Team:Minnesota/Modeling
From 2009.igem.org
| Home | The Team | The Project | Parts Submitted to the Registry | Modeling | Notebook |
|---|
Contents |
Our Modeling Goals
We want to improve the user interface for SynBioSS (see below), which was developed for last year's competition and also create mathematical models of our experimental results. Our computational output will guide future research and is the key to determining what is happening at the molecular level of our logical AND gate.
Computational Tools
Synthetic Biology Software Suite (SynBioSS)
The entire software suite can be found via the SynBioSS [http://synbioss.sourceforge.net home page]. The software suite is open source that runs on Windows, Macintosh and Linux. It does not require command-line programming and thus facilitates the modeling of biochemical reactions.
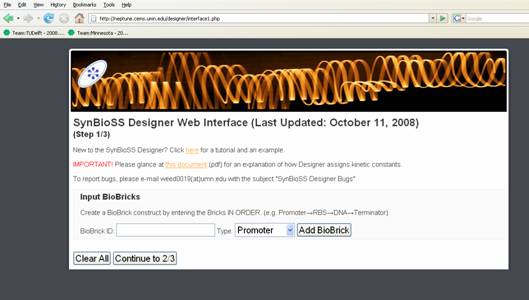
SynBioSS Designer
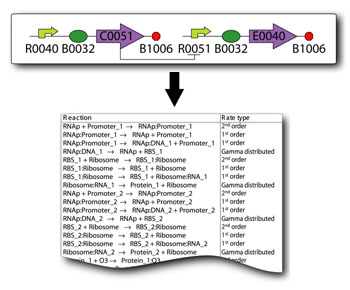
SynBioSS Designer is an application for the automatic generation of sets of biomolecular reactions. This software allows a user to input the molecular parts involved in gene expression and regulation (e.g. promoters, transcription factors, ribosomes, etc.) The software then generates complete networks of reactions that represent transcription, translation, regulation, induction and degradation of those parts. To facilitate the creation of detailed kinetic models of synthetic gene networks composed of BioBricks, we have adapted SynBioSS Designer to automatically generate a kinetic model from a construct composed entirely of BioBricks. A NetCDF or SBML file is generated for simulations.
Our goal for this year's iGEM competition is to improve the user interface for SynBioSS be...
SynBioSS wiki
The inaccessibility of requisite kinetic data complicates the generation of detailed mechanistic models. We address this barrier by creating a web accessible database curated by users in a “Wiki” format. SynBioSS Wiki is a significant extension of the open-source Mediawiki software. The Wiki stores reaction kinetic data in a formatted and searchable scheme with references to the relevant literature. This framework allows for the input of reactions whose rates are described either by elementary first & second order rate equations or any arbitrarily complex rate equation defined using MathML (e.g. Hill type reactions). Reactions may be searched via participating molecules which may be proteins, DNA sequences, small molecules, etc. Once located, reactions of interest (along with their associated kinetic data) may be collected. The completed model may be exported in SBML format for additional editing or simulation.
It is also through the SynBioSS Wiki databases that SynBioSS Designer can access and proliferate kinetic information related to the simulation of BioBricks, thus extending the utility of the database for the benefit of the greater modeling community. To jumpstart the process, we have entered the known biomolecular interactions in the expression and regulation of well-studied operons, such as the lactose, the tetracycline and the arabinose operons.
 "
"