Team:UCSF/Project
From 2009.igem.org
| Home | The Team | The Project | Parts Submitted to the Registry | Our summer experience | Notebook | Human Practices |
|---|
(Or you can choose different headings. But you must have a team page, a project page, and a notebook page.)
Contents |
Overall project
Your abstract
Motivation: why is this useful?
We envision a cellular robot that could travel to practically any site in the human body. This would provide a flexible platform that could be used for a variety of therapeutic tasks. The first step toward achieving this goal is to broaden the range of possible chemotactic targets for our cells. Ideally, we could connect virtually any input to chemotaxis in a generalized way.
Approach:
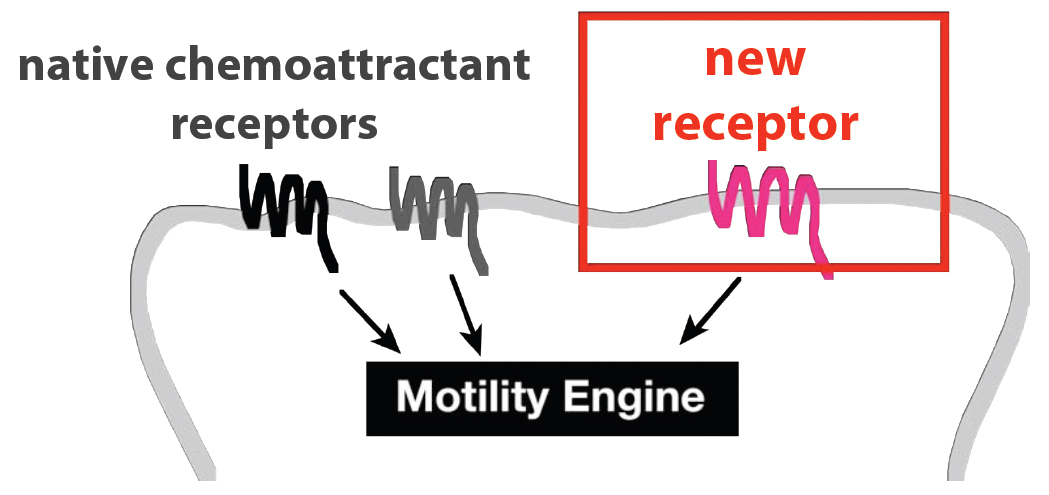
Neutrophils sense most of their chemotactic signals through [http://en.wikipedia.org/wiki/G_protein_coupled_receptor G protein-coupled receptors] (GPCRs). The spectrum of chemical signals to which these cells respond is therefore determined, at least in part, by the set of GPCRs they express. Can this spectrum be broadened arbitrarily by the introduction of new GPCRs? We tested this idea by transiently expressing 23 exogenous GPCRs in HL-60 (neutrophil-like) cells.
These cells were then tested for their ability to migrate toward ligands for the new GPCRs in multiwell [http://en.wikipedia.org/wiki/Chemotaxis_assay#Two-chamber_techniques Boyden chamber] assays. We measured the fold change in % of cells migrating toward the new ligand (with vs without added GPCR) at the peak response. We refer to this ratio as the "Migration Index." For receptors that appeared to activate a migration response (Migration Index > 3), we also conducted time-lapse microscopy to determine whether the cell movement was directed toward the gradient of ligand.
Results:
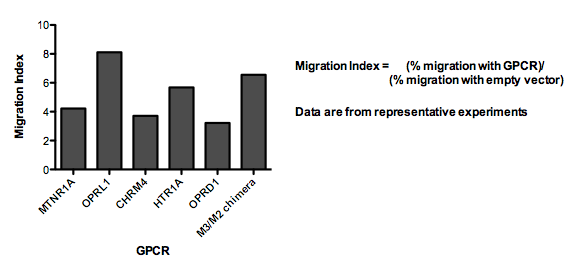
6 of the GPCRs we transiently expressed in our cells resulted in a Migration Index > 3.
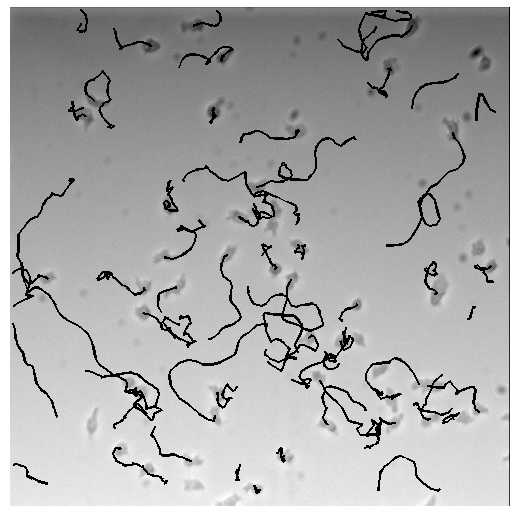
Here, we show an example of one of these receptors (M3/M2 chimera) mediating directional migration up a stable, linear gradient of ligand:
Part 2: Speed: engineering accelerators and brakes: A cellular cruise control by modulating cell polarity with feedback loops
Motivation: why is this useful?
