Team:TorontoMaRSDiscovery/Modeling
From 2009.igem.org
Contents |
Objective
Model
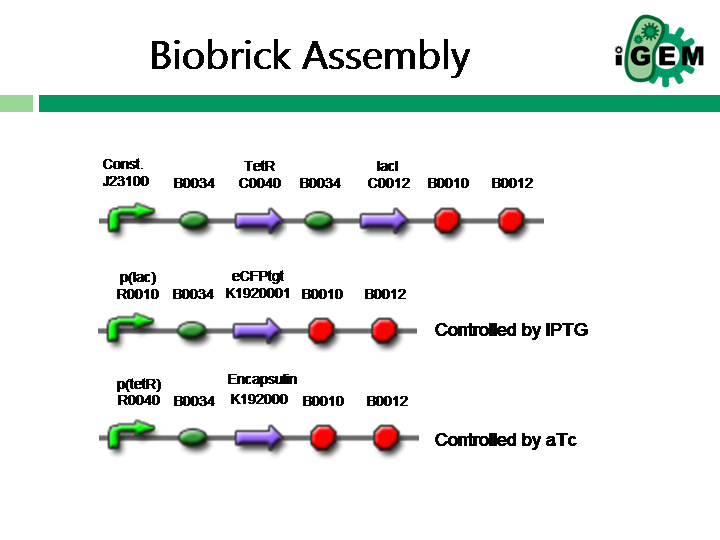
The construct that needs to be modelled is shown below. Our submission this year may/may not have all of the parts assebled.
The construct is represented by the reactions shown below Click here for excel file. The reaction framework was mainly generated using the [http://synbioss.sourceforge.net/ SynBioSS Designer (a great modelling tool!)], which also provides "default" parameter values. These parameter values have been gathered from various literature [http://neptune.cems.umn.edu/designer/designer_defaults.pdf sources], and should be taken as approximations on the appropriate order of magnitude. All units were taken to be (1 / molarity^n-1 * s), where n is the order of the reaction. For example, for the reaction 2 lacI --> lacI2, n = 2, thus the rate constant has the units 1/molality*s.
The pathways for the expressed proteins, eCFPt and Encapsulin, need to be modelled in more detail. For example, in this basic model eCFPt is degraded via a first order reaction and Encapsulin makes a dimer complex. However, in reality fluorescent proteins have been shown to degreade via Micheales-menton kinetics and Encapsulin is thought to make a complex of about 60 monomers. These issues will be addressed in future work.
| Reaction | Rate Constant |
| Constitutive Expression | |
| --> lacl4 | 1E-10 |
| --> tetR2 | 1E-10 |
| Multimerization | |
| 2 lacI --> lacI2 | 1.00E+09 |
| lacI2 --> 2 lacI | 10 |
| 2 lacI2 --> lacI4 | 1.00E+09 |
| lacI4 --> 2 lacI2 | 10 |
| 2 tetR --> tetR2 | 1.00E+09 |
| tetR2 --> 2 tetR | 10 |
| 2 Enc --> Enc2 | 100 |
| Encapsulation | |
| Enc2 + eCFPt --> Enc2:eCFPt | 100 |
| Transcription | |
| RNAp + BBa_R0010 + LacI_binding_site --> RNAp:BBa_R0010:LacI_binding_site | 10000000 |
| RNAp:BBa_R0010:LacI_binding_site --> RNAp + BBa_R0010 + LacI_binding_site | 0.057 |
| RNAp:BBa_R0010:LacI_binding_site --> RNAp:BBa_R0010:LacI_binding_site* | 0.1 |
| RNAp:BBa_R0010:LacI_binding_site* --> RNAp:DNA_eCFPt + BBa_R0010 + LacI_binding_site | 30 |
| RNAp:DNA_eCFPt --> RNAp + mRNA_eCFPt | 0.035 |
