Team:Wash U/Biological Parts
From 2009.igem.org
(→Modeling the Gene Regulatory Network) |
|||
| Line 136: | Line 136: | ||
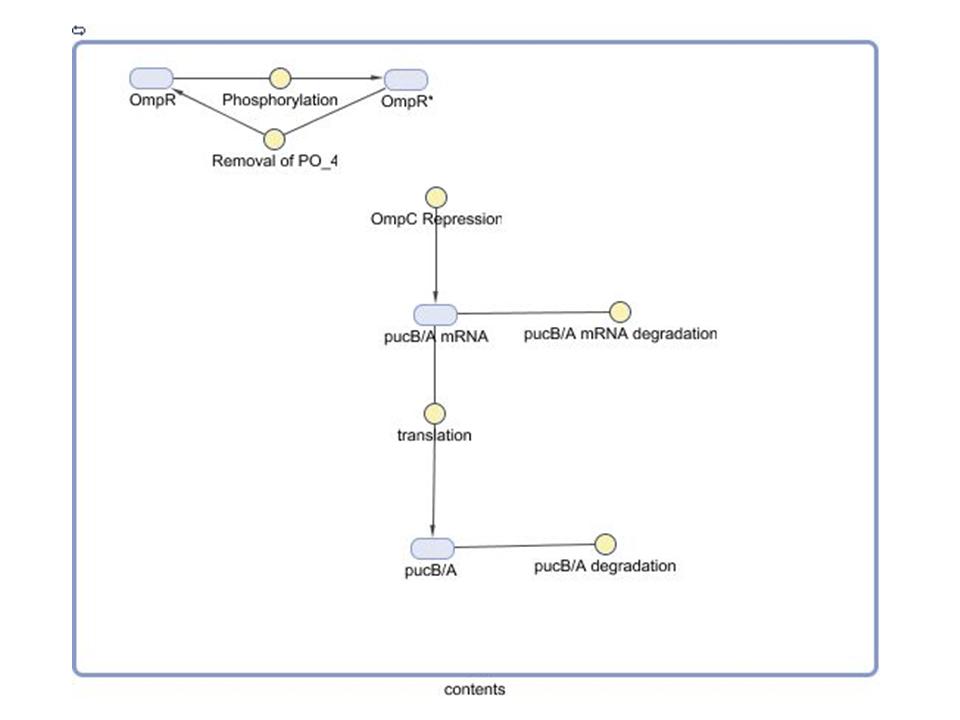
| - | == Modeling the Gene Regulatory Network == | + | == '''Modeling the Gene Regulatory Network''' == |
<font size="2"> | <font size="2"> | ||
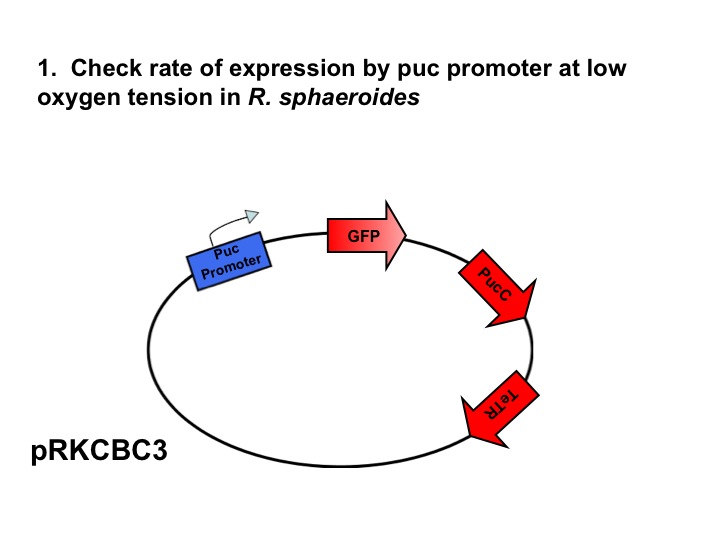
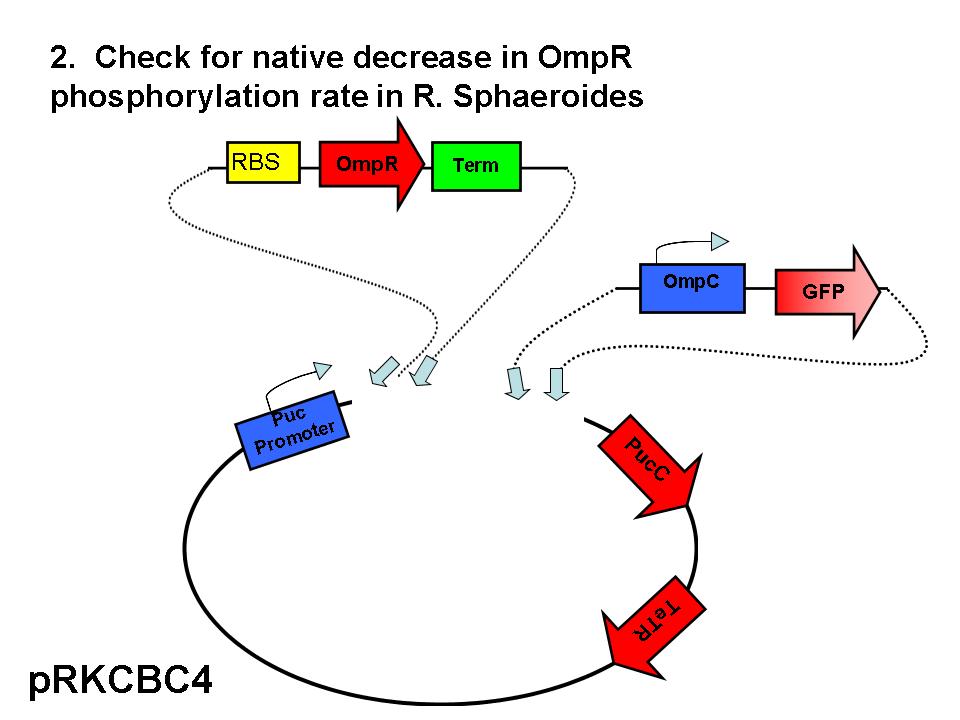
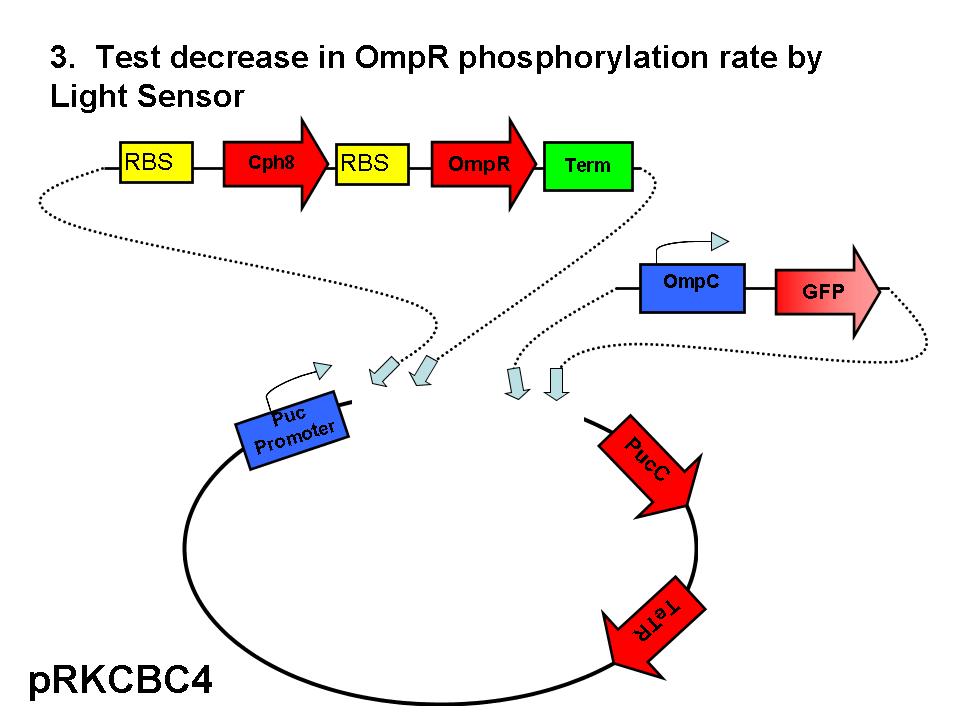
