/Project/Receptor/Antibody
From 2009.igem.org
(→Lpp-OmpA-linker-(receptor)) |
(→PelB-(receptor)-C_IgAP) |
||
| Line 53: | Line 53: | ||
|T|B0010||= | |T|B0010||= | ||
|T|B0012||}} | |T|B0012||}} | ||
| - | [[ | + | [[image:NYMU PelB C-IgAP.jpg|500px|]] |
==== PelB: pelB leader sequence ==== | ==== PelB: pelB leader sequence ==== | ||
A sequence of amino acids which when attached to a protein, directs the protein to the periplasmic membrane of ''E. coli'', where the sequence is removed by pelB peptidase. | A sequence of amino acids which when attached to a protein, directs the protein to the periplasmic membrane of ''E. coli'', where the sequence is removed by pelB peptidase. | ||
Latest revision as of 20:02, 21 October 2009
Contents |
The autotransporter secretion system
Only Receptors that are exposed to the outside of E. coli can catch the virus. E.coli is a gram-negative bacterium that has an inner membrane and an outer membrane, and a periplasmic space between the inner and outer membranes. Because of this, we need the autotransporter secretion system to construct our receptor proteins on E. coli 's surface.
We chose to implement two different autotransporter mechanisms (OmpA and C-IgAP) because we wanted to test and compare their secretion abilities. We suspected that the C-IgAP system is more efficient than the OmpA system.
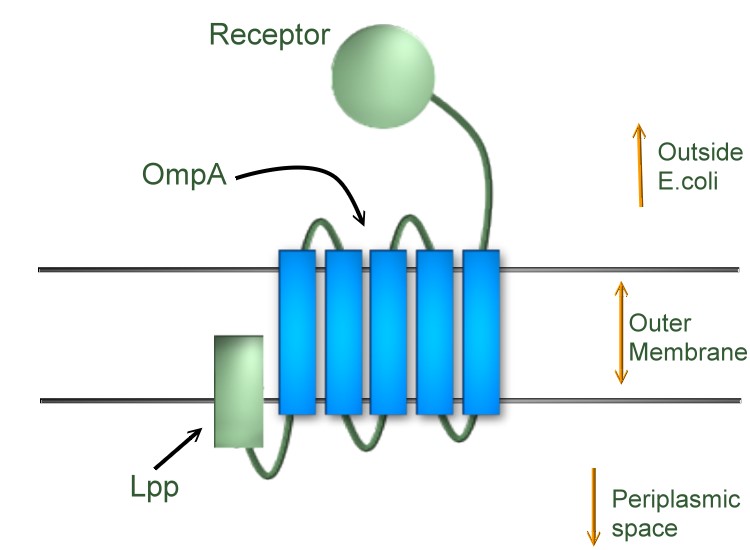
Lpp-OmpA-linker-(receptor)
We used an existing BioBrick part: [http://partsregistry.org/Part:BBa_K103006 BBa_K103006] (iGEM08_Warsaw). The part contains Lpp, OmpA and a linker, so right after the linker, we can fuse with our receptor.
1 catatgaaag ctactaaact ggtactgggc gcggtaatcc tgggttctac 51 tctgctggca ggttgctcca gcaacgctaa aatcgatcag ggaattaacc 101 cgtatgttgg ctttgaaatg ggttacgact ggttaggtcg tatgccgtac 151 aaaggcagcg ttgaaaacgg tgcatacaaa gctcagggcg ttcaactgac 201 cgctaaactg ggttacccaa tcactgacga cctggacatc tacactcgtc 251 tgggtggcat ggtatggcgt gcagacacta aatccaacgt ttatggtaaa 301 aaccacgaca ccggcgtttc tccggtcttc gctggcggtg ttgagtacgc 351 gatcactcct gaaatcgcta cccgtctgga ataccagtgg accaacaaca 401 tcggtgacgc acacaccatc ggcactcgtc cggacaacgg cggaggttct 451 ggaggaggga gctc
Lpp: Lipoprotein signal peptide
When fused with another protein downstream, the signal peptide should anchor the protein to the outer membrane of E. coli, facing the periplasmic space.
OmpA: Outer membrane protein A
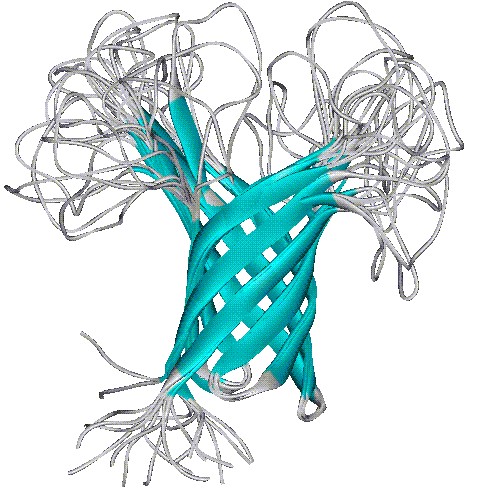
Five transmembrane domains of OmpA, crossing the outer membrane of E. coli, forming a barrel shape.
