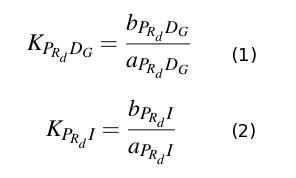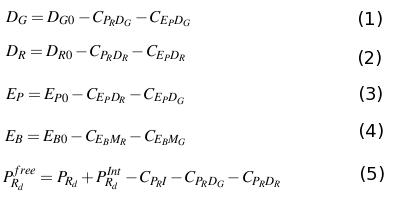Team:Bologna/Modeling
From 2009.igem.org
| HOME | TEAM | PROJECT | SOFTWARE | MODELING | WET LAB | PARTS | HUMAN PRACTICE | JUDGING CRITERIA |
|---|
A. Einstein
Contents |
Introduction
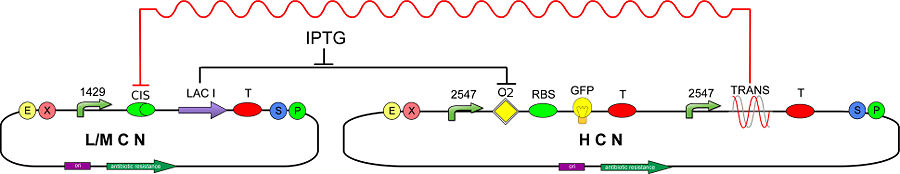
We developed a mathematical model to simulate the response of the testing circuit (Fig. 1).
Mathematical Model
Transcription and translation processes are considered similar to a second order kinetics like an enzymatic reaction: RNA polymerase and ribosome perform enzymes' role, while gene promoter and RBS sequence act as substrates. The