Team:Uppsala-Sweden/Ethanol
From 2009.igem.org
The Pathway
The idea to produce ethanol with the use of cyanobacteria was originally proposed and already accomplished by Ming-De Deng and John R. Coleman. [1] We from the Uppsala iGEM Team decided to try to improve the production of ethanol by interfering with the metabolic pathways of Synechocystis sp. PCC 6803 by an Antisense RNA and a protein-mediated approach. Hence we build a construct that encodes for ethanol production in our host organism. But first of all the some theory about the molecular mechanism.
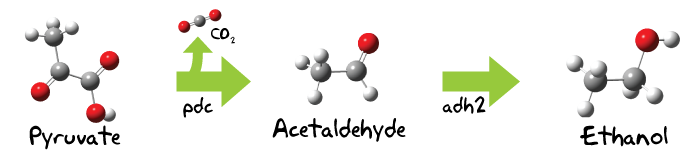
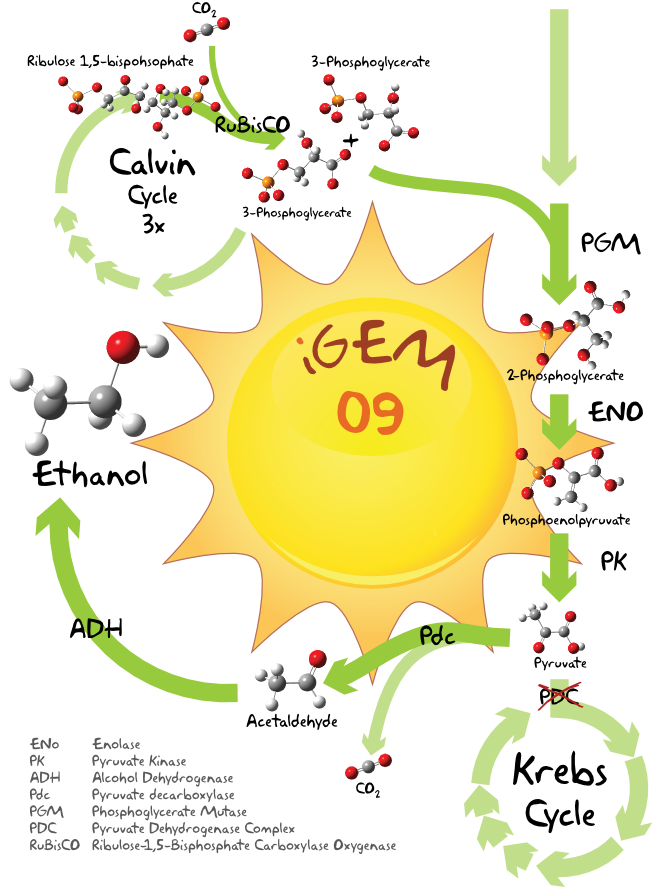
Light energy is used in the Calvin cycle to transform water and carbon dioxide into organic compounds, among those is Ribulose-1,5-bisphosphate, which immediately splits into two 3-Phosphoglycerate molecules. [2] These two molecules can now be converted to 2-Phosphoglycerate. Enolase and pyruvatekinase finally catalyze then the reaction over phosphoenolpyruvate to pyruvate. [3] This pyruvate molecule is now substrate for the ethanol production.
blabal mechanism
References
[1] [http://www.ncbi.nlm.nih.gov/pubmed/9925577 Ethanol synthesis by genetic engineering in cyanobacteria. Deng MD, Coleman JR. Appl Environ Microbiol. 1999 Feb;65(2):523-8]
[2] Jeremy M. Berg, John L. Tymoczko, L. Stryer Biochemistry 5th Edition; Ch.21.1 p826-838<i> 653
[3] Jeremy M. Berg, John L. Tymoczko, L. Stryer <i>Biochemistry 5th Edition; Ch.16.1 p653<i>
[4] [http://biocyc.org/META/NEW-IMAGE?type=REACTION&object=RXN-7643 MetaCyc Reaction: 4.1.1.1]
[5] [http://biocyc.org/META/NEW-IMAGE?type=REACTION&object=RXN-7657 MetaCyc Reaction: 1.1.1.1]
 "
"