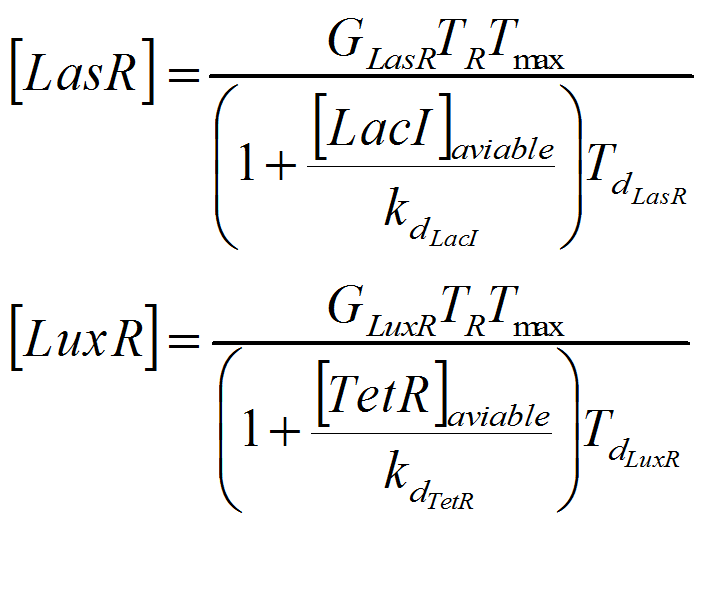Team:IPN-UNAM-Mexico/Modeling/SecondAp
From 2009.igem.org
(New page: {{Template:IPN-UNAM-Mexico}} == 50pxSecond approach: Activator-Inhibitor dynamics on a single cell == Based on the design of the biobricks network we tried to m...) |
m (→The integrated system) |
||
| Line 39: | Line 39: | ||
To build up the entire model we put together all the previous submodels, adding the diffusion of the AHLs and introducing a reporter moduel under the same regulation as lasI. | To build up the entire model we put together all the previous submodels, adding the diffusion of the AHLs and introducing a reporter moduel under the same regulation as lasI. | ||
| - | This system was built on [http://www.celldesigner.org/ Cell Designer ] software to explicitly denoting all the described interactions. Each coefficient of the kinetic rules was gathered from the literature and past years [ | + | This system was built on [http://www.celldesigner.org/ Cell Designer ] software to explicitly denoting all the described interactions. Each coefficient of the kinetic rules was gathered from the literature and past years [https://2007.igem.org/title=ETHZ team wikis]. |
For this model. We tried to be very accurate on the description of our system, and the parameters and the kinetic laws we used are described below. | For this model. We tried to be very accurate on the description of our system, and the parameters and the kinetic laws we used are described below. | ||
| - | + | [[Image:Ex01.png|400px|center]] | |
| + | [[Image:Ex02.png|400px|center]] | ||
| + | [[Image:Ex03.png|400px|center]] | ||
| + | [[Image:Ex04.png|400px|center]] | ||
{| class="prettytable" | {| class="prettytable" | ||
Latest revision as of 23:13, 21 October 2009
 Second approach: Activator-Inhibitor dynamics on a single cell
Second approach: Activator-Inhibitor dynamics on a single cell
Based on the design of the biobricks network we tried to model the behavior it would have within a single cell. To do this we constructed the kinetic laws describing the relationships between the components of our network and IPTG and aTc. We modeled the interactions of the AHLs with their respective targets as allosteric regulators affecting the production rate of the catalyzing enzymes, thus affecting the synthesis of the AHLs. This catalysis was assumed to be a first order kinetics depending only on the concentration of the catalyst.
The diffusion rate of the AHLs was reduced to a first order kinetics considering the membrane area of the bacteria and assuming the complete absence of the AHL outside the cell, therefore depending only on the production within the cell.