| RNAp + BBa_R0040 + TetR_1 + TetR_2 --> RNAp:BBa_R0040:TetR_1:TetR_2 | 10000000 |
| RNAp:BBa_R0040:TetR_1:TetR_2 --> RNAp + BBa_R0040 + TetR_1 + TetR_2 | 0.057 |
| RNAp:BBa_R0040:TetR_1:TetR_2 --> RNAp:BBa_R0040:TetR_1:TetR_2* | 0.1 |
| RNAp:BBa_R0040:TetR_1:TetR_2* --> RNAp:DNA_Enc + BBa_R0040 + TetR_1 + TetR_2 | 30 |
| RNAp:DNA_Enc --> RNAp + mRNA_Enc | 0.0375 |
| Translation | |
| rib + mRNA_eCFPt --> rib:mRNA_eCFPt | 100000 |
| rib:mRNA_eCFPt --> rib:mRNA_eCFPt_1 + mRNA_eCFPt | 33 |
| rib:mRNA_eCFPt_1 --> rib + eCFPt | 0.1154 |
| rib + mRNA_Enc --> rib:mRNA_Enc | 100000 |
| rib:mRNA_Enc --> rib:mRNA_Enc_1 + mRNA_Enc | 33 |
| rib:mRNA_Enc_1 --> rib + Enc | 0.1234 |
| Protein-DNA | |
| lacI4 + nsDNA --> lacI4:nsDNA | 1000 |
| lacI4:nsDNA --> lacI4 + nsDNA | 1.6225 |
| lacI4 + LacI_binding_site --> lacI4:LacI_binding_site | 1.00E+09 |
| lacI4:LacI_binding_site --> lacI4 + LacI_binding_site | 0.005 |
| tetR2 + nsDNA --> tetR2:nsDNA | 1000 |
| tetR2:nsDNA --> tetR2 + nsDNA | 1.6225 |
| tetR2 + TetR_1 --> tetR2:TetR_1 | 1.00E+09 |
| tetR2:TetR_1 --> tetR2 + TetR_1 | 0.005 |
| tetR2 + TetR_2 --> tetR2:TetR_2 | 1.00E+09 |
| tetR2:TetR_2 --> tetR2 + TetR_2 | 0.005 |
| Protein - Effector - DNA | |
| lacI4 + IPTG --> lacI4:IPTG | 50000000 |
| lacI4:IPTG --> lacI4 + IPTG | 0.1 |
| lacI4:IPTG + LacI_binding_site --> lacI4:IPTG:LacI_binding_site | 1.00E+09 |
| lacI4:IPTG:LacI_binding_site --> lacI4:IPTG + LacI_binding_site | 0.7 |
| lacI4:LacI_binding_site + IPTG --> lacI4:IPTG:LacI_binding_site | 1000000 |
| lacI4:IPTG:LacI_binding_site --> lacI4:LacI_binding_site + IPTG | 0.4 |
| lacI4:IPTG + nsDNA --> lacI4:IPTG:nsDNA | 1000 |
| lacI4:IPTG:nsDNA --> lacI4:IPTG + nsDNA | 1.6225 |
| lacI4:nsDNA + IPTG --> lacI4:IPTG:nsDNA | 1000 |
| lacI4:IPTG:nsDNA --> lacI4:nsDNA + IPTG | 1.6225 |
| tetR2 + aTc --> tetR2:aTc | 50000000 |
| tetR2:aTc --> tetR2 + aTc | 0.1 |
| tetR2:aTc + TetR_1 --> tetR2:aTc:TetR_1 | 1.00E+09 |
| tetR2:aTc:TetR_1 --> tetR2:aTc + TetR_1 | 0.7 |
| tetR2:TetR_1 + aTc --> tetR2:aTc:TetR_1 | 1000000 |
| tetR2:aTc:TetR_1 --> tetR2:TetR_1 + aTc | 0.4 |
| tetR2:aTc + TetR_2 --> tetR2:aTc:TetR_2 | 1.00E+09 |
| tetR2:aTc:TetR_2 --> tetR2:aTc + TetR_2 | 0.7 |
