Team:MoWestern Davidson/project
From 2009.igem.org

Overview
The Satisfiability (Sat) Problem:
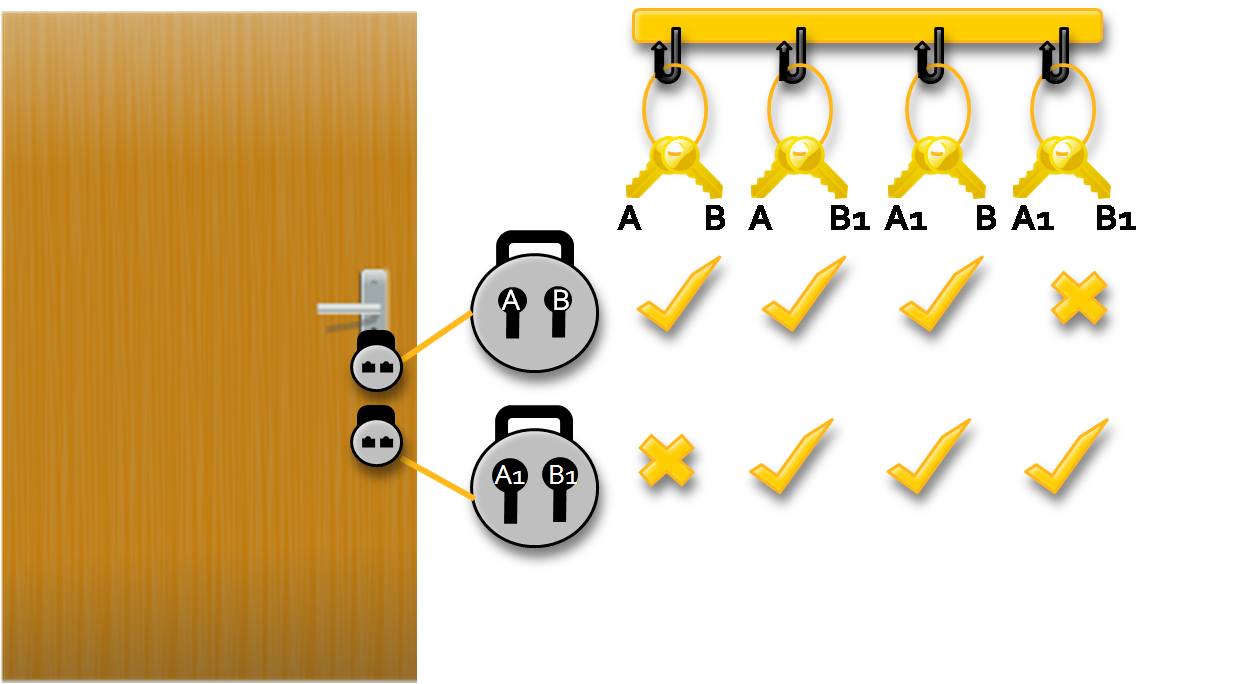
A member of the NP Complete family (the most challenging of the non-deterministic polynomial time problems), The Sat problem can be compared to an analogy of locks and keys.
Imagine a door with two locks(see figure1). each lock can accept two different keys, but the door will only open when there is a key to open both locks. A janitor with two sets of keys  wants to find the combination of keys that will open the door
wants to find the combination of keys that will open the door
Our team translated this np-complete problem into a modified biological process. In cells, the process of transcription codes DNA into RNA, and then the RNA is translated into a protein that is expressed once completed. Just as an English sentence requires a specific grouping of letters to form words, DNA must be read in 3 nucleotide groupings called codons to continue the process successfully. Transfer RNAs (tRNAs) are RNA molecules with 3 nucleotide anticodon loops that match specifically to codon sequences and supply the amino acids according to those sequences. These amino acids are required to construct the protein. If nucleotides are inserted or deleted, the reading frame is shifted, causing a frameshift mutation. That frameshift mutation causes the tRNA to supply the wrong amino acid in the peptide chain that then gives an unanticipated protein, or no protein.
With this biological background, the connection can be made between the “door and key” analogy and the biological interpretation. Our “doors” are engineered reporter gene DNA sequences including the 5 nucleotide frameshift mutation “keyhole”, resulting in our frameshift leaders (FSL). The “keys” that may or may not open the door are engineered 5 nucleotide anticodon suppressor tRNAs. Opening the door, thus solving the MAX SAT problem, would allow the cell to convey this outcome in the expression of the reporter gene (fluorescence/antibiotic resistance). The idea of using suppressor tRNAs to bypass designed mutation to solve a SAT problem is an extension using suppressor logic from papers by Thomas J. Magliery, J. Christopher Anderson and Peter G. Schultz {Magliery et. al. & Anderson et. al.}. This team worked with 2,3,4,5, and 6 nucleotide frameshift mutations. Their research found that 4 and 5 base codon suppression was most efficient in their mission of expanding the genetic code. The difference in the original amino acid and the modified 5mer amino acid is the addition of the 2 nucleotides.
 "
"