Team:BCCS-Bristol/Project
From 2009.igem.org
| Line 1: | Line 1: | ||
{{:Team:BCCS-Bristol/Header}} | {{:Team:BCCS-Bristol/Header}} | ||
| + | <html> | ||
<h1>VESECURE</h1> | <h1>VESECURE</h1> | ||
| - | + | </html> | |
{|align = "center" width = "60%" | {|align = "center" width = "60%" | ||
Revision as of 22:57, 20 October 2009
iGEM 2009
VESECURE
 | |
| Click above to read more about our team's modelling approach and the simulations we performed to assess the effectiveness of OMV based communication. | Click above to read more about our wet-lab work . |
Project Abstract
We aim to construct a system for directed delivery of proteins into cells by outer membrane vesicle (OMV) protein secretion. Lab work will be informed by an enhanced version of the BSim agent-based modelling framework [1], with a greatly expanded feature set and increased biological realism.
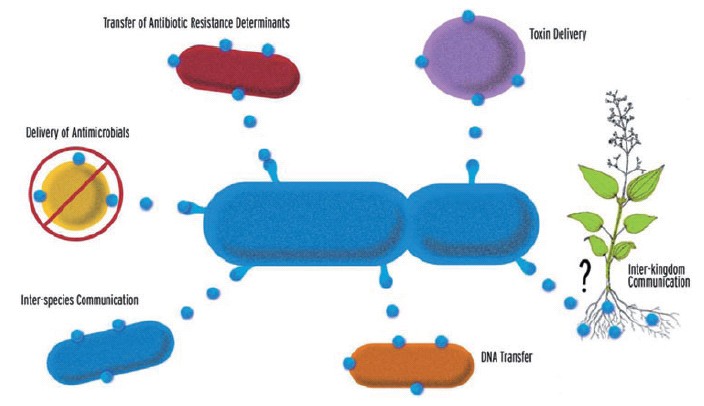
Gram-negative bacteria naturally produce outer member vesicles (OMVs): spherical, bilayered proteolipids from 20-200nm in diameter. OMVs can carry outer membrane, periplasmic and cytoplasmic proteins, DNA, RNA and other factors associated with virulence. They have been implicated in the delivery of toxins to host cells, in the transfer of proteins and genetic materials between bacterial cells and in cell-to-cell signalling [2]. Synthetic biology offers the potential to engineer these mechanisms for human benefit, for example, the injection of toxic cocktails of proteins into cancerous cells or the replacement of proteins missing as a result of genetic defects [3]. These systems may be further enhanced by localised delivery of OMVs, for example, via the control of magnetotactic bacteria using external magnetic fields [5].
We intend to design Biobricks allowing the secretion of any protein in OMVs via fusion with novel, non-toxic partners known to be enhanced in OMVs. We hope that the Biobricks will become widely used as a simple and effective means of protein secretion, whilst also conveying the unique engineering advantages of vesicular encapsulation.
The mechanisms of magnetotaxis and OMV secretion will be implemented into BSim, the award-winning modelling framework developed by last year’s Bristol team, to assist in experimental design and to test speculative ideas. Another key new feature will allow users to ‘plug in’ gene regulatory network models and see their effect at the population level.
References
[1] https://2008.igem.org/Team:BCCS-Bristol/Modeling, true 30th July 2009
[2] Global proteomic profiling of native outer membrane vesicles derived from Escherichia coli, doi:10.1002/pmic.200700196
[3] Engineered Bacterial Outer Membrane Vesicles with Enhanced Functionality, doi:10.1016/j.jmb.2008.03.076
[4] Special delivery: vesicle trafficking in prokaryotes, doi:10.1111/j.1365-2958.2006.05272.x
[5] Controlled manipulation and actuation of micro-objects with magnetotactic bacteria, doi:10.1063/1.2402221
 "
"