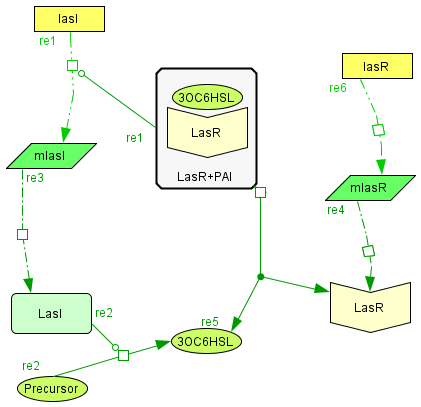
The Activator module
| The activator module is a representation of the kinetiks involved in the same module on the BioBricks design. In this module the transcription and translation of the gene lasR to the protein LasR induces the association with its activator, wich cathalyzes the transcriprion of las. The increasing of its translation finally leads to an increase of the cathalytic production of the activator. |
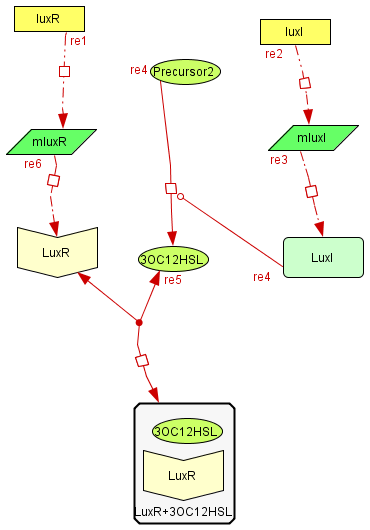
The Inhibitory module
| The inhibitory module consists on two separated genes luxI and luxR producing ther respective proteins. LuxI is the cathalizer of the Lux operon activator, and LuxR is the receptor for this activator. The way this system inhibits is by the binding of the complex LuxR and its activator to a promoter repressible by this complex. |
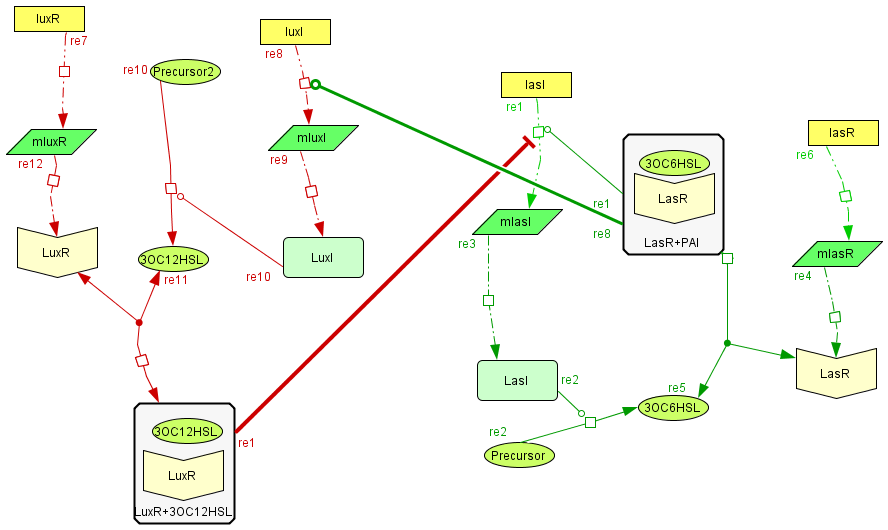
Activator-Inhibitor interaction
| The interaction between the activator and the inhibitor is given by the inhibition on the transcription of lasI and by the induction on the transcription of the luxI gene. This interaction qualitatively behaves as the theoretical Activator-Inhibitor system. |
