Team:NYMU-Taipei/Project/Promoter Strength Testing
From 2009.igem.org
Revision as of 22:42, 21 October 2009 by Blackrabbit (Talk | contribs)
- All experiments were done in E. coli DH5alpha.
- Maturation rate of GFP is 6.5mins [1]
- All our analysis was done in excel documents: File:NYMU-promotertesting.zip and done based on the paper by Leveau and Lindow [2].
Contents |
2009-08-25 Reporting Assay
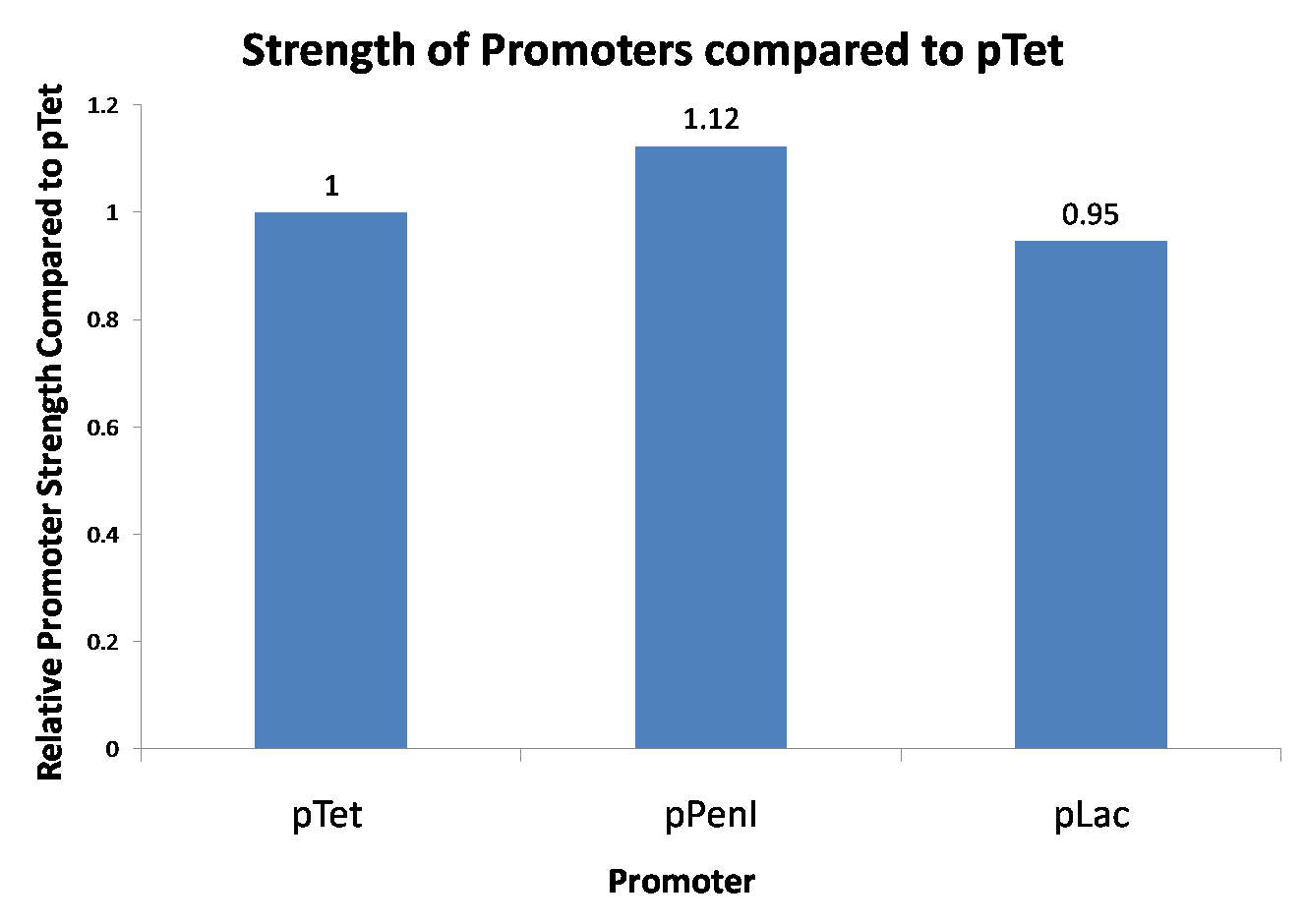
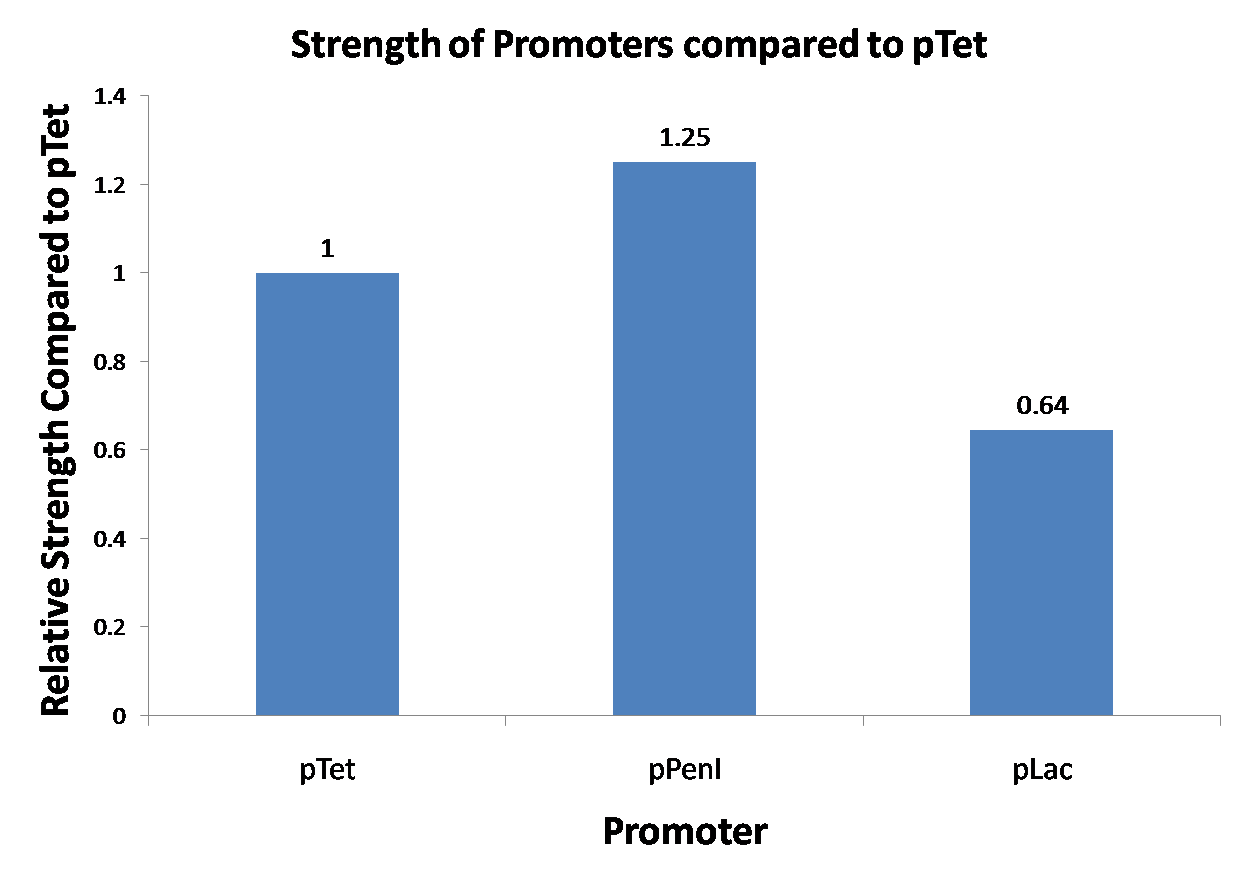 | Figure 1. This is the graph of our promoter strength testing. Since we did not do biological replicates, we are not able to confirm the validity of the result. |
