Team:Bologna/Project
From 2009.igem.org
| Line 7: | Line 7: | ||
<br><br> | <br><br> | ||
<font face="Calibri" font size="4" color="#000000"> | <font face="Calibri" font size="4" color="#000000"> | ||
| - | The project aims to | + | The project aims to design a new device to control the synthesis of a protein in Escherichia coli regardless of the protein to be controlled. This "general-purpose" standard device acts on translation to allow a faster silencing of protein expression if compared to standard regulated promoters. We named this device <b>T-Rex</b> (<b>T</b>rans <b>R</b>epressor of <b>Ex</b>pression). <br>It consists of two new BioBricks: |
| - | + | ||
<br><br> | <br><br> | ||
<ul> | <ul> | ||
| - | <li><font color="#000080"><b>CIS-repressing</b></font>, to be assembled upstream of the target protein coding sequence | + | <li><font color="#000080"><b>CIS-repressing</b></font>, to be assembled upstream of the target protein coding sequence. It contains a ribosomal binding site <font color="#228b22"><b>(RBS)</b></font>; |
</ul> | </ul> | ||
<br> | <br> | ||
<ul> | <ul> | ||
| - | <li><font color="#000080"><b>TRANS-repressor</b></font>, to | + | <li><font color="#000080"><b>TRANS-repressor</b></font>, complementary to the CIS-repressing and placed under the control of a different promoter. In order to have a better repression, it contains also a <font color="#228b22"><b>RBS cover</b></font> in two versions of different length (4 and 7 nucleotides). <br>The longer version covers also 3 nucleotides of the Shine-Dalgarno sequence. |
</ul> | </ul> | ||
<br> | <br> | ||
Revision as of 16:35, 21 October 2009
| HOME | TEAM | PROJECT | SOFTWARE | MODELING | WET LAB | PARTS | HUMAN PRACTICE | JUDGING CRITERIA |
|---|
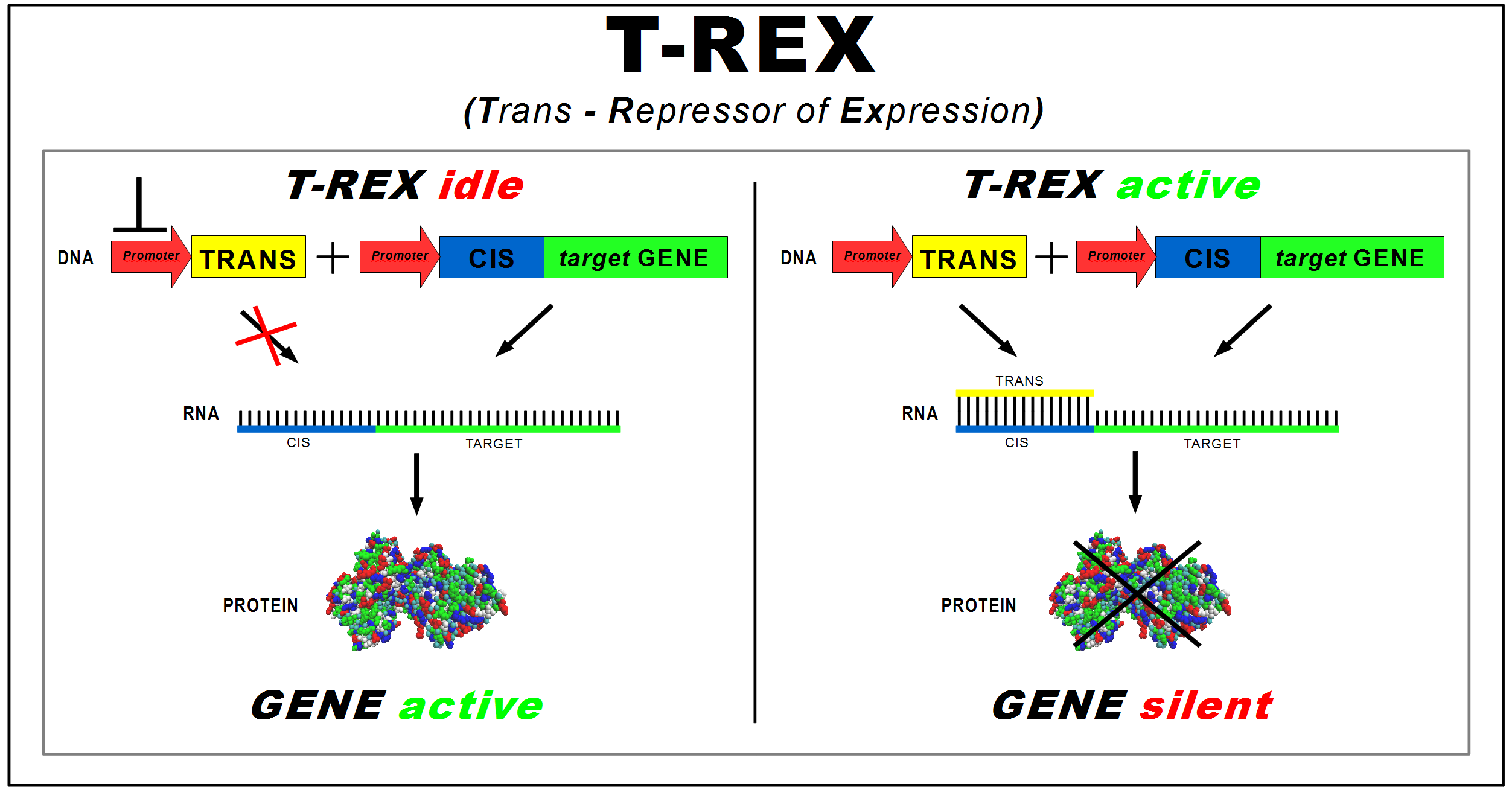
(Trans-Repressor of Expression)
The project aims to design a new device to control the synthesis of a protein in Escherichia coli regardless of the protein to be controlled. This "general-purpose" standard device acts on translation to allow a faster silencing of protein expression if compared to standard regulated promoters. We named this device T-Rex (Trans Repressor of Expression).
It consists of two new BioBricks:
- CIS-repressing, to be assembled upstream of the target protein coding sequence. It contains a ribosomal binding site (RBS);
- TRANS-repressor, complementary to the CIS-repressing and placed under the control of a different promoter. In order to have a better repression, it contains also a RBS cover in two versions of different length (4 and 7 nucleotides).
The longer version covers also 3 nucleotides of the Shine-Dalgarno sequence.
Transcription of the target gene produces a mRNA strand, starting with the Cis element, which is translated into proteins by ribosome. Trans’ promoter induction produces a transcript that binds with the Cis part. The RNA duplex prevents ribosome from binding to RBS, repressing protein synthesis. Thus, the TRANS-repressor amount regulates the gene mRNA translation rate.
The Parts
We choose to use 50bp non-coding sequences as CIS-repressing and TRANS-repressor complementary parts, and we developed a bioinformatic tool (BASER) to design them, in order to minimize unwanted interactions with genomic mRNA, reduce the probability of intra-molecular annealing and of partial/shifted hybridization.
| CIS-repressing | |||
| Prefix | non-coding TRANS target | RBS | Suffix |
| GAATTCGCGGCCGCTTCTAGAG | AACACAAACTATCACTTTAACAACACATTACATATACATTAAAATATTAC | AAAGAGGAGAAA | TACTAGTAGCGGCCGCTGCAG |
| TRANS-repressor (4) | |||
| Prefix | RBS cover | non-coding TRANS | Suffix |
| GAATTCGCGGCCGCTTCTAGAG | CTTT | GTAATATTTTAATGTATATGTAATGTGTTGTTAAAGTGATAGTTTGTGTT | TACTAGTAGCGGCCGCTGCAG |
| TRANS-repressor (7) | |||
| Prefix | RBS cover | non-coding TRANS | Suffix |
| GAATTCGCGGCCGCTTCTAGAG | CCTCTTT | GTAATATTTTAATGTATATGTAATGTGTTGTTAAAGTGATAGTTTGTGTT | TACTAGTAGCGGCCGCTGCAG |
The Genetic Circuits
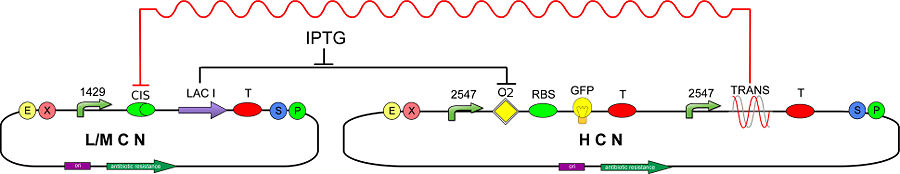
In order to test and characterize our T-REX device, we developed the following genetic circuit:
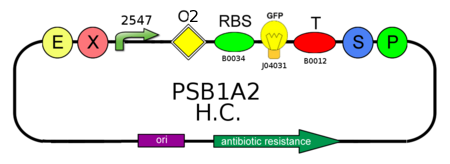
The circuit above represents the ON configuration of our T-Rex device: TRANS-repressor is constitutively expressed and CIS-repressing is assembled upstream the LacI gene (BBa_C0012), while the reporter protein GFP (BBa_J04031) is assembled under the control of another promoter, regulated by LacI natural operator O2.