3D-structure of OmpA
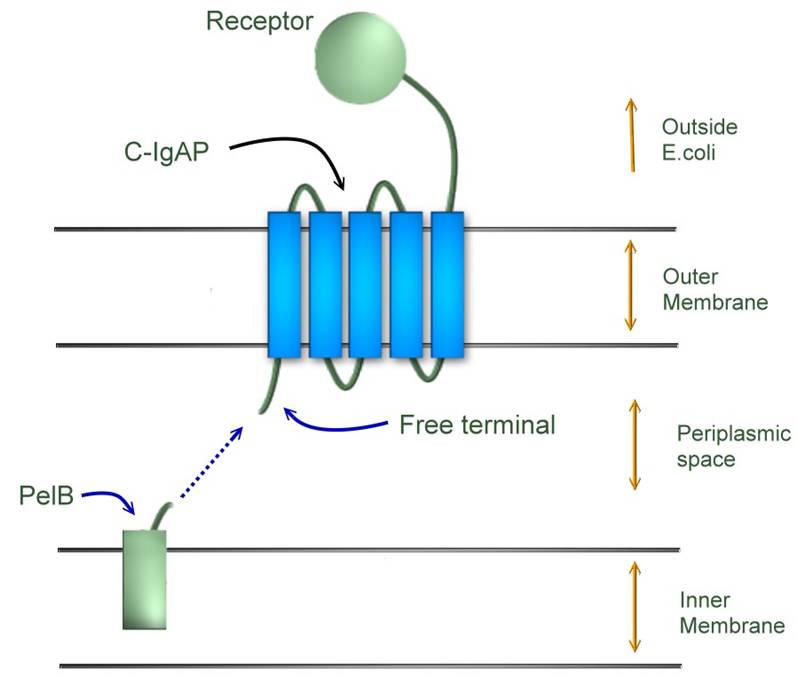
PelB-(receptor)-C_IgAP
- Proteins secreted by Gram-negative bacteria must cross the lipid bilayers of the inner (IM) and outer (OM) membranes as well as the periplasmic space containing the peptidoglycan layer. Insertion of the β-domain into the OM precedes translocation of the passenger protein. Secretion of the N-passenger domain of the β-autotransporters by using a single-chain antibody (scFv) as a reporter of the process
PelB: pelB leader sequence
A sequence of amino acids which when attached to a protein, directs the protein to the periplasmic membrane of E. coli, where the sequence is removed by pelB peptidase.
- [http://partsregistry.org/Part:BBa_J32015 BBa_J32015]
1 atgaaatacc tgctgccgac cgctgctgct ggtctgctgc tcctcgctgc 51 ccagccggcg atggcc
C-IgAP: Carboxy-terminal (β-) domain of Immunoglobulin A1 (IgA1) proteases1
PCR from genome of Neisseria gonorrhoeae FA1090(provide by Dr.Hsing-Ju Wu, Academia sinica/Institute of Biological Chemistry)
- C-IgAP [http://partsregistry.org/Part:BBa_J32018 BBa_J32018]
- The mechanism autotransporters, has been examined using a single-chain antibody (scFv) as a reporter passenger domain to monitor the translocation process. Fusion of a scFv to the IgA protease allowed us to investigate the passage of the chimeric protein through the periplasm, its insertion into the outer membrane and the movement of the N-terminal moiety towards the cell surface. The binding activity of the scFv to its target antigen entirely rely on the formation of disulphide bonds.
- The leader sequence of C-IgAP is cleaved at the membrane by a signal peptidase, releasing the mature polyprotein into the periplasm. When in the periplasm, the beta-domain of the protein inserts into the outer membrane to form a beta-barrel structure. After formation of the beta-barrel pore, the passenger domain is translocated to the cell surface through the pore. Once at the cell surface, several possibilities may occur:
- the protein remains intact as a large polyprotein that has a carboxy-terminal membrane-bound domain and an aminoterminal domain extending into the environment.
- the protein either gains autoproteolytic activity and cleaves itself or it is cleaved by an outer membrane protease, but in this case the passenger domain remains in contact with the cell surface via noncovalent interactions with the b-domain.
- the passenger domain gains its autoproteolytic activity or is cleaved by an outer membrane protease and is released into the environment.