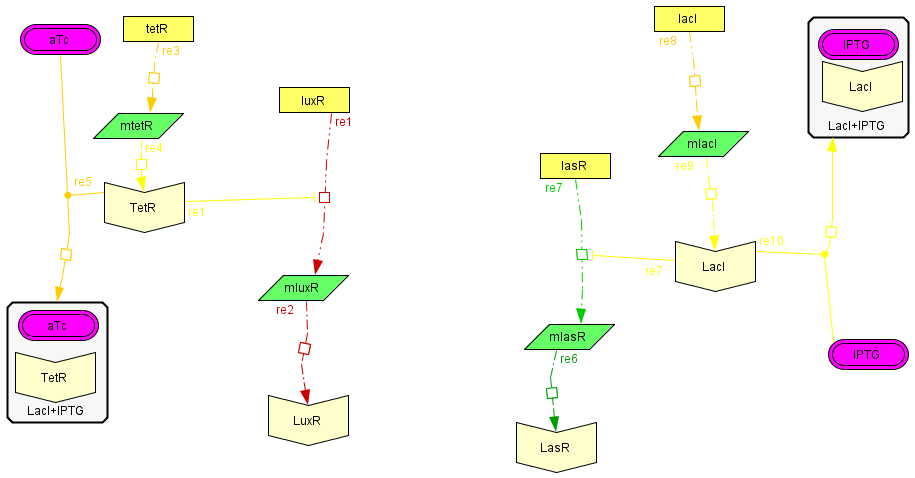
The regulatory modules
| To introduce the regulatory modules on this model we explicitely described the mechanistic description of the Lac and Tet inversor coupled to their regulatory targets: lasR and luxR, respectively;wich are essential components of the activator-inhibytor system. In this way when we add IPTG or aTc the inhibition on the transcription of the genes is removed, increasing the production of the proteins. We also observed that the concentration of the complex (LacI+IPTG or Tet+aTc) eventually reaches to a steady state, allowing us to consider this variables as constant in the following model. The amount of free inhibitor varies depending on the amount of IPTG or aTc we add, so we can regulate the degree of inhibition acting on the activator and inhibitory module |
The integrated system
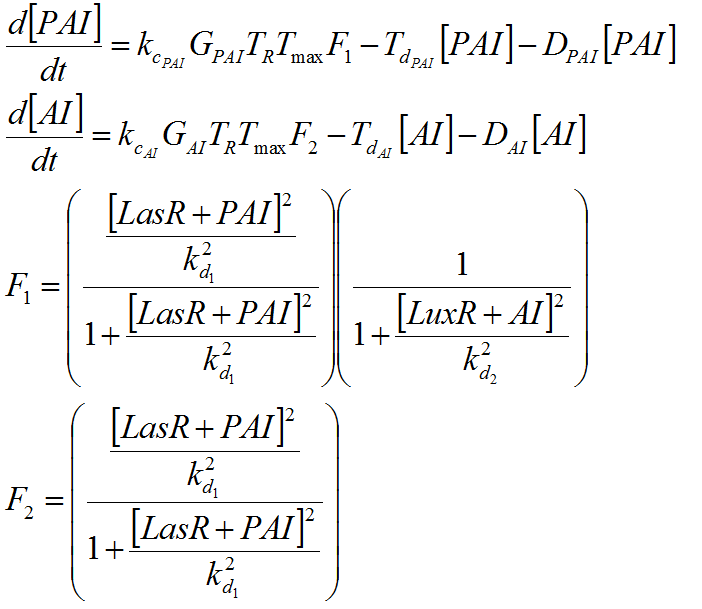
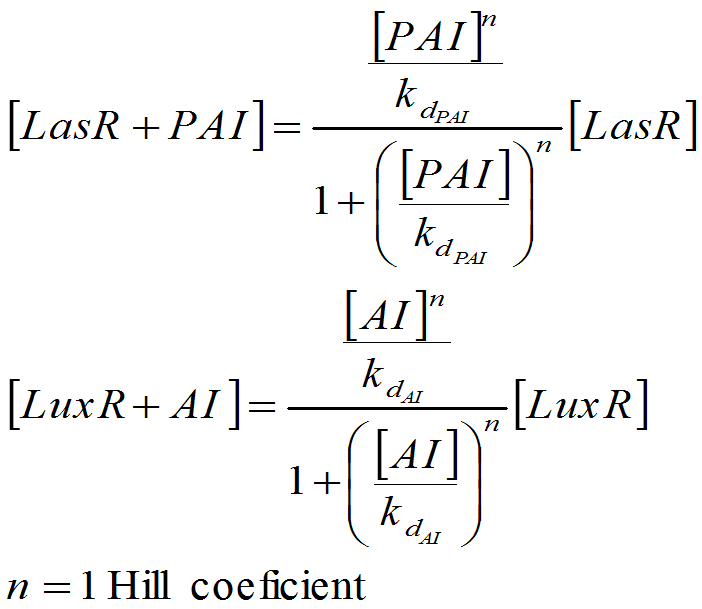
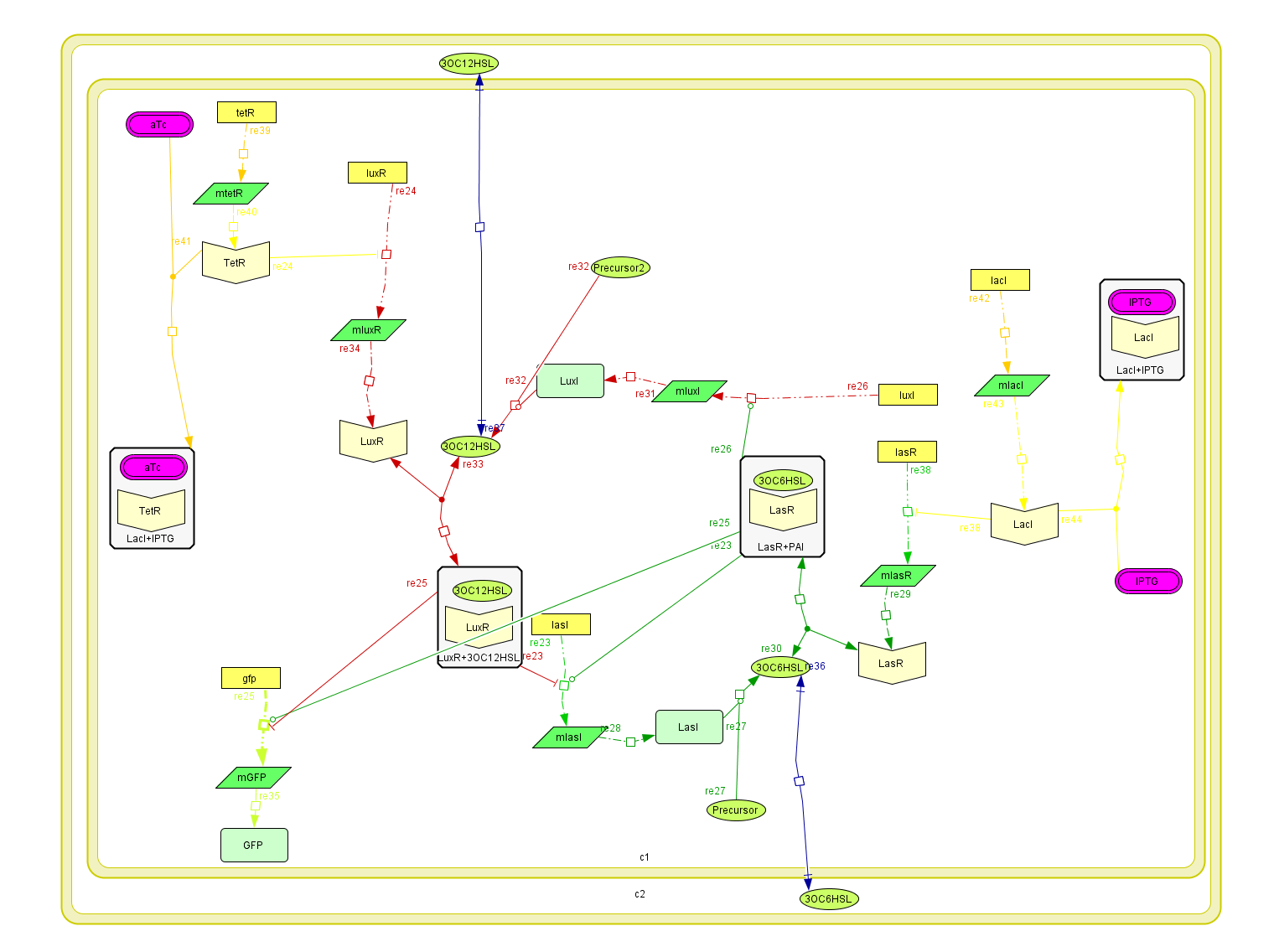
To build up the entire model we put together all the previous submodels, adding the diffusion of the AHLs and introducing a reporter moduel under the same regulation as lasI.
