|
|
Introduction
Implementing a "Timer" Function!
- Our project is to make a sort of "timer", where gene activation is triggered after a certain time.
- Our approach to this goal is to create a series of transcription factors (TFs) that can be activated by the same inducer molecule but with a different sensitivity. In an environment where the inducer concentration gradually levels up, these TFs switches on one by one according to the order of sensitivity.
- We believe such collection of such TF variants would be useful for the timing control in biological function at will, and thereby contribute to the synthetic biology & iGEM community.
- Using a set of TF variants, we aimed to draw an "animated picture": a picture that pop up one by one.
|
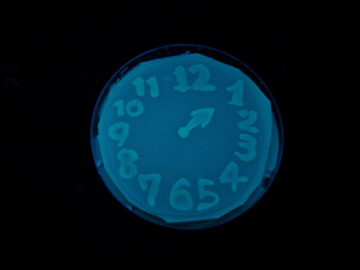
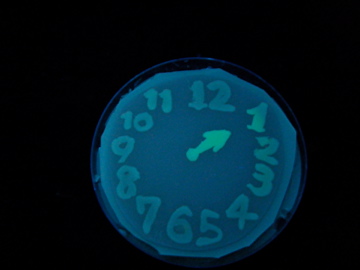
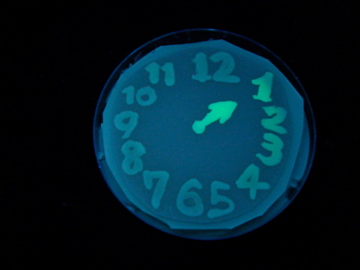
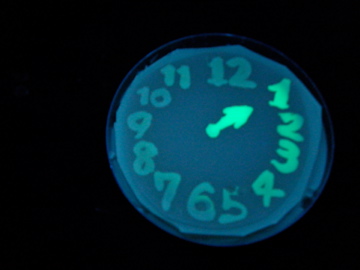
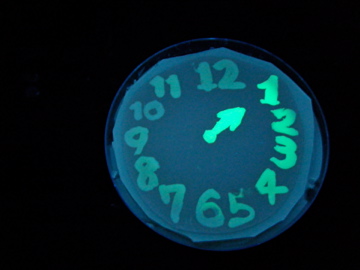
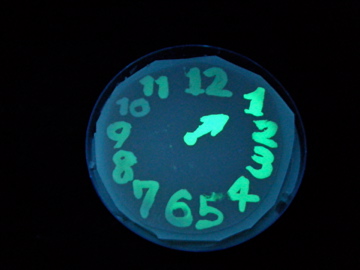
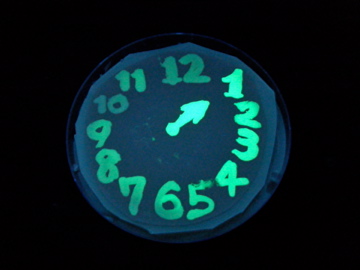
Fig. 1 Completion drawing of our bacterial timer
|
Project Design
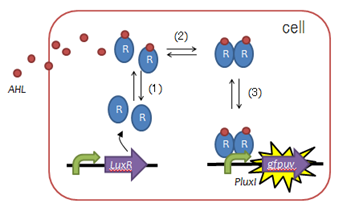 Fig. 2 Function of LuxR :(1)LuxR protein generation, (2)AHL binding domain, (3)dimerization, (4)DNA binding domain - To create such a timer, we utilized LuxR, a protein used to mediate cell-cell communication during quorum sensing, as shown in the figure on the left.
- By inserting point-mutations in the luxR gene, we sought to create LuxR proteins requiring a variety of lengths of time to activate downstream transcription.
Experiments, Results & Discussion
Experiments
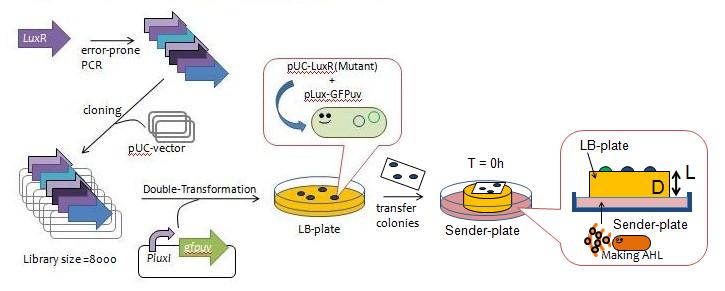 Fig.3 Y Directed evolution to get some delayed-LuxR mutants - We created a mutant library of LuxR (size = 8000) with the use of error-prone PCR and incorporated the resulting coding regions into expression vectors.
- The vectors were then transformed into a strain of E.coli JW1226 cells harboring plux-gfp and grew approximately 200 colonies.
- The colonies were lifted off the agar plate with a nitrocellulose filter and transfered to a plate containing AHL in order to observe the evolution of GFP fluorescence over time (Fig. 4).
- We selected 13 colonies that were slow to display fluorescence, which we designated as delayed-LuxR mutants.
 Fig. 4 display fluorescence
Characterization of LuxR Mutants
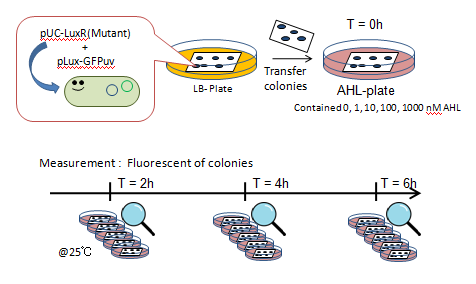 Fig.5 Characterization of LuxR Mutants Experiments (Fig.5)
- JW1226 bacterial strains co-transformed with plasmids containing either wild-type or mutant (13 variants) LuxR and pLux-GFP were cultured at 37℃ for 12h.
