Team:NYMU-Taipei/Project/Receptor/Sialic Acid
From 2009.igem.org
| Home | Project Overview: | Chassis | Receptors | Removal | Experiments and Parts | F.A.Q | About Us |
Motivation & Design Background
Influenza has been a pathogen infecting the human body since centuries ago. Recently, H1N1 became prevalent around the world, so we hope our design could make some contribution towards fighting against it. Among so many mechanisms which play important role in the infection process of Influenza virus, sialic acid is the most important one that gets involved in this infection process. There are many different types of sialic acid exist in different species[1]. For example, on the human cell membranes are alpha-2,6-sialic acid that mostly play an important role in the infection of virus. Others such as alpha-2,8-sialic acid is essential for the pathogen of some other kind of bacteria like E.coli K1. The main difference is the enzyme that is used in the process of producing the sialic acid. Sialyltransferase is the enzyme that changes alpha-2,8-sialic acid to alpha-2,6-sialic acid[3][4][5]. Which is aimed at transferring NeuNAc, the substrate that is made from other two enzymes--UDP-GlcNAc-2-Epimerase and NeuNAc Synthase onto cell membrane’s proteins or lipid in the position of alpha- 2,6 or alpha-2,8 sialic acid. Several animal viruses, including the influenza virus, attach to their host cells through interactions with oligosaccharides displayed on the host cell surface. The HA (hemagglutinin) protein is essential for influenza viruses to enter and infect healthy body cells. The residues of Neu5Ac (a type of sialic acid) is situated at the ends of the oligosaccharide chains of many plasma glycoproteins. The oligosaccharide ligands that bind to influenza viruses' HA is Neu5Ac(alpha2-6)Gal(beta1-4)Glc. We try to modify the lipopolysaccharide on the membrane of bacteria because there is already alpha-2,8-sialic acid at the ends of the oligosaccharide chains of plasma glycoproteins. The technique we use is the synthesis of glycoprotein.[2]
Materials and Methods