binding between enzyme and substrate leads to the formation of a complex, yielding to the final product: mRNA for the polymerase-promoter complex and protein for the ribosome-RBS complex.
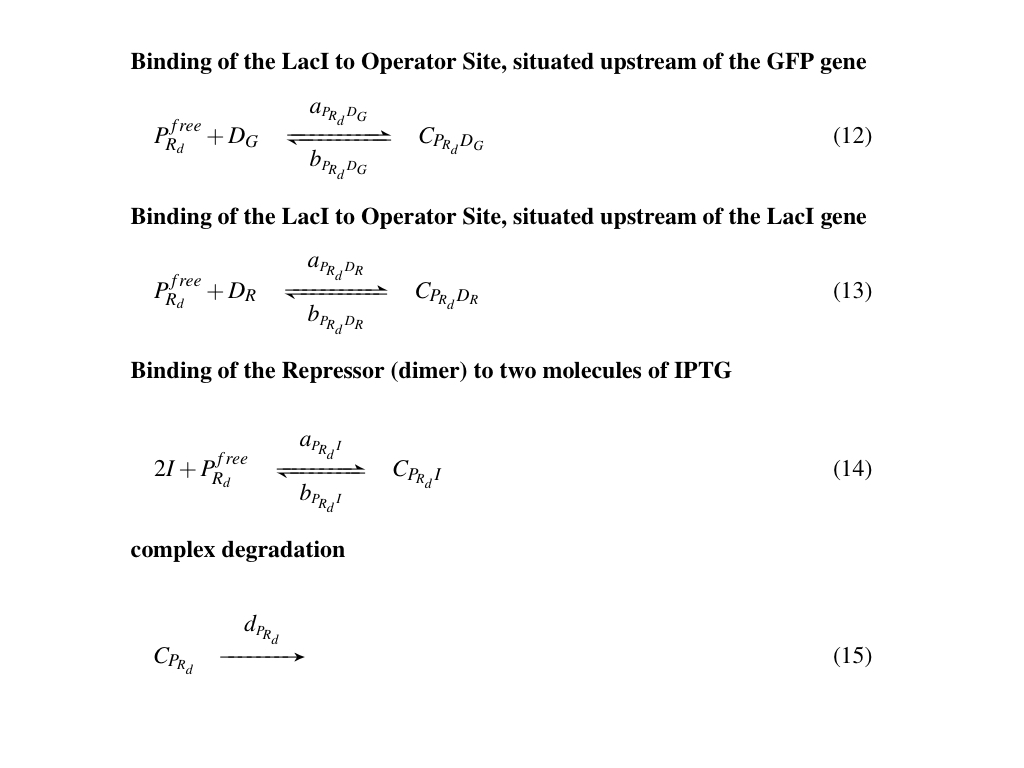
Reactions
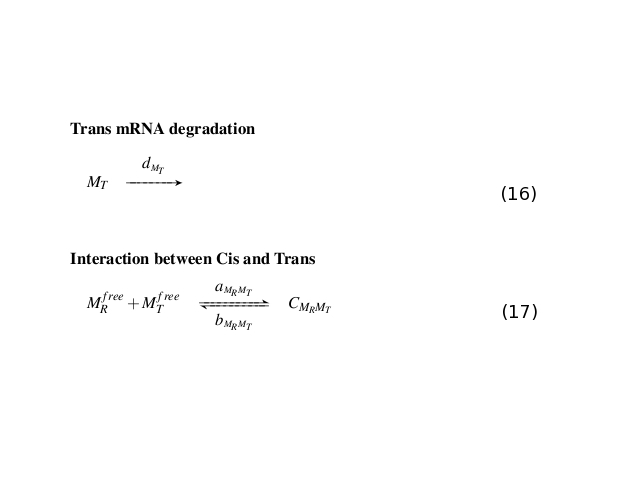
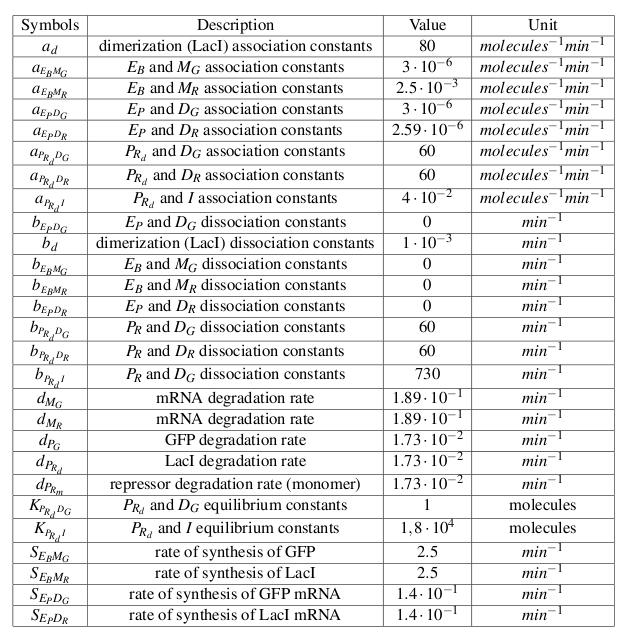
All the biochemical reactions occurring in the testing circuit are listed in Fig. 2, Fig. 3 and Fig. 4
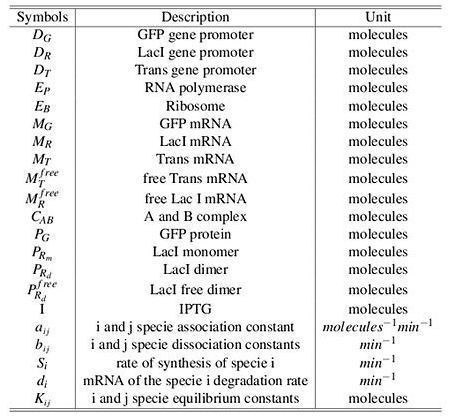
Symbol definitions are listed in Table 1
Differential Equations
The differential equations describing the above biochemical reaction are obtained appling the law of mass action.