- Using a pin, the culture was transfered to a nitrocellulose filter, placed on an agar plate, cultured at 37℃ for an additional 12h.
- The nitrocellulose filter was then transfered to a solid medium containing 0, 1, 10, 100, or 1000 nM AHL.
- Designating this the start point, we observed fluorescence through the naked eye at 30 minute intervals for 4 hours under UV light of 365nm (Funakoshi UVGL-58 at long wave mode).
Switching Score at near-end point (6h)
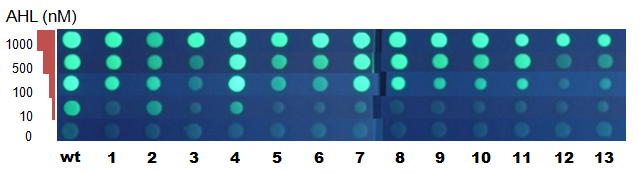 Fig.6 Fluorescence photos at 6h after incubation - Mutant LuxRs showed different threshold in AHL concentration required for switching.
- Many (not all) of the delayed mutants isolated in our screening turned out to be with lower sensitivity to AHL compared with WT.
- Threshold value of [AHL] was similar (- 10nM) to WT for LuxRs (#2, #4). Most others requied about 10x higher concentration for switching. Mutants #1, #12, and #13 required 100x higher concentration of AHL (1uM).
- The result indicates that one of the major 'solution' to make apparent delay in color development was to down-tune the sensitivity of LuxRs.
Time-course color development on the plate with fixed (100 nM) conc. of AHL.
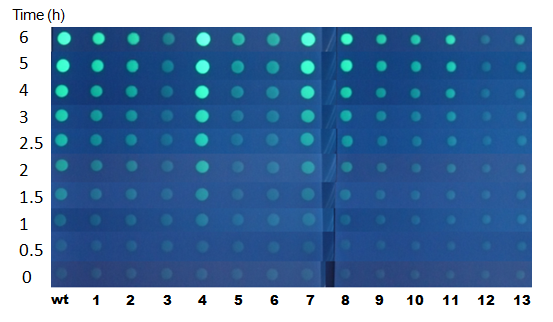 Fig.7 fluorescence photos on 100 nM AHL medium plate
- Upon full induction, the intensity of GFP fluorescence reaches plateau in 90 mins.
- At the end point of this experiment (6h after induction), clone #1 and #2 showed similar apparent fluorescence with WT. However, they were significantly slower in color development. Note that the switching threshold of clone #2 appears almost the same with WT at the end point. This indicates that this clone is as sensitive to AHL as wildtype but less efficient in activating the LuxP promoter.
- Many others exhibited slower color development, and they did not reach the maximum level of fluorescence.
}We have conducted this experiments in various other concentration of AHL. The difference in the rate of color development was maximum on the plate with 100 nM AHL.
- Further analysis (transfer curves in different time points) is currently ongoing. We hope it would provide more information on the molecular basis on how they achieve the delayed phenotype.
Genotypes
- The following lists where point-mutations were incorporated into the luxR gene.
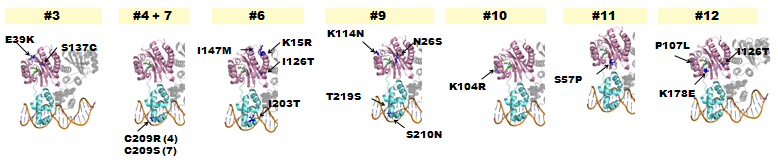 Fig. 8 Hypothetical positions of residues in TraR corresponding to those found to modulate acyl-HSL specificity in LuxR(Nassser et al., 2007). The crystal structure of the LuxR homologue TraR (PDB 1L3L) has been determined (Zhang et al., 2002). Pink represents AHL-binding domain and cyan represents DNA-binding domain. - A delayed GFP expression (in comparison to wild-type LuxR) phenotype was observed from colonies housing genes in which either the AHL-binding domain or the DNA binding domain had been mutated.
Demonstration
- Double-Transformed (pUC-LuxR Mutants series-ColE1 and pLux-GFPuv-P15A) and cultured on LB-Amp-Cm-plate. (37°C,12h)
- Pick and cultured in 10mL of LB-Amp-Cm (37°C,12h).
- Drawing picture with those liquid culture on nitrocellulose Filter.
- Those nitrocellulose Filter placed on LB-Amp-Cm-plate.
- Cultured at 37°C, 12h.
- Transfer pictures on special plate.
- Exposed to UV (365nm) light once every 30 minutes to observe GFP fluorescence.
- For the determination of best condition for E coli painting, Click here!
Conclusions
- By using error-prone PCR, we have created a LuxR library.
- With a simple and convenient screening method, we have isolated various LuxR mutants which confer delayed switching behavior in GFP signals.
- By introducing these LuxR variants together with reporter genes (such as GFP) under the control of Lux promoter, we created bacteria 'ink's that develop their color with unique delay-time.
- We conducted painting with these bacteria inks thereby created animated pictures.
- Characterization of LuxR mutants is ongoing. But our preliminary data showed that some of the variants turned out to be the one with less sensitivity to AHLs, and others seemed to be as sensitive as wild-type LuxR but seemed less efficient somewhere in the downstream process.
- We created variety of Biobricks during the course of this projects. Some of them are characterized and sent to the HQ, and many more are almost ready for shipping!
|

 "
"