The neuS gene in E.coli K1 is coding for alpha-2,8-sialyltransferase, and there is also some kinds of photobacteria that have genes coding for alpha-2,6-sialyltransferase. We performed PCR to clone the neuD, neuB, neuA, neuC, neuE genes, from one operon in E.coli K1, and are coding for enzymes that are in charge of sialic acid synthesis. On the other hand, we do PCR for cloning sialyltransferase gene from photobacteria. After performing these PCR, and make sure all the PCR products' length are correct, doing restriction enzyme digestion and then ligation, finally transformation are the next work. We use colony PCR to make sure the colony is what we want. We find the appropriate promoter(which can promote quantitative expression of genes)and ribosome binding site to ligate with the coding sequences-neu genes.
sialic acid alpha2,6 organism
- Photobacterium damselae
sialic acid alpha2,8 organism
- Escherichia coli O1:K1 / APEC with [Complete proteome] in NCBI and ENSEMBL
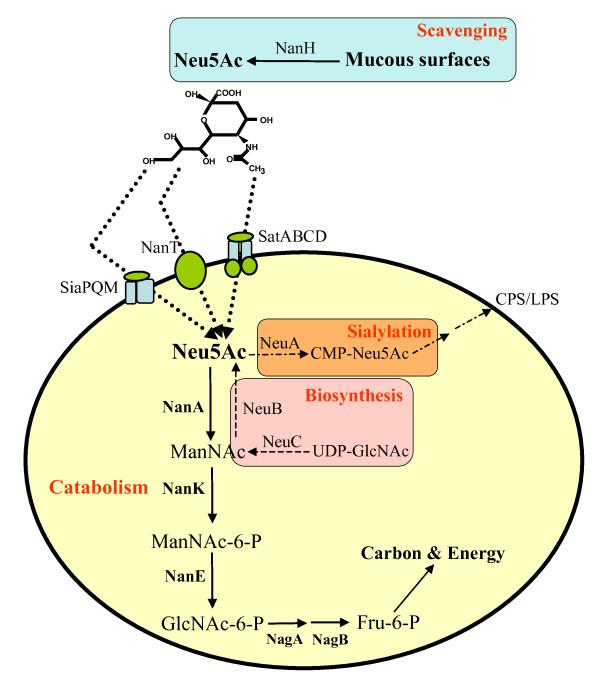
The comparison of the process of sialic acid synthesis between bacteria and mammals
- FigureA. This picture shows the catabolism of sialic acid among bacteria.[7]
- Sialic acid biosynthesis genes(neuA,neuB,neuC)among the bacteria encoding the Nan cluster(the genes encoding NanA, NanB, NanE are clustered together)
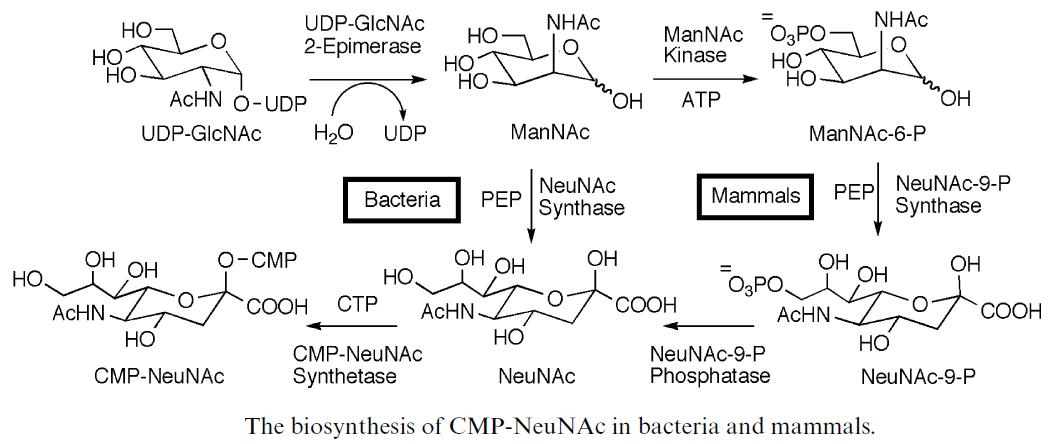
- FigureB. The differences of biosynthesis of sialic acid between the bacteria and mammals.[6]
Circuit Design
neu genes cluster descriptions:- neuD(623bp)--sugar O-acyltransferase, sialic acid O-acetyltransferase NeuD family
- neuB(1040bp)--N-acetylneuraminate synthase
- neuA(1256bp)--CMP-N-acetylneuraminic acid synthetase
- neuC(1175bp)--UDP-N-acetyl-D-glucosamine 2-epimerase,UDP-hydrolysing
- neuE(1175bp)--polysialic acid biosynthesis protein
- sialyltransferase 0160(2743bp)--Photobacterium damsela gene for sialyltransferase 0160
| pTet R0040 |
RBS B0034 |
kps genes cluster |
B0015 |
| pTet R0040 |
RBS B0034 |
kpsM、kpsT |
B0015 |
Gene Sequences
[http://www.ncbi.nlm.nih.gov/nuccore/454079?ordinalpos=1&itool=EntrezSystem2.PEntrez.Sequence.Sequence_ResultsPanel.Sequence_RVDocSum Escherichia coli polysialic acid gene cluster region 2 (neuD and neuB) genes, complete cds]
atgagtaa aaagttaata
121 atatttggtg cgggtggttt ttcaaaatct ataattgaca gcttaaatca taaacattac