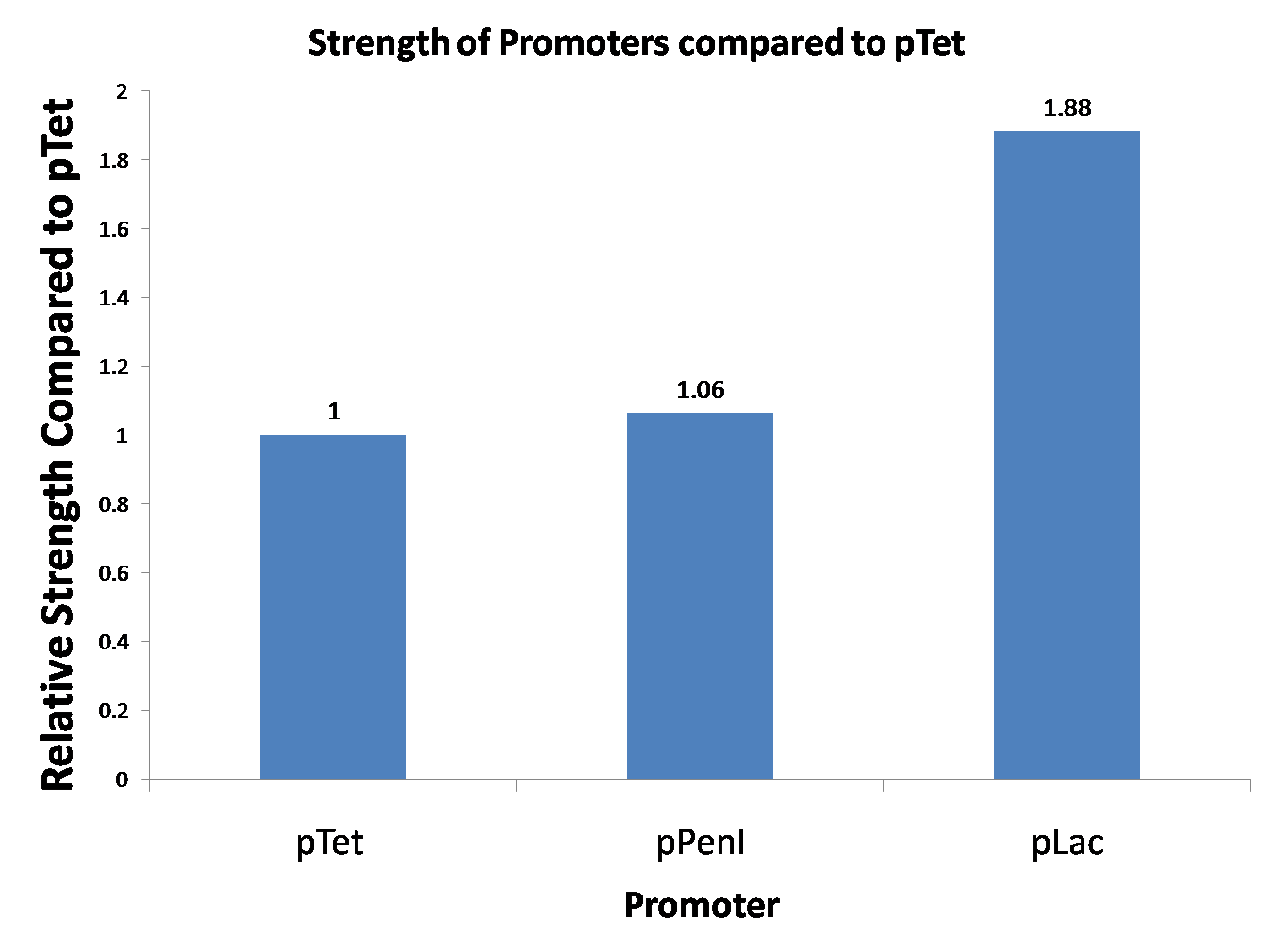
2009-09-07 Reporting Assay
2009-09-18 Reporting Assay
2009-09-29 Reporting Assay
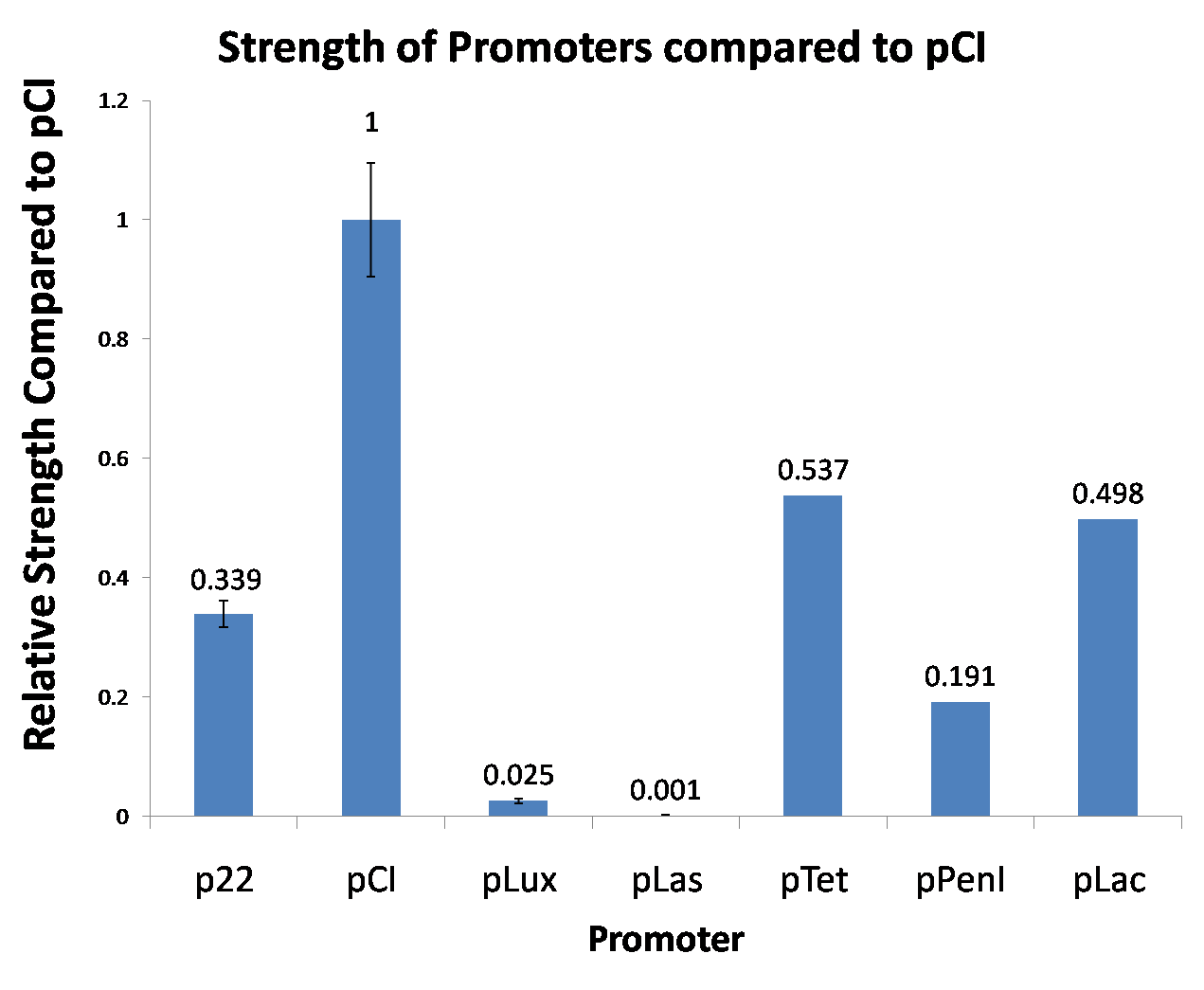
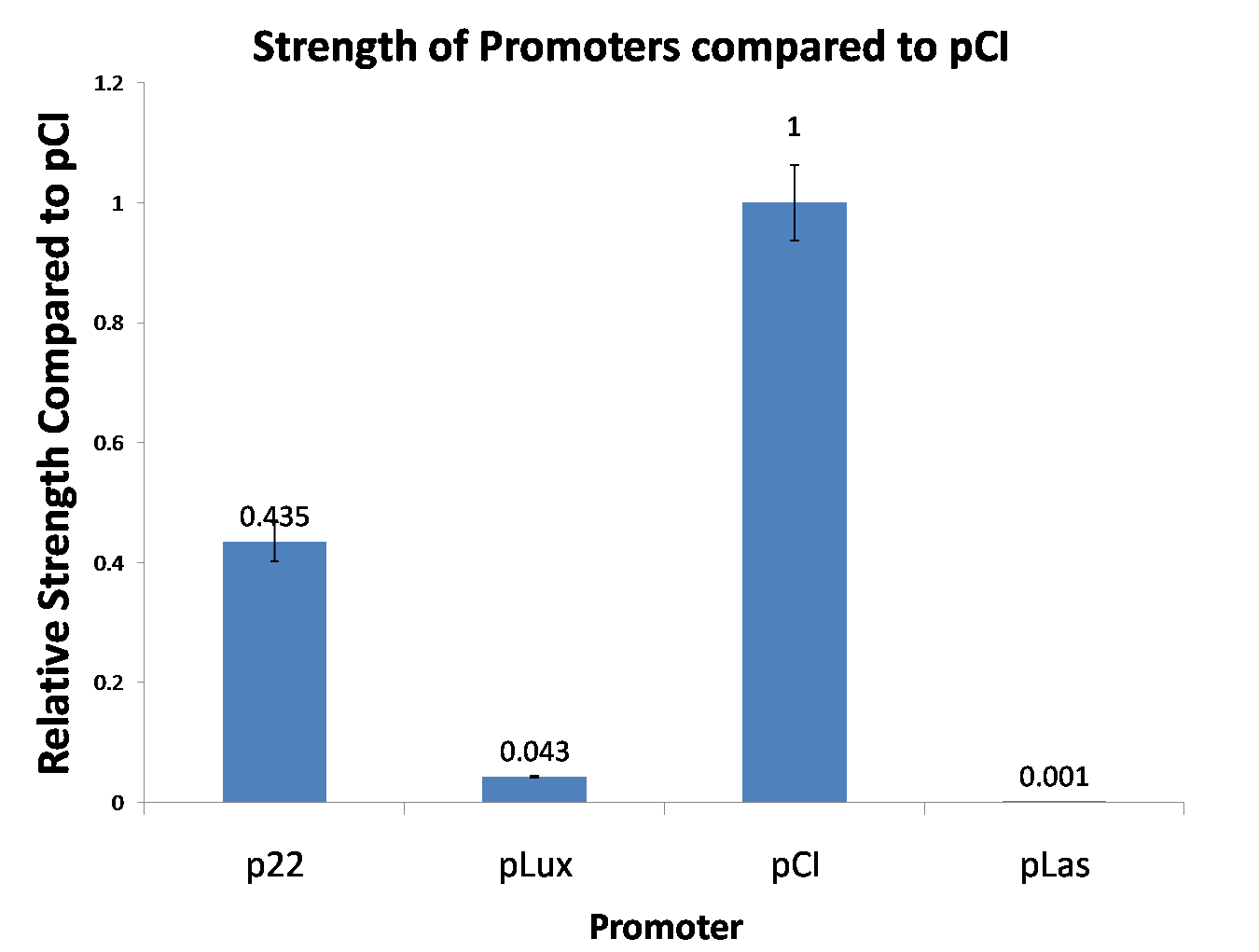 | Figure 4. We tested new promoters and measured their strength relative to pCI. This time we did a three-fold biological