Team:Minnesota/Modeling
From 2009.igem.org
| Home | The Team | The Project | Parts Submitted to the Registry | Modeling | Notebook |
|---|
Contents |
Our Modeling Goals
We want to improve the user interface for SynBioSS (see below), which was developed for last year's competition and also create mathematical models of our experimental results. Our computational output will guide future research and is the key to determining what is happening at the molecular level of our logical AND gate
Computational Tools
Synthetic Biology Software Suite (SynBioSS)
The entire software suite can be found via the SynBioSS [http://synbioss.sourceforge.net home page].
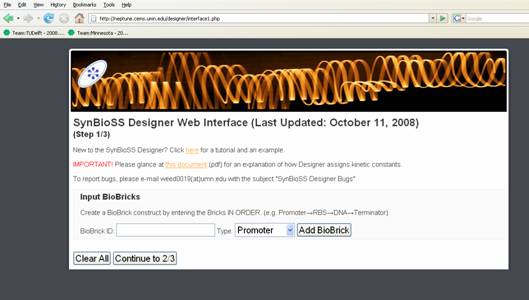
SynBioSS Designer
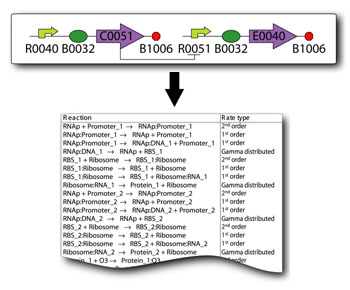
SynBioSS Designer is an application for the automatic generation of sets of biomolecular reactions. This software allows a user to input the molecular parts involved in gene expression and regulation (e.g. promoters, transcription factors, ribosomes, etc.) The software then generates complete networks of reactions that represent transcription, translation, regulation, induction and degradation of those parts. To facilitate the creation of detailed kinetic models of synthetic gene networks composed of BioBricks, we have adapted SynBioSS Designer to automatically generate a kinetic model from a construct composed entirely of BioBricks. A NetCDF or SBML file is generated for simulations.
Our goal for this year's iGEM competition is to improve the user interface for SynBioSS be...
 "
"