1 aaagctgagg aagaagagca ccgtcaaaca gcccaatccc agccgcaacg 51 ccgcaaacgc cgtgccgcac cgcaggatta tatggcagtt tcccaagacc 101 gtccgaaacg tcgcggacgc agatctactc tgccggcacc gccctcgcca 151 tcatttgatt catcagctta cgcagcaccc agggccttgc ataatccgga 201 ctggtatgaa aatgattatg aagaaatccc cttggatgcg ctggaagatg 251 aagatgtatc cgaatcggtt gacacatcag acaaacagcc tcaagacaat 301 acggaacttc atgaaaaagt tgaggcagtg agtttgcaac caagagccgc 351 gcagccgcga acccaagccg ccgcgcaagc cgatgcagtc agcaccaata 401 ctaactcggc tttatctgac gcaatggcaa gcacgcaatc tatcttgttg 451 gatacaggtg cttcattaac acggcacatt gcacaaaaat cacgcgctga 501 tgccgaaaaa aacagtgttt ggatgtcaaa caccggttat ggccgtgatt 551 atgcttccgc acaatatcgc cggtttagtt cgaaacgcac gcaaacacaa 601 atcggcattg accgcagctt gtccgaaaat atgcagatag gcggagtatt 651 gacttactct gacagtcagc atacttttga tcaggcgggc ggcaaaaata 701 cttttgtgca agccaacctt tatggtaagt attatttaaa tgatgcttgg 751 tatgtggccg gcgatattgg tgcgggcagc ttgagaagcc ggttacaaac 801 gcagcaaaaa gcaaacttta accgaacaag catccaaacc ggccttactt 851 tgggcaatac gctgaaaatc aatcaattcg agattgtccc tagtgcgggt 901 atccgttaca gccgcctgtc atctgcagat tacaagttgg gtgacgacag 951 tgttaaagta agttctatgg cagtgaaaac actaacggcc ggactggatt 1001 ttgcttatcg gtttaaagtc ggcaacctta ccgtaaaacc cttgttatct 1051 gcagcttact ttgccaatta tggcaaaggc ggcgtgaatg tgggcggtaa 1101 atccttcgcc tataaagcag ataatcaaca gcaatattca gcaggcgccg 1151 cgttactgta ccgtaatgtt acattaaacg taaatggcag tattacaaaa 1201 ggaaaacaat tggaaaaaca aaaatccgga caaattaaaa tacagattcg 1251 tttctaa
But this part is not exist in plate. We need to PCR this sequence from Neisseria gonorrhoeae by ourself.
PCR the C-IgAP outer primer gcttctagag AAAGCTGAGGAAGAAGAGC 53deg, CG:47%,10+19=29bp ctgcagcggccgctactagta TTAGAAACGAATCTGTATTTTAATTTGT 52deg, CG:21%, 21+28=49bp
There have some problem:Pst1 cutting site...So we have to mutate these site dierectly.
1 aaagctgagg aagaagagca ccgtcaaaca gcccaatccc agccgcaacg 51 ccgcaaacgc cgtgccgcac cgcaggatta tatggcagtt tcccaagacc 101 gtccgaaacg tcgcggacgc agatctactc tgccggcacc gccctcgcca 151 tcatttgatt catcagctta cgcagcaccc agggccttgc ataatccgga 201 ctggtatgaa aatgattatg aagaaatccc cttggatgcg ctggaagatg 251 aagatgtatc cgaatcggtt gacacatcag acaaacagcc tcaagacaat 301 acggaacttc atgaaaaagt tgaggcagtg agtttgcaac caagagccgc 351 gcagccgcga acccaagccg ccgcgcaagc cgatgcagtc agcaccaata 401 ctaactcggc tttatctgac gcaatggcaa gcacgcaatc tatcttgttg 451 gatacaggtg cttcattaac acggcacatt gcacaaaaat cacgcgctga 501 tgccgaaaaa aacagtgttt ggatgtcaaa caccggttat ggccgtgatt 551 atgcttccgc acaatatcgc cggtttagtt cgaaacgcac gcaaacacaa 601 atcggcattg accgcagctt gtccgaaaat atgcagatag gcggagtatt 651 gacttactct gacagtcagc atacttttga tcaggcgggc ggcaaaaata 701 cttttgtgca agccaacctt tatggtaagt attatttaaa tgatgcttgg 751 tatgtggccg gcgatattgg tgcgggcagc ttgagaagcc ggttacaaac 801 gcagcaaaaa gcaaacttta accgaacaag catccaaacc ggccttactt 851 tgggcaatac gctgaaaatc aatcaattcg agattgtccc tagtgcgggt 901 atccgttaca gccgcctgtc atctgcagat tacaagttgg gtgacgacag 951 tgttaaagta agttctatgg cagtgaaaac actaacggcc ggactggatt 1001 ttgcttatcg gtttaaagtc ggcaacctta ccgtaaaacc cttgttatct 1051 gcagcttact ttgccaatta tggcaaaggc ggcgtgaatg tgggcggtaa 1101 atccttcgcc tataaagcag ataatcaaca gcaatattca gcaggcgccg 1151 cgttactgta ccgtaatgtt acattaaacg taaatggcag tattacaaaa 1201 ggaaaacaat tggaaaaaca aaaatccgga caaattaaaa tacagattcg 1251 tttctaa