Simulations
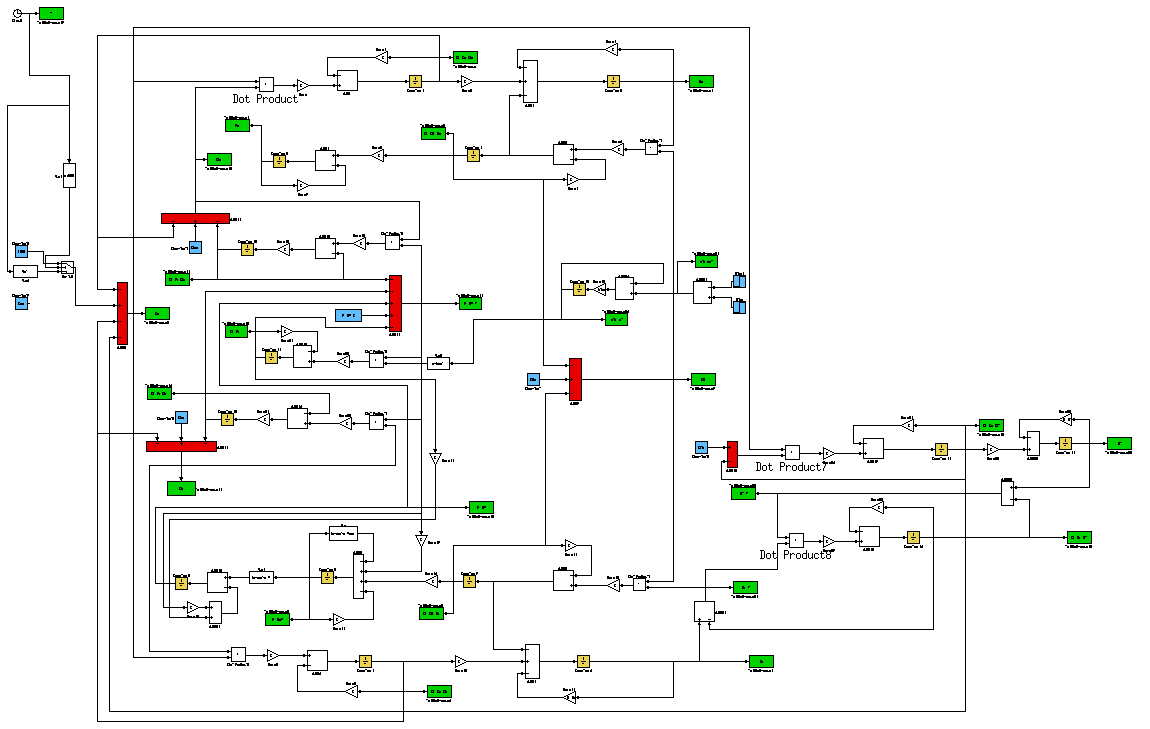
To simulate the model we implemented the equation in Simulink (Figure 3 and Figure 4).
T-REX device
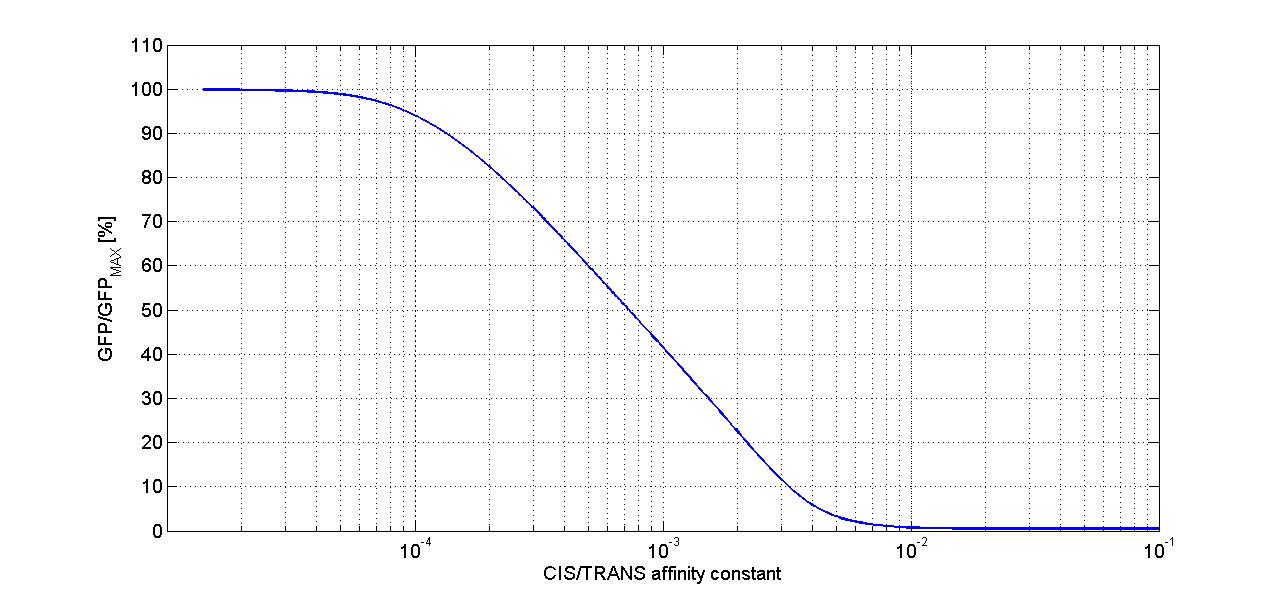
In the below figure there's the T-REX device behaviour simulated with the mathematical model. In particular the figure number 10 outlines how the affinity between CIS and TRANS influences the production of GFP.
Results of the model simulations are shown in the wet lab parts characterization.
 "
"