181 gagttaatag gatttatcga taaatataaa agtggttatc atcaatcata tccaatatta
241 ggtaatgata ttgcagacat cgagaataag gataattatt attattttat tgggataggc
301 aaaccatcaa ctaggaagca ctatttaaac atcataagaa aacataatct acgcttaatt
361 aacattatag ataaaactgc tattctatca ccaaatatta tactgggtga tggaattttt
421 attggtaaaa tgtgtatact taaccgtgat actagaatac atgatgccgt tgtaataaat
481 actaggagtt taattgaaca tggtaatgaa ataggctgct gtagcaatat ctctactaat
541 gttgtactta atggtgatgt ttctgttgga gaagaaactt ttgttggtag cgtgactgtt
601 gtaaatggcc agttgaagct aggctcaaag agtattattg gttctgggtc ggttgtaatt
661 agaaatatac caagtaatgt tgtagttgct gggactccaa caagattaat tagggggaat
721 gaatgagtaa tatatatatc gttgctgaaa ttggttgcaa ccataatggt agtgttgata
781 ttgcaagaga aatgatatta aaagccaaag aggccggtgt taatgcagta aaattccaaa
841 catttaaagc tgataaatta atttcagcta ttgcacctaa ggcagagtat caaataaaaa
901 acacaggaga attagaatct cagttagaaa tgacaaaaaa gcttgaaatg aagtatgacg
961 attatctcca tctaatggaa tatgcagtca gtttaaattt agatgttttt tctacccctt
1021 ttgacgaaga ctctattgat tttttagcat ctttgaaaca aaaaatatgg aaaatccctt
1081 caggtgagtt attgaattta ccgtatcttg aaaaaatagc caagcttccg atccctgata
1141 agaaaataat catatcaaca ggaatggcta ctattgatga gataaaacag tctgtttcta
1201 tttttataaa taataaagtt ccggttggta atattacaat attacattgc aatactgaat
1261 atccaacgcc ctttgaggat gtaaacctta atgctattaa tgatttgaaa aaacacttcc
1321 ctaagaataa cataggcttc tctgatcatt ctagcgggtt ttatgcagct attgcggcgg
1381 tgccttatgg aataactttt attgaaaaac atttcacttt agataaatct atgtctggcc
1441 cagatcattt ggcctcaata gaacctgatg aactgaaaca tctttgtatt ggggtcaggt
1501 gtgttgaaaa atctttaggt tcaaatagta aagtggttac agcttcagaa aggaagaata
1561 aaatcgtagc aagaaagtct attatagcta aaacagagat aaaaaaaggt gaggtttttt
1621 cagaaaaaaa tataacaaca aaaagacctg gtaatggtat cagtccgatg gagtggtata
1681 atttattggg taaaattgca gagcaagact ttattccaga tgaattaata attcatagcg
1741 aattcaaaaa tcagggggaa taa
[http://www.ncbi.nlm.nih.gov/nuccore/146943?ordinalpos=1&itool=EntrezSystem2.PEntrez.Sequence.Sequence_ResultsPanel.Sequence_RVDocSum E.coli CMP-N-acetylneuraminic acid synthetase (neuA) gene, complete cds]
atgagaa caaaaattat tgcgataatt ccagcccgta
61 gtggatctaa agggttgaga aataaaaatg ctttgatgct gatagataaa cctcttcttg
121 cttatacaat tgaagctgcc ttgcagtcag aaatgtttga gaaagtaatt gtgacaactg
