Team:MIT/Projects/Project2
From 2009.igem.org
Light-inducible protein localization in Yeast
Objective
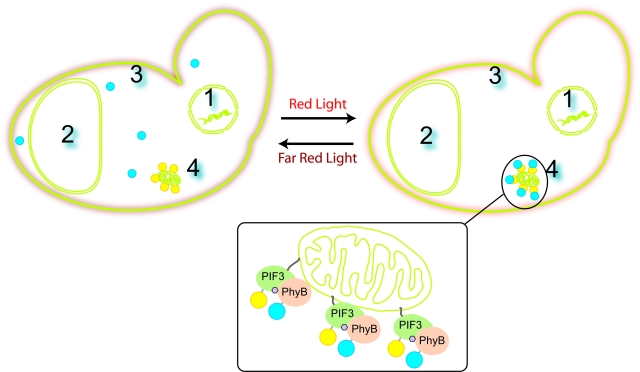
We set out to create a quick, reversible switch to control the localization to various targets within a yeast cell, such as the nuclear membrane, plasma membrane, mitochondrial membrane, and vacuole.
Motivation
To fully control cellular functioning, it would be useful to have a modular system that can provide both temporal and spatial control over any protein of interest within a cell. In other words, we should be able to control both when a gene is expressed and where the protein is functioning. The existing PhyB-PIF3 system provides temporal control over gene expression by acting as a transcriptional regulator. However, by adapting the PhyB-PIF3 system to control protein targeting, we can have both temporal and spatial control over protein function. Such a system could be used as a rapid on-off switch for localization-dependent processes. For example, because protein targeting is important in control of the cell cycle, the system could be used to control cell cycling and differentiation. Furthermore, reversible control of protein localization would be useful to study signal transduction and as a biophysical tool to study diffusion rates within a cell.
Background on PhyB-PIF3 System
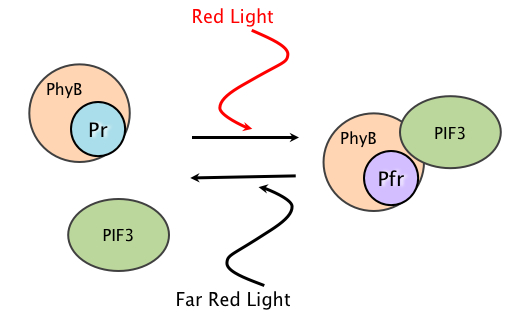
The light switching mechanism that we use exploits the machinery by which plants and algae respond to varying light conditions. Phytochromes are light sensitive proteins that are composed of a polypeptide bound to a small pigmented molecule called a chromophore. They have two conformations: the inactive Pr form and the active Pfr form. When red light is present, the phytochrome switches from the Pr to the Pfr form. When in the Pfr form, the phytochrome can bind to proteins called phytochrome interacting factors. In nature, the resulting complex can bind to DNA and act as a transcriptional regulator. When far red light is present, the phytochrome dissociates from the phytochrome interacting factor, making the switching mechanism reversible. Peter H. Quail's lab at University of California Berkeley developed a synthetic biological switch using a phytochrome called PhyB, which is bound to a chromophore called phycocyanobilin (PCB). Red light causes PhyB to bind to a phytochrome interacting factor called PIF3, and far-red light causes them to unbind. Our system harnesses this switching mechanism to control post-translational processing instead of transcriptional regulation.
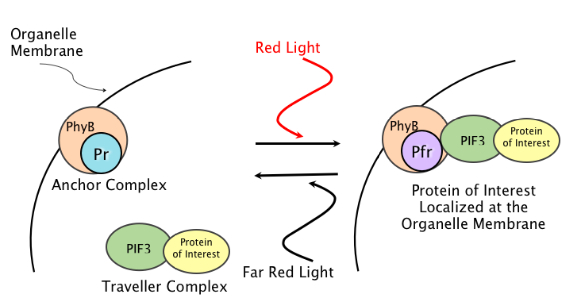
Mechanism
To localize our protein of interest, either PhyB or PIF3 is anchored to the target while the other is the traveler construct that diffuses throughout the cell. The anchored construct contains a signal sequence, which causes it to be localized constitutively to the target. The traveler, on the other hand, is fused to the protein of interest. When pulsed with red light, the PhyB will switch to the Pfr conformation. As a result, the as the traveler diffuses throughout the cell, PhyB and PIF3 will bind, which attaches the traveler to the anchor. As the traveler becomes bound to the anchor, the protein of interest becomes localized to the target. Under far red light, the anchor and the traveler dissociate, the the protein of interest again can diffuse throughout the cell. In our system, PIF3 is tagged with Yellow Fluorescent Protein (YFP) and PhyB is tagged with Cyan Fluorescent Protein (CFP) so that the localization and delocalization can be observed using fluorescence microscropy.
Constructs
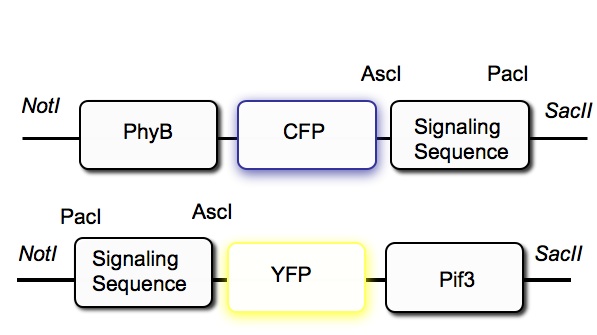
The diagram below shows the DNA constructs designed to reversibly control protein targeting in yeast. PhyB is fused with CFP, while PIF3 is fused with YFP. Plasmids with these two fusion constructs serve as our base constructs into which signal sequences are to be added. For the anchor constructs, a signal sequences is added to direct the resulting protein to the target for localization. Whether PIF3 or PhyB is used as the anchor depends on which signal sequence is used to direct the protein to the target. For signal sequences that function on the N-terminal of the protein, PhyB is used as the anchor and PIF3 as the traveler. For C-terminal signal sequences, PIF3 is used as the anchor and PhyB as the traveler.