This system was built on [http://www.celldesigner.org/ Cell Designer ] software to explicitly denoting all the described interactions. Each coefficient of the kinetic rules was gathered from the literature and past years team wikis.
For this model. We tried to be very accurate on the description of our system, and the parameters and the kinetic laws we used are described below.
| Constant | Value | Original Units |
| Kc PAI | 0.45
| |
| G PAI | 300
| copies |
| Tmax PAI | 0.01
| mM/h |
| TR PAI | 0.556
| 1/s |
| Td PAI | 0.007
| per protein |
| D PAI | 4.30E-06
| cm2/s |
| kd1 (PAI) | 250
| nM |
| kd2 (AI) | 250
| nM |
| kPAI | 0.5
| uM |
| Kc LasR | 0.45
| |
| G LasR | 300
| copies |
| Tmax LasR | 0.01
| mM/h |
| TR LasR | 0.556
| 1/s |
| kd LacI | 800
| nM |
| kAI | 0.5
| uM |
| Kc LuxR | 0.45
| |
| G LuxR | 300
| copies |
| Tmax LuxR | 0.01
| mM/h |
| TR LuxR | 0.556
| 1/s |
| kd TetR | 179
| pM |
| Kc AI | 0.45
| |
| G AI | 300
| copies |
| Tmax AI | 0.01
| mM/h |
| TR AI | 0.556
| 1/s |
| Td AI | 0.007
| per protein |
| D AI | 6.00E-06
| cm2/s |
| [Iptg] | 1
| mM |
| [aTc] | 3
| uM |
| [LacI] inicial T | 0.008
| mM |
| [TetR] inicial T | 0.008
| mM |
| kd iptg lacI | 893
| pM |
| kd aTc tetR | 1.3
| uM |





 "
"