modify
1 aaagctgagg aagaagagca ccgtcaaaca gcccaatccc agccgcaacg 51 ccgcaaacgc cgtgccgcac cgcaggatta tatggcagtt tcccaagacc 101 gtccgaaacg tcgcggacgc agatctactc tgccggcacc gccctcgcca 151 tcatttgatt catcagctta cgcagcaccc agggccttgc ataatccgga 201 ctggtatgaa aatgattatg aagaaatccc cttggatgcg ctggaagatg 251 aagatgtatc cgaatcggtt gacacatcag acaaacagcc tcaagacaat 301 acggaacttc atgaaaaagt tgaggcagtg agtttgcaac caagagccgc 351 gcagccgcga acccaagccg ccgcgcaagc cgatgcagtc agcaccaata 401 ctaactcggc tttatctgac gcaatggcaa gcacgcaatc tatcttgttg 451 gatacaggtg cttcattaac acggcacatt gcacaaaaat cacgcgctga 501 tgccgaaaaa aacagtgttt ggatgtcaaa caccggttat ggccgtgatt 551 atgcttccgc acaatatcgc cggtttagtt cgaaacgcac gcaaacacaa 601 atcggcattg accgcagctt gtccgaaaat atgcagatag gcggagtatt 651 gacttactct gacagtcagc atacttttga tcaggcgggc ggcaaaaata 701 cttttgtgca agccaacctt tatggtaagt attatttaaa tgatgcttgg 751 tatgtggccg gcgatattgg tgcgggcagc ttgagaagcc ggttacaaac 801 gcagcaaaaa gcaaacttta accgaacaag catccaaacc ggccttactt 851 tgggcaatac gctgaaaatc aatcaattcg agattgtccc tagtgcgggt 901 atccgttaca gccgcctgtc atTtgcagat tacaagttgg gtgacgacag 951 tgttaaagta agttctatgg cagtgaaaac actaacggcc ggactggatt 1001 ttgcttatcg gtttaaagtc ggcaacctta ccgtaaaacc cttgttatct 1051 gcCgcttact ttgccaatta tggcaaaggc ggcgtgaatg tgggcggtaa 1101 atccttcgcc tataaagcag ataatcaaca gcaatattca gcaggcgccg 1151 cgttactgta ccgtaatgtt acattaaacg taaatggcag tattacaaaa 1201 ggaaaacaat tggaaaaaca aaaatccgga caaattaaaa tacagattcg 1251 tttctaa
mutant primer 1. C-IgAP_mt1_F:gcctgtcatTtgcagattacaag C-IgAP_mt1_R:cttgtaatctgcaAatgacaggc 2. C-IgAP_mt2_F:ctgcCgcttactttgcc C-IgAP_mt2_R:ggcaaagtaagcGgcag
Experiments
PCR the C-IgAP from genome of Neisseria gonorrhoeae FA1090(Pfu)
Reference
- [http://www3.interscience.wiley.com/journal/119079102/abstract?CRETRY=1&SRETRY=0 Probing secretion and translocation of a β-autotransporter using a reporter single-chain Fv as a cognate passenger domain]
- [http://jvi.asm.org/cgi/content/abstract/77/24/13396 Neutralization of Enteric Coronaviruses with Escherichia coli Cells Expressing Single-Chain Fv-Autotransporter Fusions]
- [http://www.sciencedirect.com/science?_ob=ArticleURL&_udi=B6TD0-3TX4KWY-H&_user=1576506&_coverDate=09%2F01%2F1998&_fmt=full&_orig=search&_cdi=5184&view=c&_acct=C000053839&_version=1&_urlVersion=0&_userid=1576506&md5=d3089b24652222e68aaf9dfcf1d44bda&ref=full The great escape: structure and function of the autotransporter proteins]
- [http://www.nature.com/emboj/journal/v21/n9/abs/7594432a.html Export of autotransported proteins proceeds through an oligomeric ring shaped by C-terminal domains]
 "
"