Just like we have control over speed in a car – we can brake or accelerate – it would be useful to engineer such behavior into our cellular nanorobots. Just think about it: We could speed cells up so that they reach their targets faster and stop them once they have arrived or do not behave properly.
Background:
For these experiments we chose Dictyostelium discoideum cells to test our prototypical brakes and accelerators quickly. We expect that our brakes and accelerators can be used in a plug and play fashion because Dicty’s way of movement is very similar to a neutrophil’s:
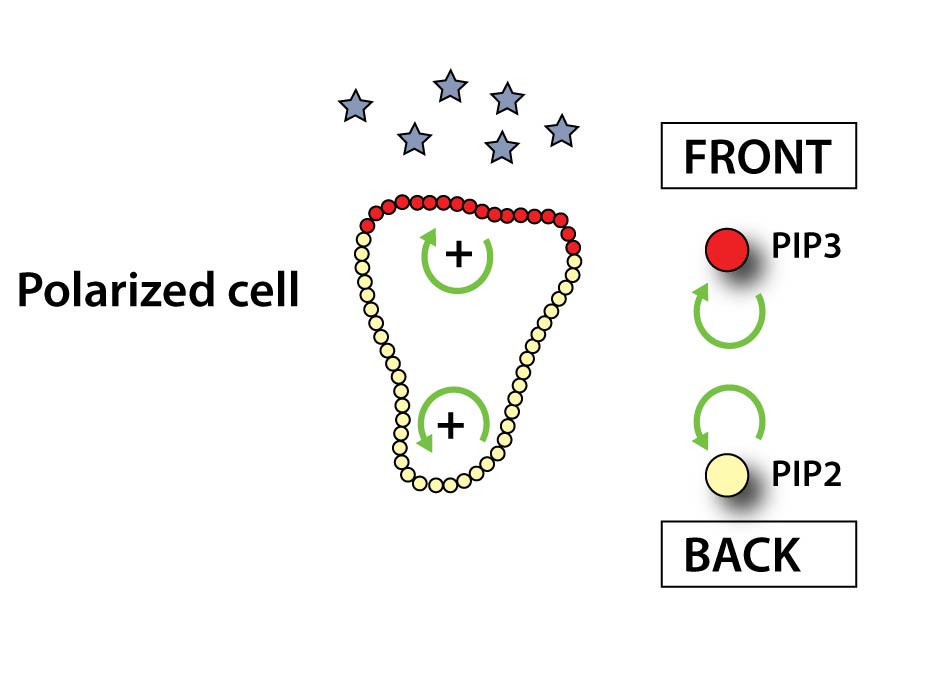
When a receptor binds chemoattractant, it induces the conversion of PIP2 to PIP3 (two signaling lipids in the plasma membrane) at the front of our cells. In a positive feedback loop PIP3 triggers the formation of more PIP3 at the front while similarly PIP2 leads to more PIP2 production at the sides and rear of the cell. This system sets the axis of polarity of the cell. The PIP3 patch at the front functions as a ‘turbo boost’ pushing the cell forward.
Approach:
Inspired by nature we tried to build accelerators and brakes by introducing our own synthetic protein based feedback loops. We designed feedback elements by fusing localization and catalytic domains involved in PIP3 production and degradation to modulate localization and level of PIP3 and PIP2 in the cell.
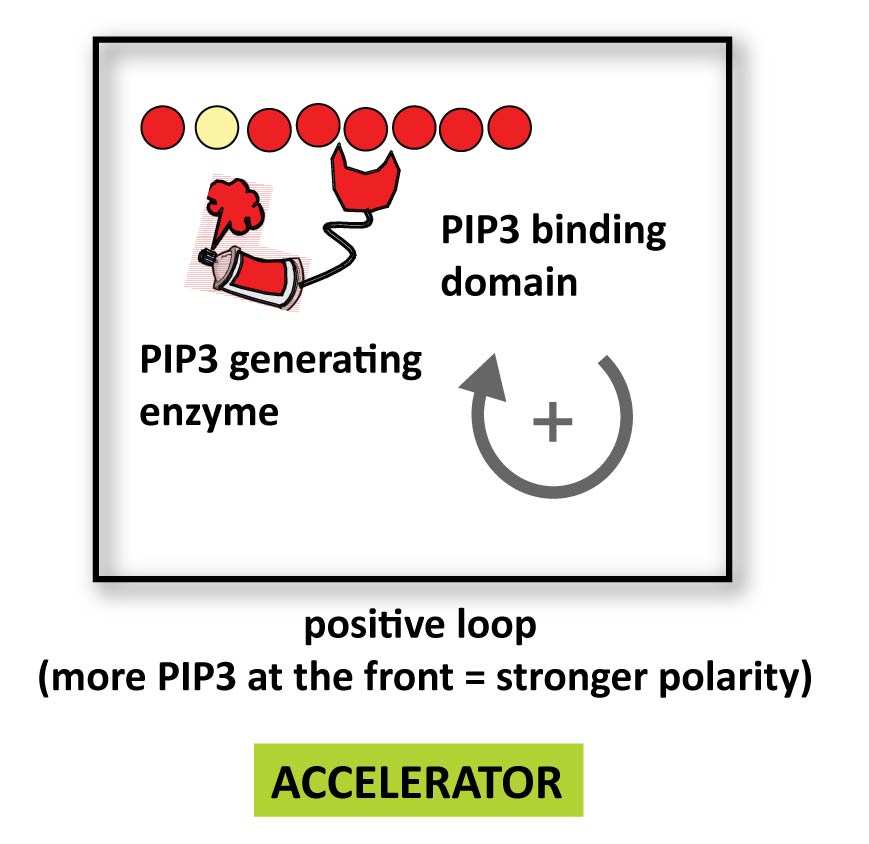
Here is an example of a positive feedback loop: a PIP3 binding localization domain fused to a PIP3 producing catalytic domain could produce more PIP3 where there is already PIP3- at the front. This might strengthen polarity and accelerate a cell.
