Team:Bologna/Project
From 2009.igem.org
Marco.cavina (Talk | contribs) |
Marco.cavina (Talk | contribs) |
||
| Line 23: | Line 23: | ||
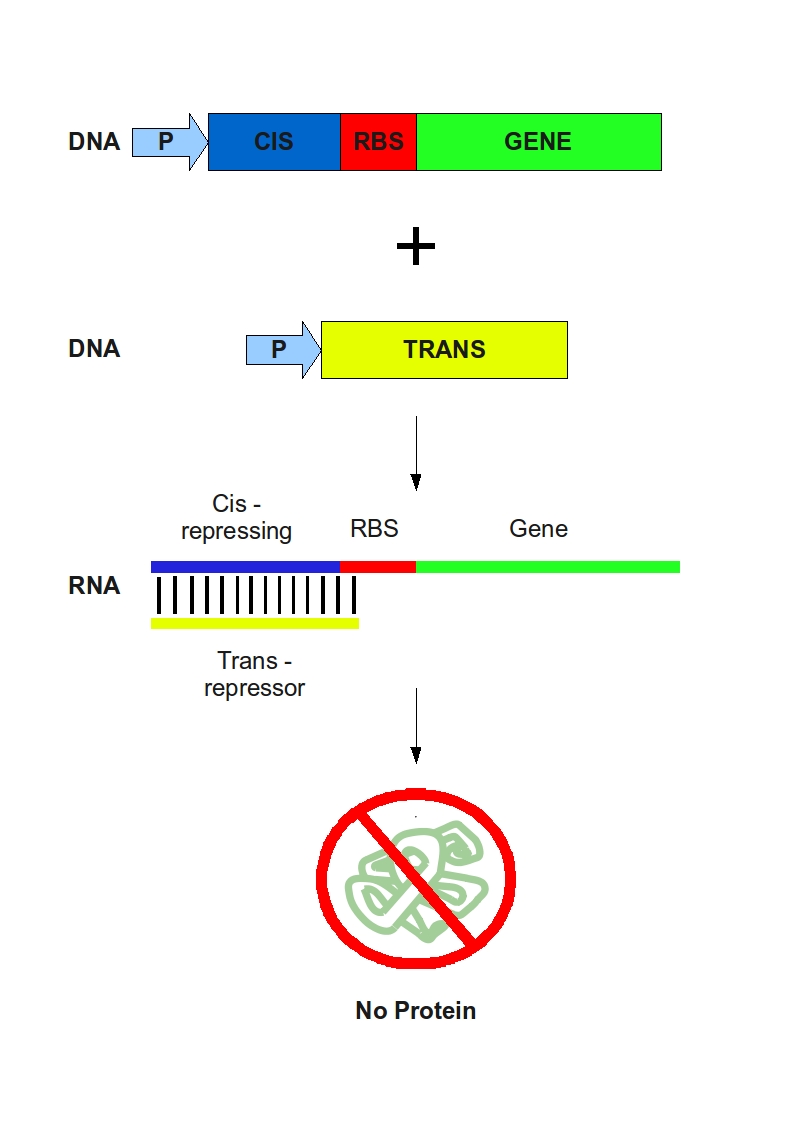
Our goal is to create a logic gate based on a post-transcriptional regulation system in Escherichia coli, using RNA to silence gene expression. We inserted a cis-acting sequence directly upstream of the rbs (ribosome binding site) to realize a "regulated rbs". We also designed the complementary trans-repressing sequence whose function is to recognize and cover the "regulated rbs" and prevent translation from it. Two versions of the trans- repressing sequence were designed with 2 different kinds of rbs covers. We want to use this short non-coding RNA segment placed in trans, to silence translation from dowstream the cis-acting sequence. | Our goal is to create a logic gate based on a post-transcriptional regulation system in Escherichia coli, using RNA to silence gene expression. We inserted a cis-acting sequence directly upstream of the rbs (ribosome binding site) to realize a "regulated rbs". We also designed the complementary trans-repressing sequence whose function is to recognize and cover the "regulated rbs" and prevent translation from it. Two versions of the trans- repressing sequence were designed with 2 different kinds of rbs covers. We want to use this short non-coding RNA segment placed in trans, to silence translation from dowstream the cis-acting sequence. | ||
We developed a [[Team:Bologna/Software|bioinformatic tool]] to research the best sequences. Using the results of our software we changed some base pairs in order to minimize the free energy of the [[RNA secondary structures]]. | We developed a [[Team:Bologna/Software|bioinformatic tool]] to research the best sequences. Using the results of our software we changed some base pairs in order to minimize the free energy of the [[RNA secondary structures]]. | ||
| + | These are the result sequences: | ||
| + | <br> | ||
{| align="center" | {| align="center" | ||
Revision as of 09:43, 9 September 2009
| HOME | TEAM | PROJECT | MODELING | WETLAB | DRY LAB | SUBMITTED PARTS | HUMAN PRACTICE |
|---|
Introduction
Our goal is to create a logic gate based on a post-transcriptional regulation system in Escherichia coli, using RNA to silence gene expression. We inserted a cis-acting sequence directly upstream of the rbs (ribosome binding site) to realize a "regulated rbs". We also designed the complementary trans-repressing sequence whose function is to recognize and cover the "regulated rbs" and prevent translation from it. Two versions of the trans- repressing sequence were designed with 2 different kinds of rbs covers. We want to use this short non-coding RNA segment placed in trans, to silence translation from dowstream the cis-acting sequence.