181 actccgaaca gtatggagca atagcagagt catatggtgc tgattttttg ctgagaccgg
241 aagaactagc aactgataaa gcatcatcat ttgaatttat aaaacatgcg ttaagtatat
301 atactgatta tgagagcttt gctttattac aaccaacttc accctttaga gattcgaccc
361 atattattga ggctgtaaag ttatatcaaa ctttagaaaa ataccaatgt gttgtttctg
421 ttactagaag caataagcca tcacaaataa ttagaccatt agatgattac tcgacactgt
481 ctttttttga ccttgattat agtaaatata atcgaaactc aatagtagaa tatcatccga
541 atggagctat atttatagct aataagcagc attatcttca tacaaagcat ttttttggtc
601 gctattcact agcttatatt atggataagg aaagctcttt agatatagat gatagaatgg
661 atttcgaact tgcaattacc attcagcaaa aaaaaaatag acaaaaaatt gacctttatc
721 aaaacataca taatagaatc aatgagaaac gaaatgaatt tgatagtgta agtgatataa
781 ctttaattgg acactcgctg tttgattatt gggacgtaaa aaaaataaat gatatagaag
841 ttaataactt aggtatcgct ggtataaact cgaaggagta ctatgaatat attattgaga
901 aagagctgat tgttaatttc ggagagtttg ttttcatctt ttttggaact aatgatatag
961 ttgttagtga ttggaaaaaa gaagacacat tgtggtattt gaagaaaaca tgccagtata
1021 taaagaagaa aaatgctgca tcaaaaattt atttattgtc ggttcctcct gtttttgggc
1081 gtattgatcg agataataga ataattaatg atttaaattc ttatcttcga gagaatgtag
1141 attttgcgaa gtttattagc ttggatcacg ttttaaaaga ctcttatggc aatctaaata
1201 aaatgtatac ttatgatggc ttacatttta atagtaatgg gtatacagta ttagaaaacg
1261 aaatagcgga gattgttaaa tga
[http://www.ncbi.nlm.nih.gov/nuccore/146945?ordinalpos=1&itool=EntrezSystem2.PEntrez.Sequence.Sequence_ResultsPanel.Sequence_RVDocSum E.coli protein p7 (neu C) gene, complete cds]
atgaaaaaaa tattatacgt aactggatct
181 agagctgaat atggaatagt tcggagactt ttgacaatgc taagagaaac tccagaaata
241 cagcttgatt tggcagttac aggaatgcat tgtgataatg cgtatggaaa tacaatacat
301 attatagaac aagataattt taatattatc aaggttgtgg atataaatat caatacaact
361 tcacatactc acattctcca ttcaatgagt gtttgcctca attcgtttgg tgattttttt
421 tcaaataaca catatgatgc ggttatggtt ttaggcgata gatatgaaat attttcagtc
481 gctatcgcag catcaatgca taatattcca ttaattcata ttcatggtgg tgaaaagaca
541 ttagctaatt atgatgagtt tattaggcat tcaattacta aaatgagtaa actccatctt
601 acttctacag aagagtataa aaaacgagta attcaactag gtgaaaagcc tggtagtgtg
661 tttaatattg gttctcttgg tgcagaaaat gctctttcat tgcatttacc aaataagcag
721 gagttggaac taaaatatgg ttcactgtta aaacggtact ttgttgtagt attccatcct