Results:
Over the summer we assembled more than 100 fusions of localization and catalytic domains and screened whether they work. How? We measured the effect our constructs have on motility of Dictyostelium cells: stronger polarity should make cells faster while weaker polarity ought to slow them down!
Here is an overview of all feedback loops we screened and the effect they had on the speed of cells. We used automated cell tracking on more than 196 hours worth of movies (note: one movie is 10 minutes!) and identified strains that moved faster or slower at a very stringent statistical cutoff (p<0.0001).
This way we were able to identify 7 brakes and 1 accelerator!
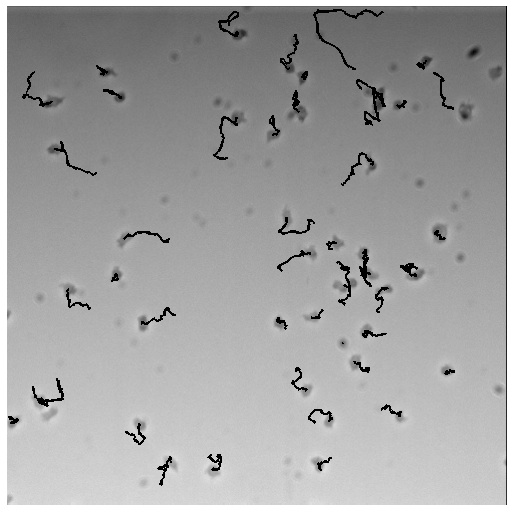
Check out the movie of one of our strong brakes (PTEN-RasC dominant active (da))[http://www.youtube.com/watch?v=_9od33Nx06Y] compared to wildtype [http://www.youtube.com/watch?v=Vtdtf8-zSRs].
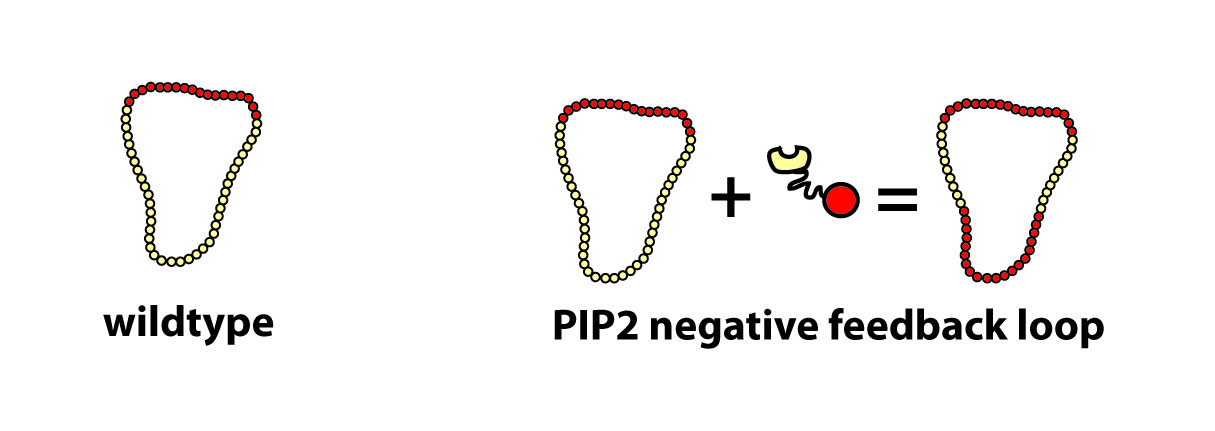
We hypothesize that this construct acts as a negative feedback loop on PIP2 - (generating PIP3 where PIP2 should be) thereby confusing the cell with multiple fronts:
This is indeed an effect of fusing the particular localization to the catalytic domain as neither of them alone has such a strong effect ( see details).
Summary and outlook:
We have screened more than 100 synthetic feedback elements for their ability to accelerate or slow down speed of cell motility. We have isolated a hand full of functional elements. Now we need to confirm the mechanism of action of these elements. In the future we would like to make them inducible by a signal from outside – like a stoplight!
Methods
- Cloning: Aar1 Shuffle - Connecting all our fragments together.
- Dicty: Transformation Protocol - Used to transform into dicty
- Dicty: Motility Assay Protocol - Used to analyze the basic motility of all cells
 "
"