| tetR2:TetR_2 + aTc --> tetR2:aTc:TetR_2 | 1000000 |
| tetR2:aTc:TetR_2 --> tetR2:TetR_2 + aTc | 0.4 |
| tetR2:aTc + nsDNA --> tetR2:aTc:nsDNA | 1000 |
| tetR2:aTc:nsDNA --> tetR2:aTc + nsDNA | 1.6225 |
| tetR2:nsDNA + aTc --> tetR2:aTc:nsDNA | 1000 |
| tetR2:aTc:nsDNA --> tetR2:nsDNA + aTc | 1.6225 |
| Degredation | |
| lacI4 --> | 0.000289 |
| tetR2 --> | 0.000289 |
| mRNA_eCFPt --> | 0.0015 |
| mRNA_Enc --> | 0.0015 |
| eCFPt --> | 0.000289 |
| Enc2 --> | 0.000289 |
| Enc2:eCFPt --> | 0.000289 |
| Dilution | |
| lacI4:nsDNA --> nsDNA | 0.000193 |
| tetR2:nsDNA --> nsDNA | 0.000193 |
| lacI4:IPTG --> IPTG | 0.000289 |
| tetR2:aTc --> aTc | 0.000289 |
| lacI4:IPTG:nsDNA --> IPTG + nsDNA | 0.000193 |
| tetR2:aTc:nsDNA --> aTc + nsDNA | 0.000193 |
Simulations
All simulations were carried out using the SimBiology Matlab Toolbox, which was freely available to iGEM teams. The File:SimBiology project file used for the following simulations can be accessed File:Here.
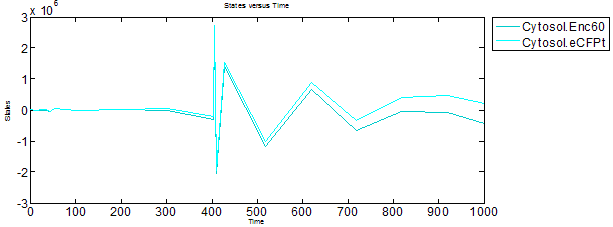
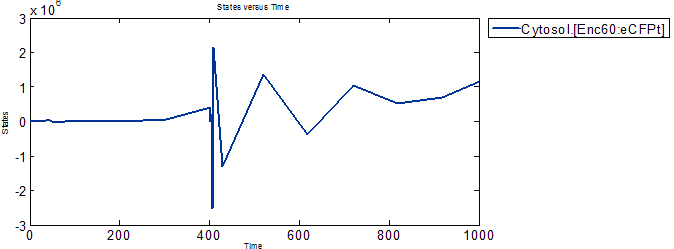
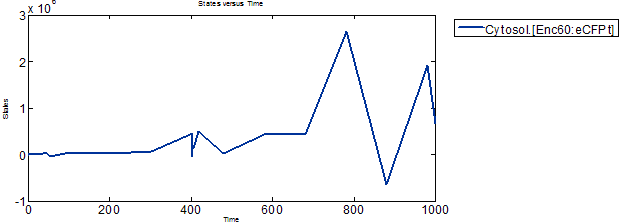
Accumulation of Proteins and Encapsulin-eCFP complex
It can be observed that the expressed proteins and Encapsulin-eCFP complex have some sort of oscillatory behavior, and that the later inverses the former. The Encapsulin and eCFPt proteins concentrations build up while the complex concentation stays low, and then the complex starts to form while the reservoirs of Encapsulin and eCFPt are depleted.
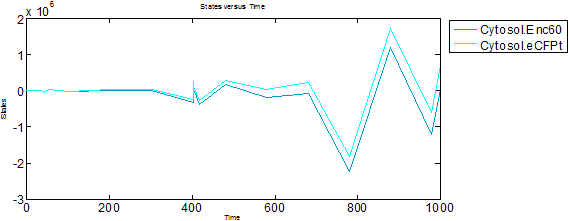
Addition of IPTG and aTc effectors
IPTG and aTc effectors bind lacI4 and tetR2 respectively, preventing them from inhibiting transcription. Predictably, when IPTG and aTc are added to the system (initial amounts changed from 0 --> 100), we observe higher peaks in protein expresssion and complex formation.
Sensitivity Analysis
It is interesting to note which parameters have the greatest effect the dynamics of the system. Since the parameters in our model are approximations from literature sources, the most senstive parameters would be leading candidates to be experimentally determined.
 "
"