replicate. The error bars are 95% confidence intervals. |
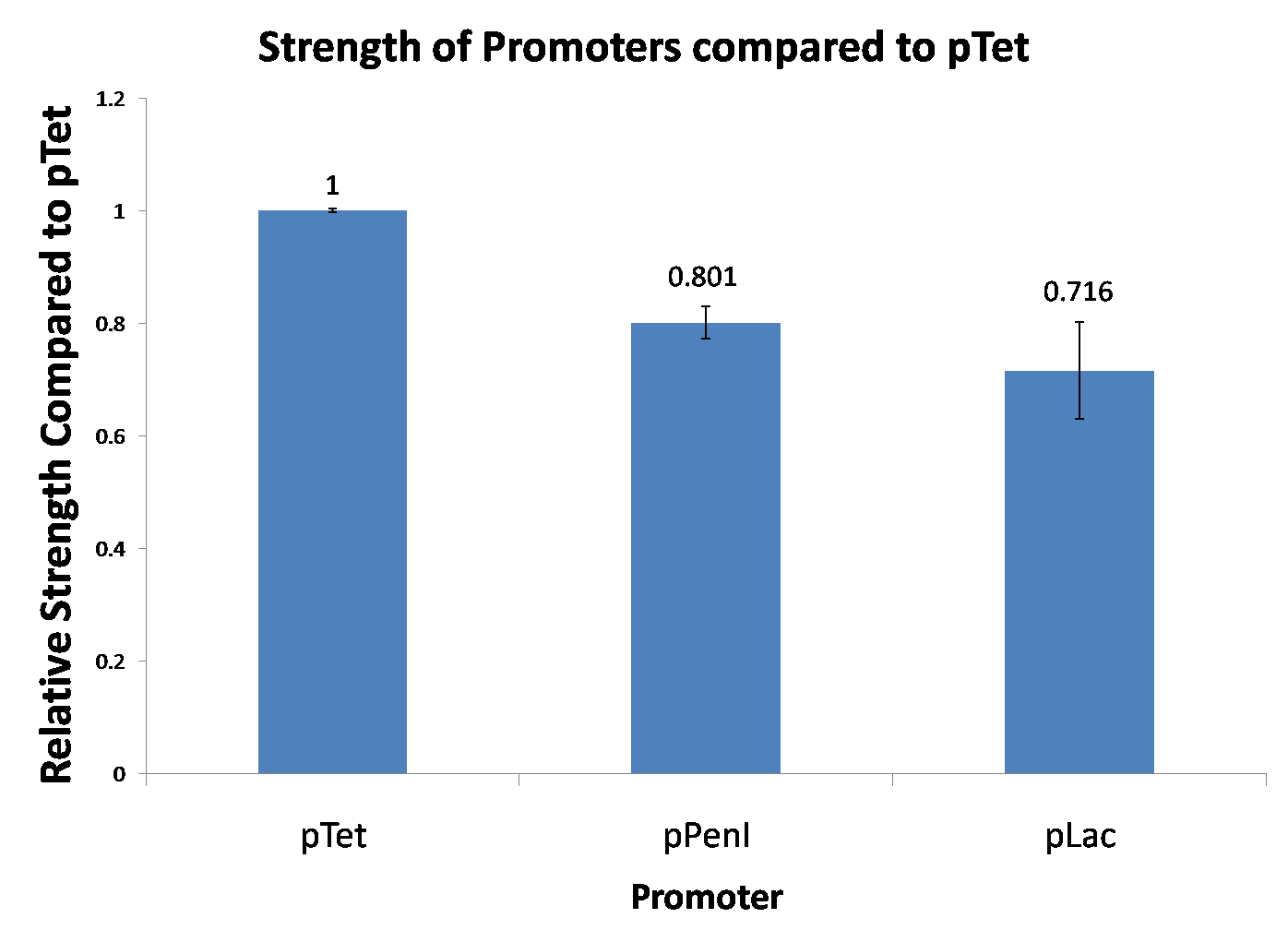
2009-10-05 Reporting Assay
2009-10-13 Reporting Assay
Conclusion
- We have tested the relative strengths of p22, pCI, pLux (basal level), pLas (basal level), pTet, pPenI, and pLac.
- We obtained relatively consistent results between the relative strengths of p22, pCI, pLux, and pLas, while obtaining not so consistent results between pTet, pPenI, and pLac.
- We think it is due to the lack of experience earlier on and confusing us which relative strengths between pTet, pPenI, and pLac were correct.
Reference
[1] Megerle JA, Fritz G, Gerland U, Jung K, Radler JO: Timing and Dynamics of Single Cell Gene Expression in the Arabinose Utilization System, Biophys J, 2008, 95(4): 2103–2115
[2] Leveau JHJ and Lindow SE, Predictive and Interpretive Simulation of Green Fluorescent Protein Expression in Reporter Bacteria, Journal of Bacteriology, 2001, 183(23): 6752-6762
 "
"