Team:DTU Denmark/introduction private securkey Dhjg1mab2ak47
From 2009.igem.org
| Line 217: | Line 217: | ||
<p align="justify"><i><b>Gene design and redox regulation</b><br> | <p align="justify"><i><b>Gene design and redox regulation</b><br> | ||
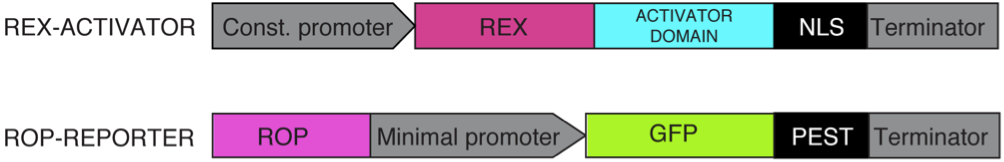
<b>A:</b> The Rex gene will be fused to an activator domain and will be transcribed constitutively leading to constant concentration of the sensor in the cell. The ROB sequence and a minimal promoter is followed by a reporter gene, which will only be transcribed when the Rexivator complex is bound to the promoter. <b>B:</b> The Rexivator only binds the ROB DNA sequence under the condition of having NAD+ bound. Under these circumstances the fused activator domain summons the RNA polymerase and the reporter gene will be transcribed</i></p><br> | <b>A:</b> The Rex gene will be fused to an activator domain and will be transcribed constitutively leading to constant concentration of the sensor in the cell. The ROB sequence and a minimal promoter is followed by a reporter gene, which will only be transcribed when the Rexivator complex is bound to the promoter. <b>B:</b> The Rexivator only binds the ROB DNA sequence under the condition of having NAD+ bound. Under these circumstances the fused activator domain summons the RNA polymerase and the reporter gene will be transcribed</i></p><br> | ||
| + | |||
| + | <font size="3"><b>Our synthetic biology project: The Redoxilator</b></font><br> | ||
| + | <p align="justify">To achieve a system that senses changing levels in the NAD<sup>+</sup>/NADH ratio in the eukaryote <i>S. cerevisiae</i>, the gene encoding the Rex protein will be fused to a yeast activator domain, resulting in a new synthetic protein: the Redoxilator. The ROP sequence - the DNA binding site Rex can bind to - will be inserted into a yeast promoter, resulting in a promoter activated by the Redoxilator.</p> | ||
| + | |||
| + | The Redoxilator system consists of two synthetic genes. One of the genes will be designed to code for a synthetic protein that activates transcription of the second gene only at a high NAD+/NADH ratio. | ||
| + | |||
| + | <p align="justify">A certain NAD<sup>+</sup>/NADH ratio will activate the Redoxilator to recognize the ROB promoter resulting in transcription of a downstream gene. In this way the ROB promoter and the Redoxilator comprises the complete sensing system. The system can be coupled to the expression of virtually any gene of interest; making transcription solely dependent on the ratio of NAD<sup>+</sup>/NADH in the cell. In our iGEM project, the system will be used for two selected applications considered highly relevant: i) in vivo monitoring of NAD<sup>+</sup>/NADH in yeast, and ii) NAD<sup>+</sup>/NADH ratio regulated production of yeast products in chemostat processes.</p><br> | ||
| + | |||
<p><b>Design specifications</b><b> </b><br /> | <p><b>Design specifications</b><b> </b><br /> | ||
| Line 289: | Line 297: | ||
</table> | </table> | ||
<br><br> | <br><br> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
</font> | </font> | ||
Revision as of 22:20, 18 October 2009

| Home | The Team | The Project | Parts submitted | Modelling | Notebook |
|
The redoxilator - Introduction - Results - Applications and perspectives The USERTM assembly standard - Principle - Proof of concept - Manual USERTM fusion primer design software - Abstract - Instructions - Output format |
The project Genetic design We have successfully designed and physically constructed a genetic system that couples the intracellular NAD+/NADH level to the gene expression of a reporter protein. The system has potentially many applications including in vivo online monitoring of the redox poise, production optimization and cancer research with yeast as a model organism (see Applications). We have demonstrated that the system functions as expected in S. cerevisiae. The NAD+/NADH ratio is sensed by a system originating in Streptomyces coellicolor. In S. coellicolor the protein REX is a repressor and controls the gene expression of multiple genes by recognizing and binding to a specific DNA-sequence termed ROP (Rex operator). NAD+ and NADH compete to associate with Rex, but only a REX:NAD+ association can bind the ROP DNA-sequence (Brekasis and Paget, 2003). In S. coellicolor REX DNA binding represses expression of target genes, by physically hindering RNA-polymerases from binding the promoter. As the transcription machinery of eukaryotes is different and more complicated, there is no guarantee that repression will be effective in eukaryotes. REX has therefore been fused to a eukaryotic transcriptional activator, a widely used technique applied for the investigation of the GAL proteins and other systems (Sadowski et al. 1988). The REX-activator fusion-protein is able to bind the ROB sequence placed upstream of a minimal eukaryotic promoter that only supports transcription upon activation, and is devoid of regulatory motifs.
Gene design and redox regulation Our synthetic biology project: The Redoxilator To achieve a system that senses changing levels in the NAD+/NADH ratio in the eukaryote S. cerevisiae, the gene encoding the Rex protein will be fused to a yeast activator domain, resulting in a new synthetic protein: the Redoxilator. The ROP sequence - the DNA binding site Rex can bind to - will be inserted into a yeast promoter, resulting in a promoter activated by the Redoxilator. The Redoxilator system consists of two synthetic genes. One of the genes will be designed to code for a synthetic protein that activates transcription of the second gene only at a high NAD+/NADH ratio.A certain NAD+/NADH ratio will activate the Redoxilator to recognize the ROB promoter resulting in transcription of a downstream gene. In this way the ROB promoter and the Redoxilator comprises the complete sensing system. The system can be coupled to the expression of virtually any gene of interest; making transcription solely dependent on the ratio of NAD+/NADH in the cell. In our iGEM project, the system will be used for two selected applications considered highly relevant: i) in vivo monitoring of NAD+/NADH in yeast, and ii) NAD+/NADH ratio regulated production of yeast products in chemostat processes. Design specifications
The genetic elements and the requirements they need to fulfill are listed in the following table. The detailed description of the used genetic elements will not be made not publicly available due to IP rights. Description and requirements of genetic elements.
|
The yeast metabolic cycle It has recently been shown by Tu et al. and Klevecz et al. that the expression of at least half of the genes monitored on a standard yeast gene chip will oscillate in a coordinated manner when grown under glucose limited conditions. The cells will shift between oxidative and reductive metabolism in a synchronized metabolic cycle with three phases: oxidative, reductive/building and reductive/ charging. As oxygen will only be consumed in the oxidative phase, the dissolved oxygen will oscillate. Many metabolites and cofactors including NADH and NAD+ will also oscillate during this cycle as NADH is converted to NAD+ when oxygen is consumed. |
| Comments or questions to the team? Please Email us -- Comments of questions to webmaster? Please Email us |
 "
"