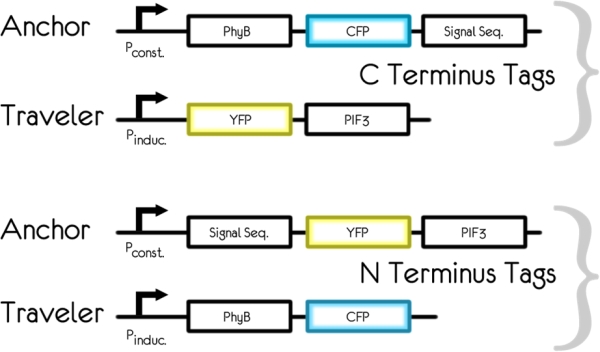
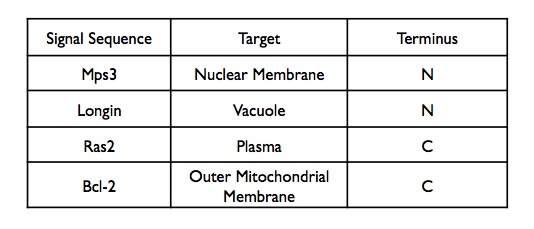
Materials and Methods
Construct Construction
The PhyB-CFP and YFP-PIF3 constructs were constructed using two methods: sequential cloning and homologous recombination. PhyB-CFP was built into pRS313 with a Myo2 promoter, while YFP-PIF3 was built into pRS415 with an ADHI promoter. Using both methods, we were able to construct the base constructs, with PhyB-CFP and YFP-PIF3 without signal sequences inserted.
Sequential Transformation Plasmid Construction
Construction of the PhyB-CFP and Pif3-YFP Plasmids began with the amplification of each of the individual sequences, PhyB, CFP, PIF3 and YFP, from the Berkley Lab Plasmids. These sequences were subsequently digested and ligated into the prS313 backbone, a plasmid that contains a constitutive Myo2 promoter and the His gene. PIF3 and CFP were inserted using NotI and SacII, while PhyB, YFP used NotI and XmaI. Once the plasmid was complete it was transformed into E. coli. The PhyB and Pif3 plasmids transformed successfully, which was confirmed through a check digest and sequencing. The corresponding fluorescent protein DNA was then incorporated into the plasmid through another round of digestion, ligation and transformation. The sequencing results for these plasmids were positive for incorporation of both DNA pieces for both of the constructs. Therefore, each of these plasmids were transformed into yeast and observed for fluorescence through microscopy.
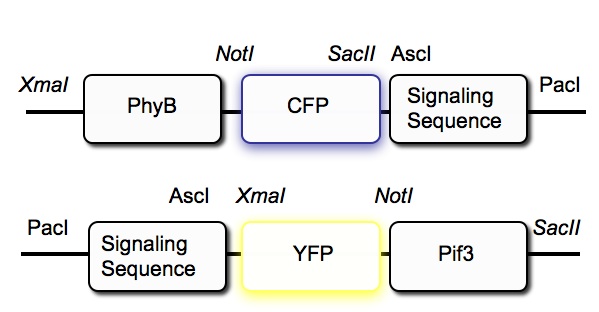
Alongside work in the base constructs, a variety of C and N terminus localization sequences were chosen to test the PhyB-PIF3 System. Each localization sequence, once amplified, would be cloned into the appropriate backbone using the restriction enzymes. N terminus tags were destined for the Pif3-YPF backbone and C terminus for the PhyB-CFP Backbone
Homologous Recombination
PhyB and CFP were PCR amplified using primers such that there was a region of homology of 40 base pairs between the PhyB product and the CFP product as well as 40 base pairs of homology with the pRS313 backbone on both the PhyB and CFP products. The pRS313 backbone was linearized using a double restriction enzyme digest. The high efficiency yeast transformation protocol was carried out such that the linearized backbone as well as the PhyB and CFP PCR products were added to the transformation mix. When crossing over between the regions of homology occured, the result is a circular plasmid with both PhyB and CFP, as shown the the diagram below. The yeast was then grown on selective media to select for strains in which the cross-over events that result in a circular plasmid occur. A similar procedure was carried out for PIF3 and YFP. PIF3 and YFP were PCR amplified so that they had a 40 base pair long region of homology with each other and with the pRS415 backbone. pRS415 was linearized and transformed into yeast with the PIF3 and YFP products. The yeast with the circular plasmid with YFP and PIF3 were then selective for using selective media.
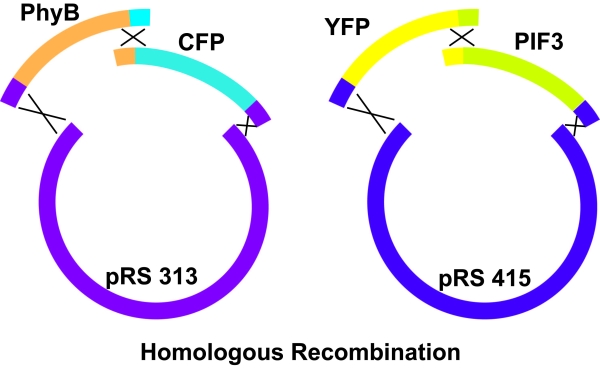
LED Arrays for Red and Far Red Light

 "
"