781 gaaacacttt ccacgcagtc ggttaatgat caaatagatg agttattgtc agcgatttct
841 ttttttaaaa atactcacga ctttattttt attggcagta acgctgacac tggttctgat
901 ataattcaga gaaaagtaaa atatttttgc aaagagtata agttcagata tttgatttct
961 attcgttcag aagattattt ggcaatgatt aaatactctt gtgggctaat tgggaactcc
1021 tcctctggtt taattgaggt tccatcttta aaagttgcaa caattaacat tggtgatagg
1081 cagaaaggcc gtgttcgtgg agccagtgta atagatgtac ccgttgaaaa aaatgcaatc
1141 gtcagaggga taaatatatc tcaagatgaa aaatttatta gtgttgtaca gtcatctagt
1201 aatccttatt ttaaagaaaa tgctttaatt aatgctgtta gaattattaa ggattttatt
1261 aaatcaaaaa ataaagatta caaagatttt tatgacatcc cggaatgtac caccagttat
1321 gactag
[http://www.ncbi.nlm.nih.gov/sites/entrez?cmd=Retrieve&db=nucleotide&dopt=GenBank&RID=6E5CM00T01R&log%24=nucltop&blast_rank=8&list_uids=62866977 Escherichia coli polysialic acid biosynthesis protein (neuE) gene, complete cds]
61 atgacatccc ggaatgtacc accagttatg actagaaaaa aagtgctttg ttttgtcttt
121 cgttatgatt ctcatttttt agctttgaaa aatatttttg agcagataga tgttgattca
181 tatgatttat ttttttgctg cttggataat tctctacaag agtttgtaaa aaaaaattta
241 gatgaaaaga tagttgtatt ctatcctgat gactttgttt gttttttcac ttttattaat
301 attgagttta ttttttgttc aacaggaggg aaggaccttc atgaaattgt taatactgta
361 agaacaaaag ataccataat tatatcttgt tttcctggca ttgtccttac ttctcagata
421 gaagctttta tttcaaaatc taatagtcac tatttactta ttaactcccc aaaagatatt
481 aaaacgtata aaaaaatttg taaaataata ggggttcctt ttaatggaat tctttttggt
541 ccaccatgga ttaaaaatgt caatatcaat gcaaaaagtg agaattcttg tcttatcgtt
601 gatcaagtta atgaaccctt gacgccaata aagaggatag aatatgcacg ttttttgatt
661 agagtaattc agaaacatcc acatatgaat tttattttta aaactcgaaa tcctcttata
721 tcaccagact caattgtttt tgatattaag gaatacattg aacgcttcga tttgaaaaat
781 ataacattta gcgatgataa tattgattct ttaatttcta aagttgaata ttgtattaca
841 atatcttctt cggtcgcaat atattgtctg gctaataaaa ttaaggttta tttaataaat
901 ggatttaatc atacttgcaa tggacaatgt tatttttcaa gatctggact tattgttgac
961 tataataagt ttaattttaa acacattcca cgtattaaaa aaaaatggat ggaggagaac
1021 ttttattact ctagggatat tcaacataag attttgaatg atattttaaa aatgccgtca
1081 aatgttaatg ttaggacttt tggaattaaa agatctacat tgattatatt gtttttgatc
1141 ttttttaatt tctttttctc attaggacca aaaaaaataa aaacattgaa aaaaatccat
1201 aaagttttat taaggtataa gaaagatgat atttga
primer design:
gaattcgcggccgcttctag atgagtaaaaagttaataatatttggtg Length=20+28, Tm=51, GC%=21% ctgcagcggccgctactagta tcaaatatcatctttcttataccttaat Length=21+28, Tm=51, GC%=21%
[http://www.ncbi.nlm.nih.gov/nuccore/2988378 Photobacterium damsela gene for sialyltransferase 0160, complete cds]
atgaa gaaaatactg acagttctat
421 ctatttttat tctttcagcg tgtaatagtg acaataccag cttgaaagaa acggtaagct
481 ctaattctgc agatgtagta gaaacagaaa cttaccaact gacaccgatt gatgctccta
541 gctctttttt atctcattct tgggagcaaa catgtggcac acctatcttg aatgaaagtg
601 acaagcaagc gatatctttt gattttgttg ctccagagtt aaagcaagat gaaaagtatt
661 gttttacttt taaaggtatt acaggcgatc ataggtatat cacaaataca acattaactg
721 ttgttgcacc tacgctagaa gtttacatcg atcatgcatc cttaccatcg ctacagcagc
781 ttatccacat tattcaagca aaagatgaat acccaagtaa tcaacgtttt gtctcttgga
841 agcgtgtaac tgttgatgct gataatgcca ataagttaaa cattcatact tatccattaa
901 aaggcaataa tacctcacca gaaatggtgg cagcgattga tgagtatgct cagagcaaaa
961 atcgattgaa tatagagttc tatacaaata cagctcatgt ttttaataat ttaccaccta
1021 ttattcaacc tttatataat aacgagaagg tgaaaatttc tcatattagt ttgtatgatg
1081 atggttcttc tgaatatgta agtttatatc aatggaaaga tacaccaaat aagatagaaa
1141 cattagaagg tgaagtatcg cttcttgcta attatttagc aggaacatct ccggatgcac
1201 caaaaggaat gggaaatcgt tataactggc ataaattata tgacactgat tattactttt
1261 tgcgcgaaga ttaccttgac gttgaagcaa acctacatga tttacgtgat tatttaggct
1321 cttccgcaaa gcaaatgcca tgggatgaat ttgctaaatt atctgattct cagcaaacac
1381 tatttttaga tattgtgggt tttgataaag agcaattgca acaacaatat tcacaatccc
1441 cactaccaaa ctttattttt accggcacaa caacttgggc tgggggggaa acgaaagagt
1501 attatgctca gcaacaagta aatgtgatta ataatgcgat caatgaaact agcccttatt
1561 atttaggtaa agactacgat ctatttttca aggggcatcc tgctggtggc gttattaacg
1621 acatcattct tggaagcttc cctgatatga tcaatattcc agccaagatt tcatttgagg
1681 tcttgatgat gacggatatg ttgcctgata cagtagctgg tattgcgagc tctctgtact
1741 tcacaattcc tgccgataaa gttaatttta ttgtatttac ttcatctgac actattactg
1801 atcgtgaaga ggctcttaaa tcaccattag tacaagtgat gctaacgttg ggtattgtta
1861 aagaaaaaga tgttctgttc tgggctgatc ataaagtaaa ctcgatggaa gttgccattg
1921 atgaagcctg tactcggatc attgcaaagc gacaaccaac cgcgagtgat ttacgcttgg
1981 ttattgctat tatcaaaaca attactgatc ttgagcgtat tggcgatgtg gcagaaagta
