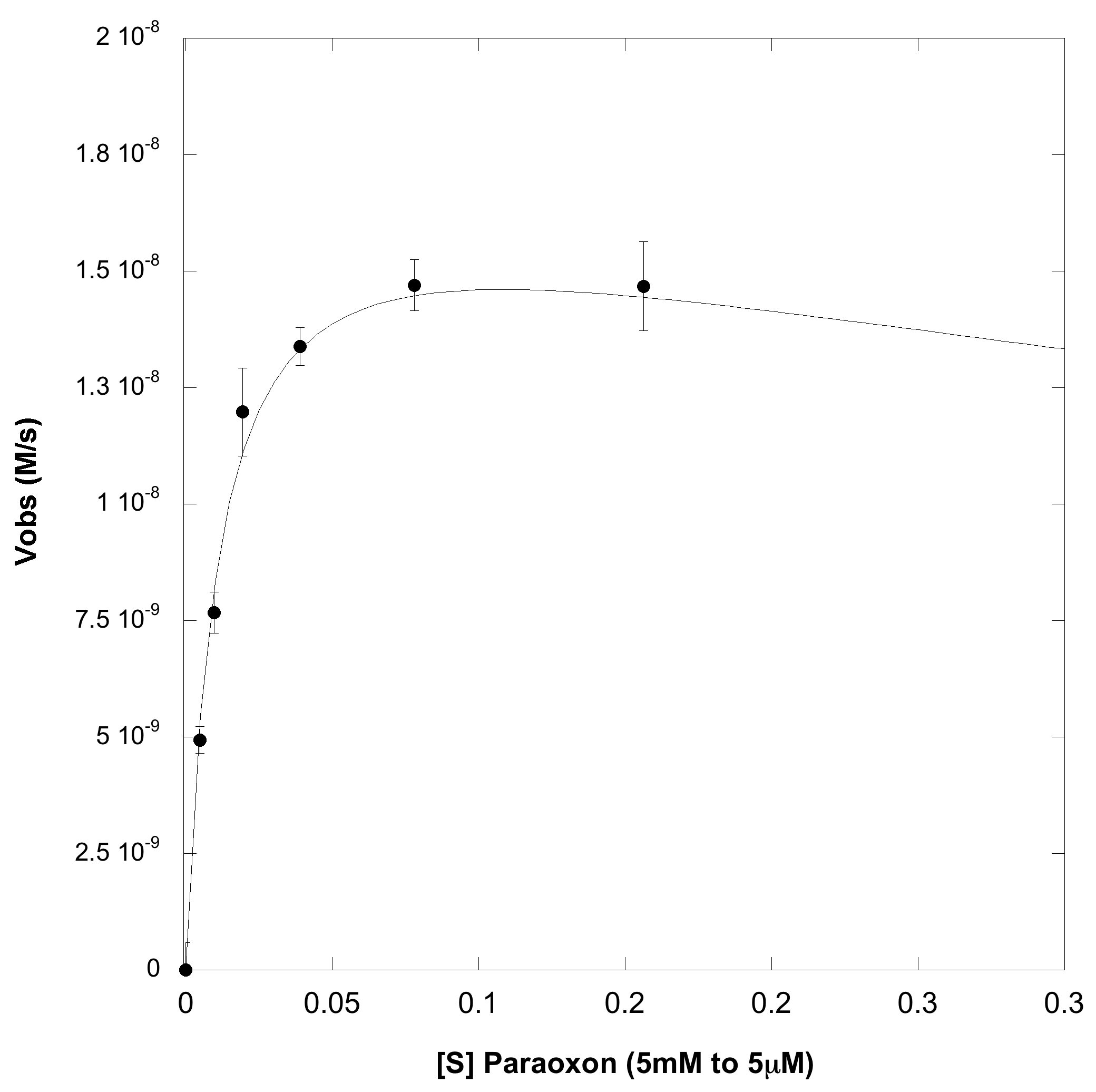
The first experiment carried out was to validate that we could express and purify functional OpdA. To do this we transformed BB# into a BL21(lacIq) cell line and followed a traditional protein production and purification procedure (ADD PROTOCOL). The resulting purified protein was then dialyzed overnight in 1x PBS to remove the elution buffer which we were worried would interfere with the activity assay. The concnetration of the dialysed protein was dteremined by meauring its absorbance at 280nM and using its predicted extinction coefficient (Value, LINK). We obtained ~1mL of 10microM protein. To determine the catalytic constants the nerve agent paroxoan was used a a substrate. As shown below, hydrolysis product of paroxoan is p-nitrophenol which has a strong absorbance at 400nM (and turns bright yellow). A serial dilution, ranging from 5 millimolar to 5 micromolar, of paroxoan was made in a reaction buffer (100mM HEPES pH=7, 500mM NaCl, 2mM CoCl2). To this a reaction OpdA was added so that it's final concentration was 1nM (dilutions were made in the reaction buffer). At all substrate concentrations no appreciable hydrolysis was observed without enzyme. The rate of hydrolysis with enzyme is shown below, the left hand plot is the full substrate range, and the right hand plot is a zoom in of the lower substrate concentrations:
From the above plot, it obvious that this enzyme efficiently catalysis paroxoan hydrolysis, but does not exhibit the usual Michaelis-Menten dynamics. It can be seen that at high enough concentrations, the enzyme actually undergoes substrate-inhibition, wherein the extra substrate actually slows the enzyme's velocity. When fit to a cononical substrate inhibition curve we obtain the following Kinetic Paramters:
kcat (M-1 s-1): ###
Km (mM): ###
Ksi (mM): ###
These parameters confirm that this is an extremely efficient enzyme, and our kinetic parameters are comparable to previously published data for this enzyme on this substrate. Also, substrate inhibition for this enzyme has been observed previously on similar substrates, so this was not an enitrely surprising results.
Since the OpdA BioBrick was characterized and worked as expected we decided to continue and insert it into the target vector. Unfortinately when OpdA-Target was expressed and purified as described above no observable paroxoan hydrolysis was observed.
Citations
The display/Nano tag was retrieved from Lamla and Erdmann: [http://www.ncbi.nlm.nih.gov/pubmed/14680960]
The secretion tag/prt B was retrieved from Palacios et al: [http://www.ncbi.nlm.nih.gov/pubmed/11157948]
The TEV Recognition Site, ENHLYFQG, was retrieved from [http://mcl1.ncifcrf.gov/waugh_tech/faq/tev.pdf]
 "
"