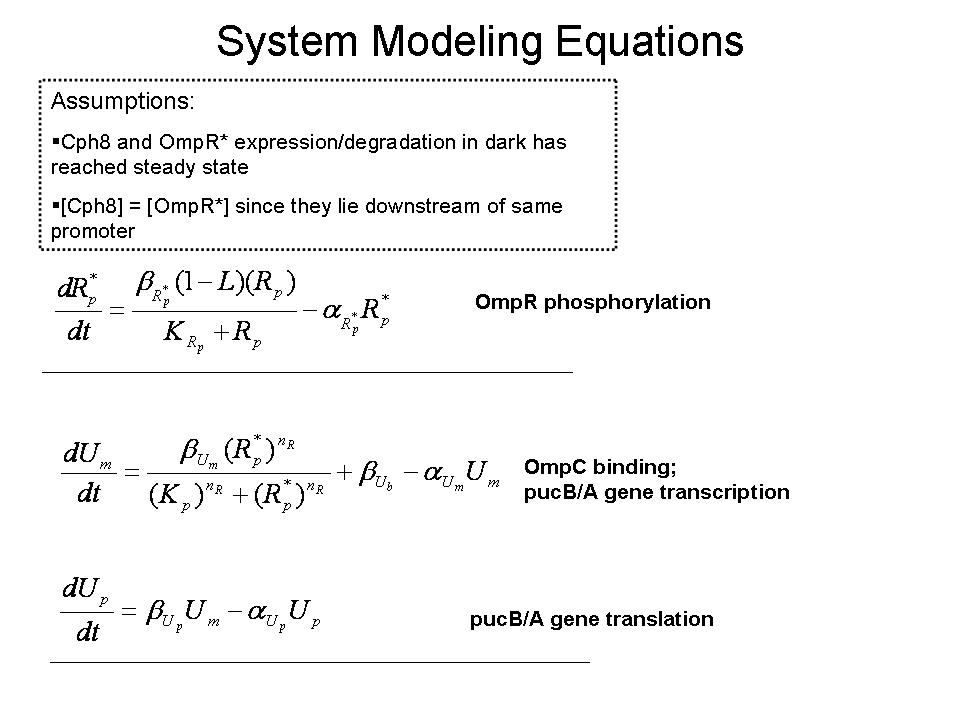
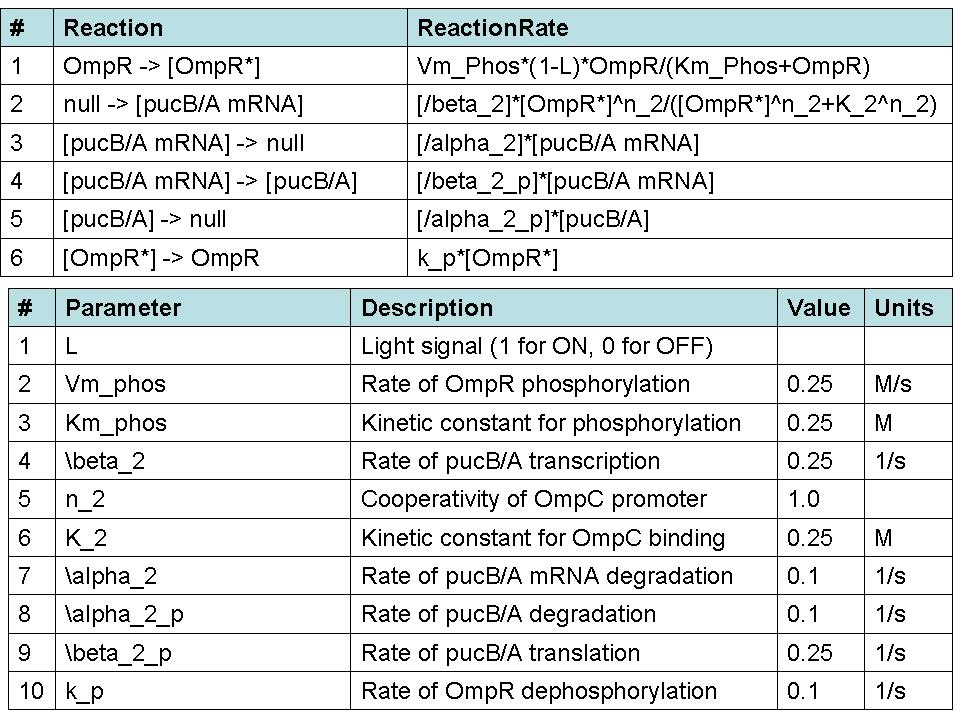
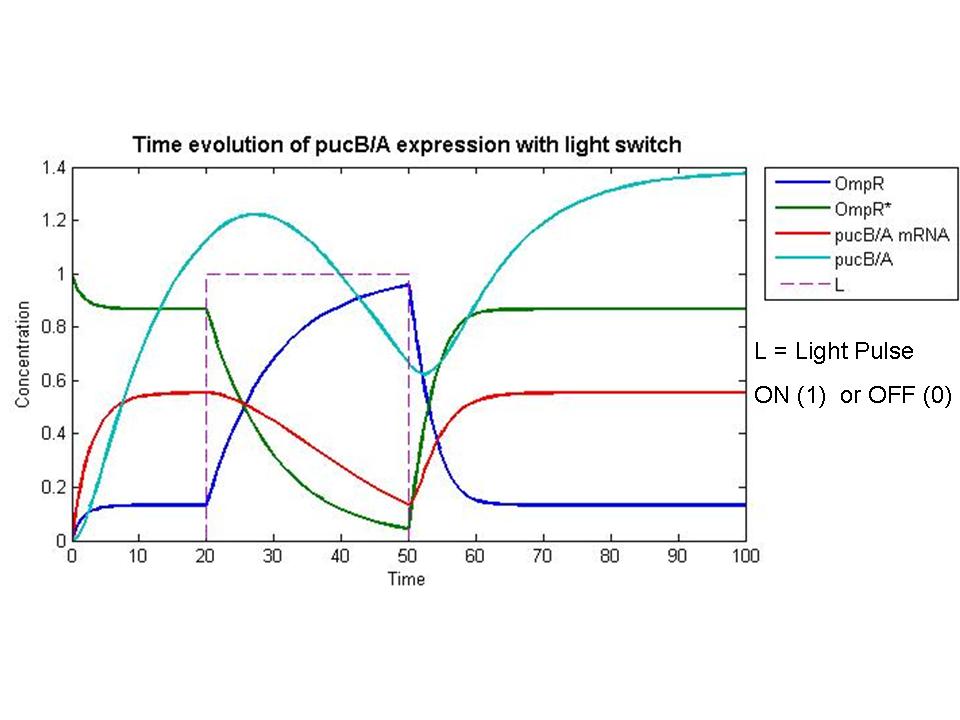
Our group seeks to assess the optimality of the synthetic system that modulates pucBA gene expression and LH2 complex assembly in ''Rhodobacter sphaeroides''. Here we employ a mathematical model of this system to generate predictions about the behavior of the active system in response to light input. Features of the system that the model may help investigate include the time scale of response to light signals, the robustness of the system in response to fluctuations in light intensity, and the translation between changes in gene expression and the absorbance spectrum of the engineered cells. | Our group seeks to assess the optimality of the synthetic system that modulates pucBA gene expression and LH2 complex assembly in ''Rhodobacter sphaeroides''. Here we employ a mathematical model of this system to generate predictions about the behavior of the active system in response to light input. Features of the system that the model may help investigate include the time scale of response to light signals, the robustness of the system in response to fluctuations in light intensity, and the translation between changes in gene expression and the absorbance spectrum of the engineered cells. | ||
Revision as of 22:36, 6 July 2009

 "
"