2041 ttgctaaagt cgcattagag agctttagta ataagcaata taacctattg gtttctttag
2101 aatctcttgg ccagcatacg gttcgaatgc tgcatgaggt gttagatgcg tttgctcgta
2161 tggatgttaa agccgcaata gaagtgtacc aagaagatga tcgaattgat caagagtatg
2221 agtcgatagt cagacagcta atggcccata tgatggaaga tccaagctca attcctaatg
2281 taatgaaagt gatgtgggcg gcacgttcta ttgagcgagt gggtgatcgc tgtcaaaaca
2341 tttgtgagta cattatctac tttgtgaagg gtaaagacgt tcgccatacc aaaccagatg
2401 attttggtac tatgctcgat taa
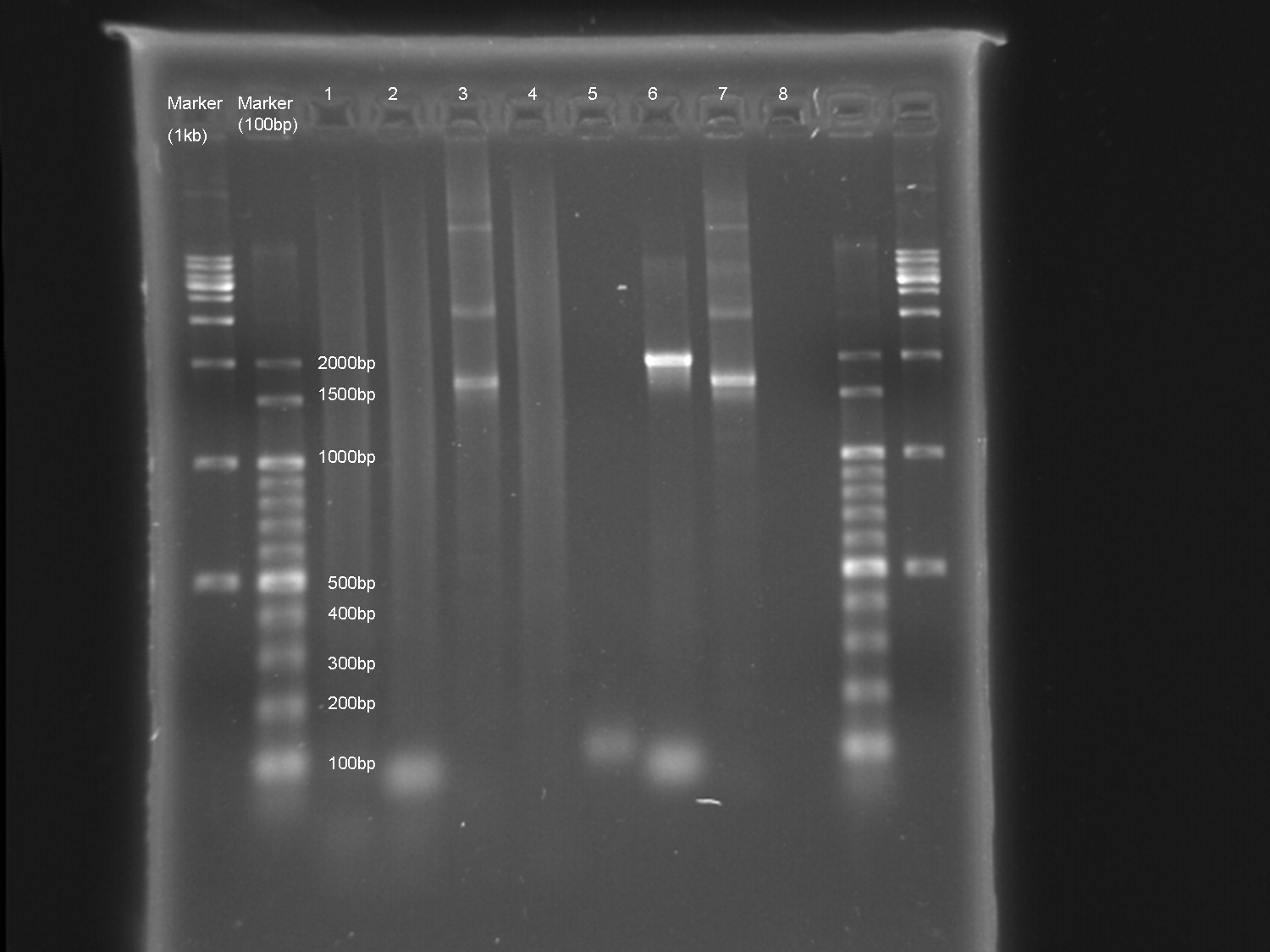
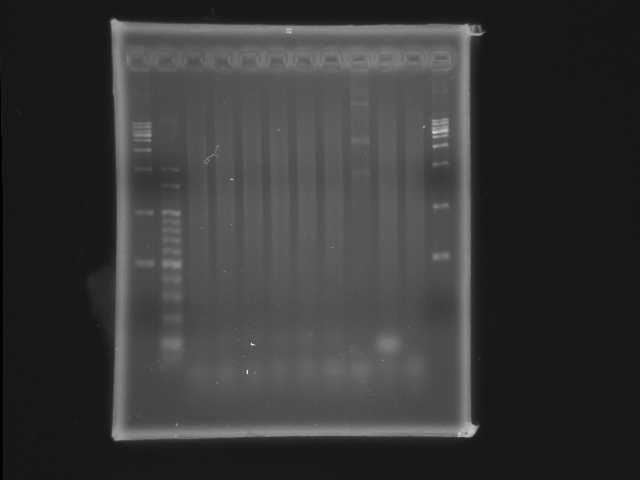
Experiment results
References
- [1]Tatsunori Iwataa, Kaori Fukuzawab, Katsuhisa Nakajimac, Sachiko Aida-Hyugaji d,Yuji Mochizukie,f, HirofumiWatanabea,f, Shigenori Tanakaa,f,:Theoretical analysis of binding specificity of influenza viral hemagglutinin to avian and human receptors based on the fragment molecular orbital method.Computational Biology and Chemistry 32 (2008) 198–211
- [2]Hai Yu, Harshal Chokhawala, Shengshu Huang, and Xi Chen:One-pot three-enzyme chemoenzymatic approach to the synthesis of sialosides containing natural and non-natural functionalities.Nat Protoc. 2006 ; 1(5): 2485–2492. doi:10.1038/nprot.2006.401.
- [3]Hai Yu, Shengshu Huang, Harshal Chokhawala,Mingchi Sun, Haojie Zheng, and Xi Chen:Highly Efficient Chemoenzymatic Synthesis of Naturally Occurring and Non-Natural a-2,6-Linked Sialosides: A P. damsela a-2,6-Sialyltransferase with Extremely Flexible Donor–Substrate Specificity.DOI: 10.1002/anie.200600572
- [4]Chin-Fen Teo,a, c Tzann-Shun Hwang,a, b, c Pei-Heng Chen,a Chih-Hung Hung,a Heau-Shan Gao,a, d Lee-Shang Chang,a, d Chun-Hung Lina, b:Synthesis of Sialyl TN Glycopeptides -- Enzymatic Sialylation by a2,6-Sialyltransferase from Photobacterium damsela.Adv. Synth. Catal. 2005, 347, 967 –972
- [5]Anne Harduin-Lepers1,2,4, Rosella Mollicone3,4,Philippe Delannoy2,4, and Rafael Oriol3,4:The animal sialyltransferases and sialyltransferase-related genes: a phylogenetic approach.Glycobiology 2005 15(8):805-817; doi:10.1093/glycob/cwi063
- [6]Bioorganic Chemistry 33 (2005) 216–228
- [7]Salvador Almagro-Moreno1,2 and E Fidelma Boyd1:Insights into the evolution of sialic acid catabolism among bacteria.BMC Evolutionary Biology 2009, 9:118doi:10.1186/1471-2148-9-118
 "
"