We developed a bioinformatic tool to research the best sequences. Using the results of our software we changed some base pairs in order to minimize the free energy of the RNA secondary structures.
These are the result sequences:
| Cis-repressing sequence, inserted upstream of the target gene | |||
| Scar | Cis-repressing | Rbs | Scar |
| TACTAGAG | AACACAAACTATCACTTTAACAACACATTACATATACATTAAAATATTAC | AAAGAGGAGAAA | TACTAGAG |
| Trans-repressor sequence with a cover of 7 bases (long version) | |||
| Scar | Cover | Trans(long) | Scar |
| TACTAGAG | CCTCTTT | GTAATATTTTAATGTATATGTAATGTGTTGTTAAAGTGATAGTTTGTGTT | TACTAGAG |
| Trans-repressor sequence with a cover of 4 bases (short version) | |||
| Scar | Cover | Trans(short) | Scar |
| TACTAGAG | CTTT | GTAATATTTTAATGTATATGTAATGTGTTGTTAAAGTGATAGTTTGTGTT | TACTAGAG |
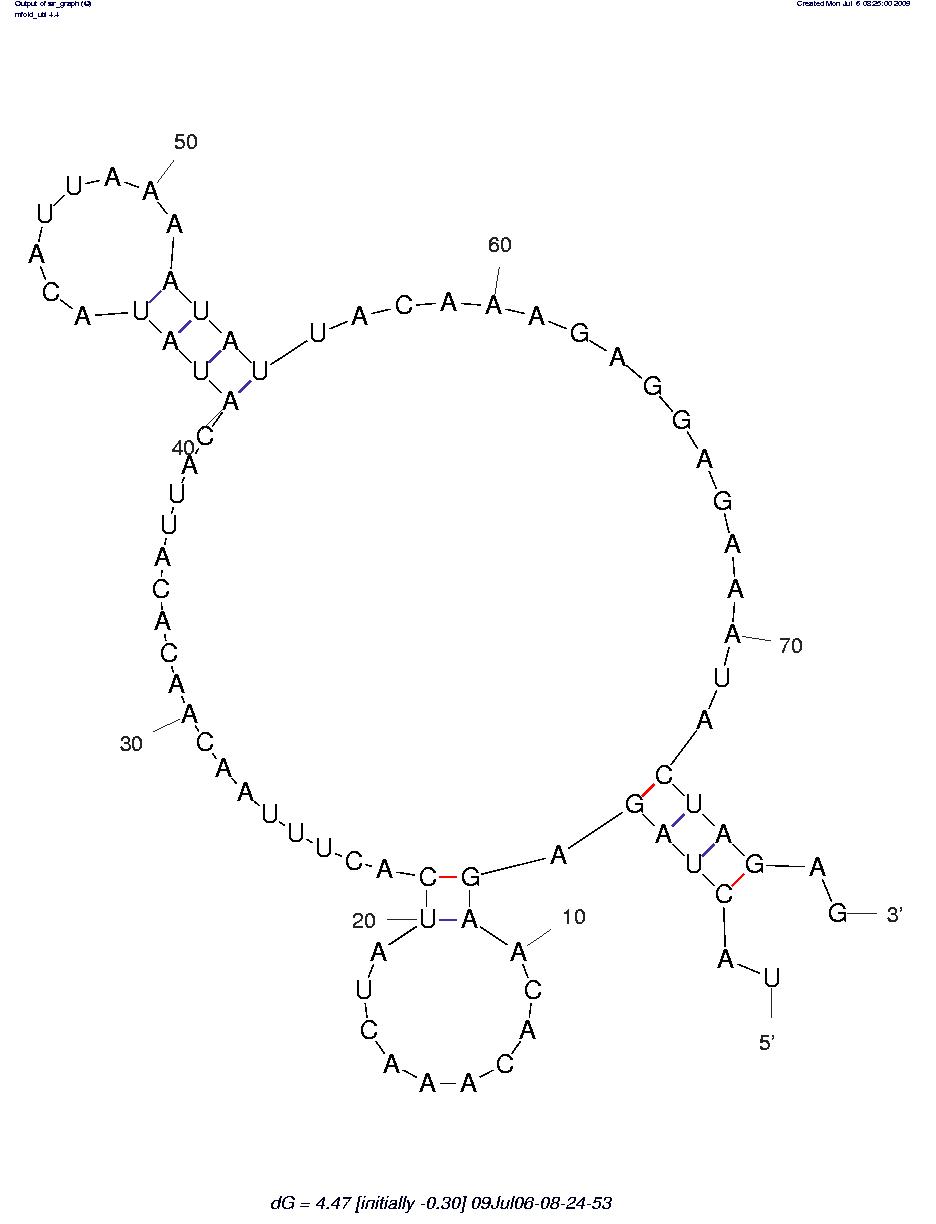
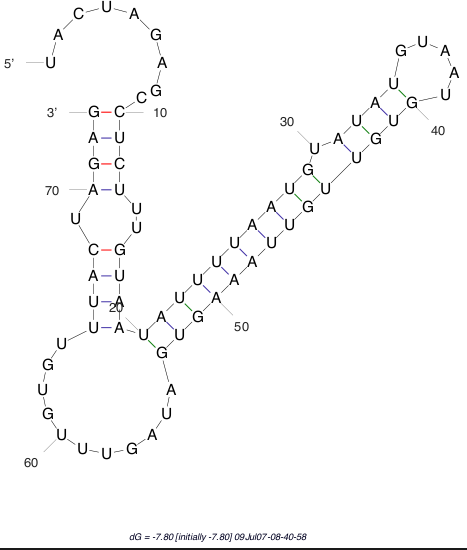
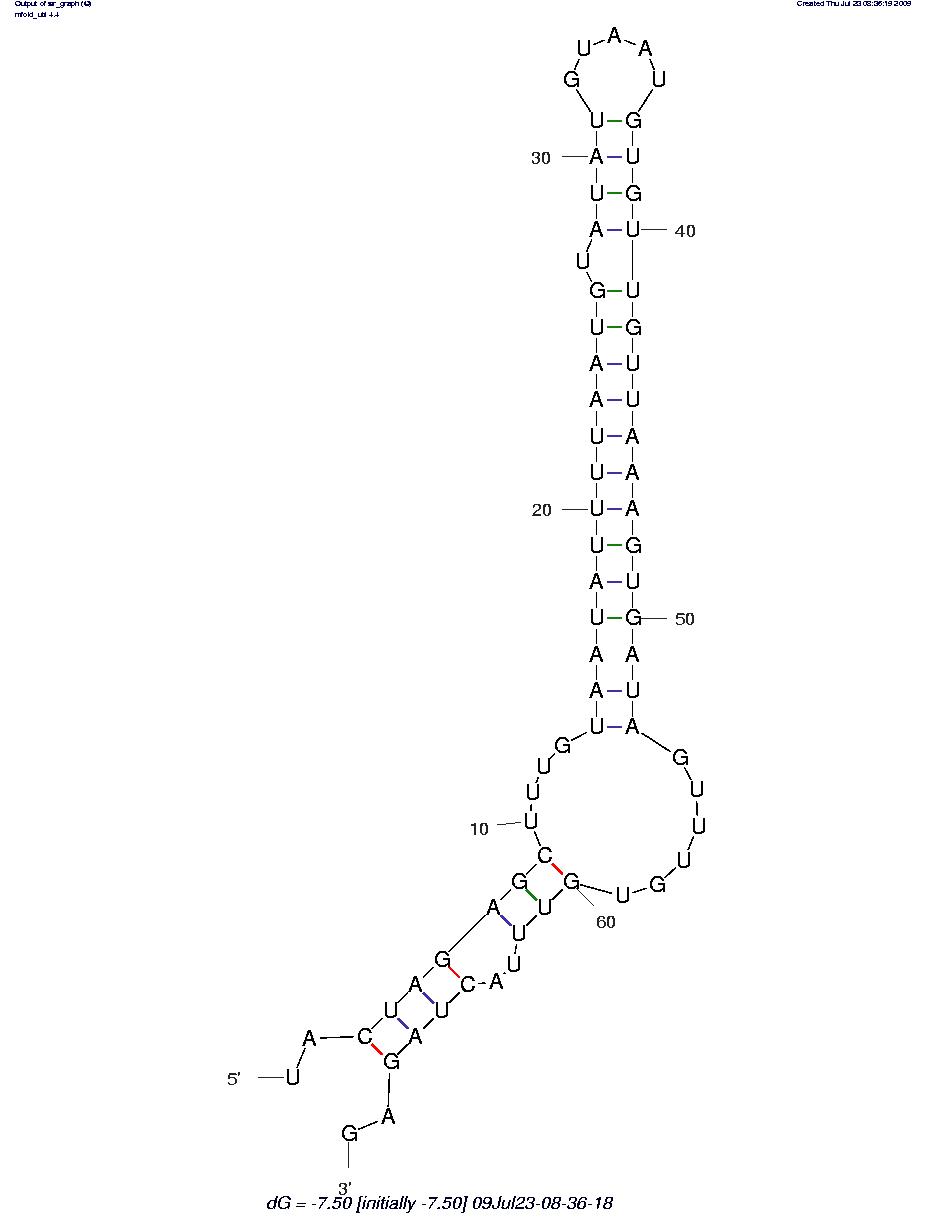
The sequence secondary structures of RNAs are obtained using the RNA mfold tool:
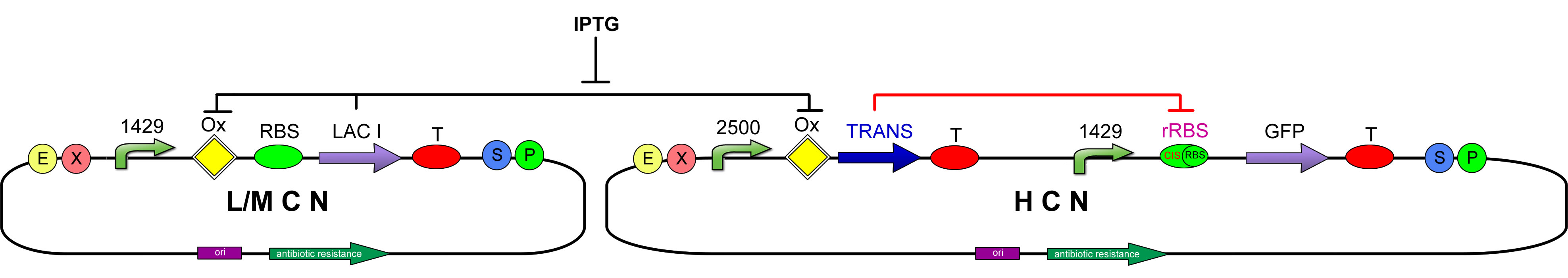
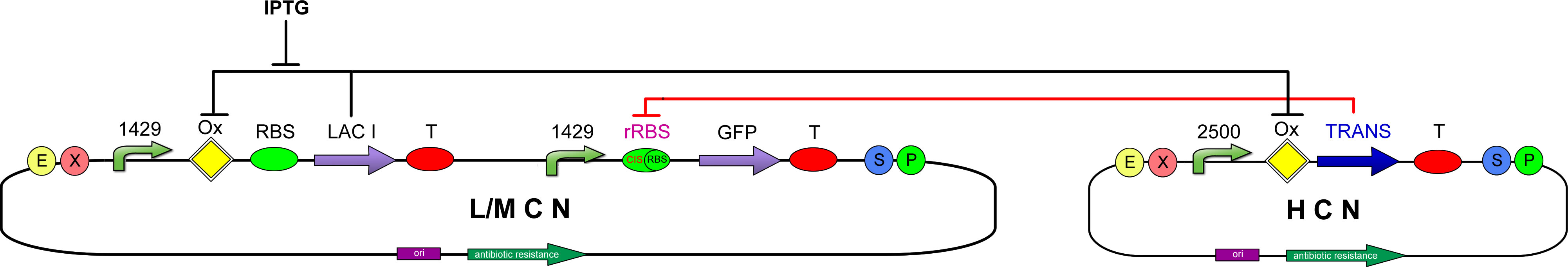
These are the three genetic circuits that we are currently studying:
 "
"