CIS-repressing and TRANS-repressor mRNAs bind together, preventing LacI translation and allowing GFP production.
Using some IPTG, the repression action of T-Rex become easily, because the concentration of free LacI in the cells is reduced.
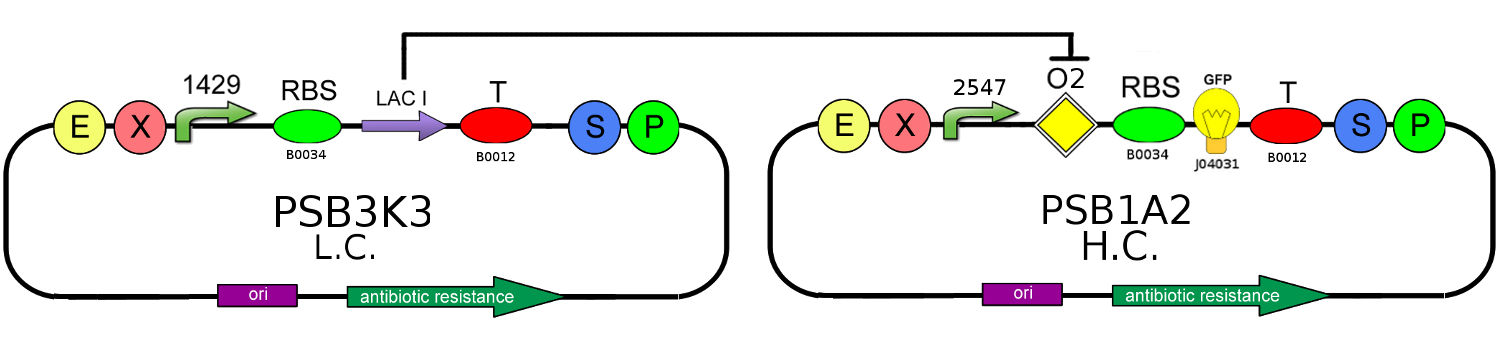
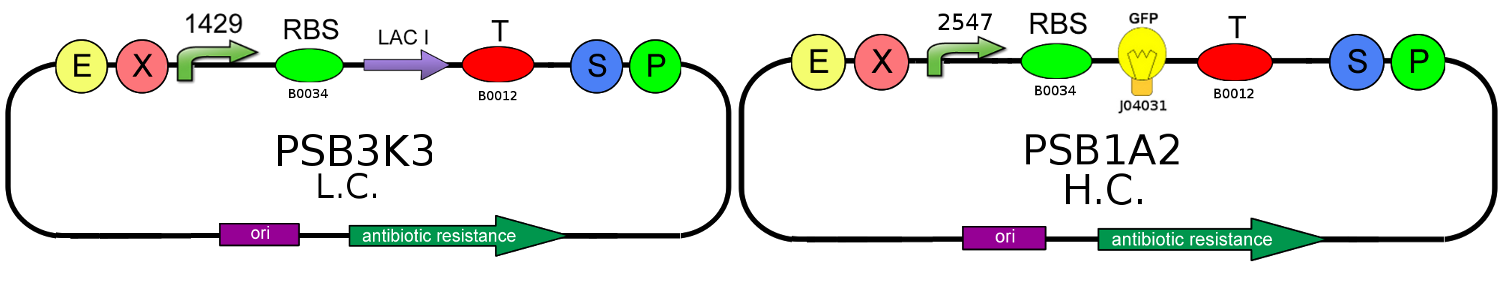
To prove T-Rex repression, GFP production level must be compared with the level produced by the OFF configuration of our T-Rex device, that is the same circuit in absence of TRANS-repressor:
In this second case, LacI is regularly translated and, binding with its O2 natural operator, it blocks GFP production.
We choose to use a low copy number plasmid and a weak promoter (J23118) for the CIS-repressing/LacI production, and a high copy number plasmid and a strong promoter (J23100) for the TRANS-repressing, to make reporter production's variations easily to be detected.
Before realizing the whole T-Rex device, we decided to perform some preliminary tests, in order to characterize every single BioBrick involved in it.
pSB1A2 vs pSB3K3
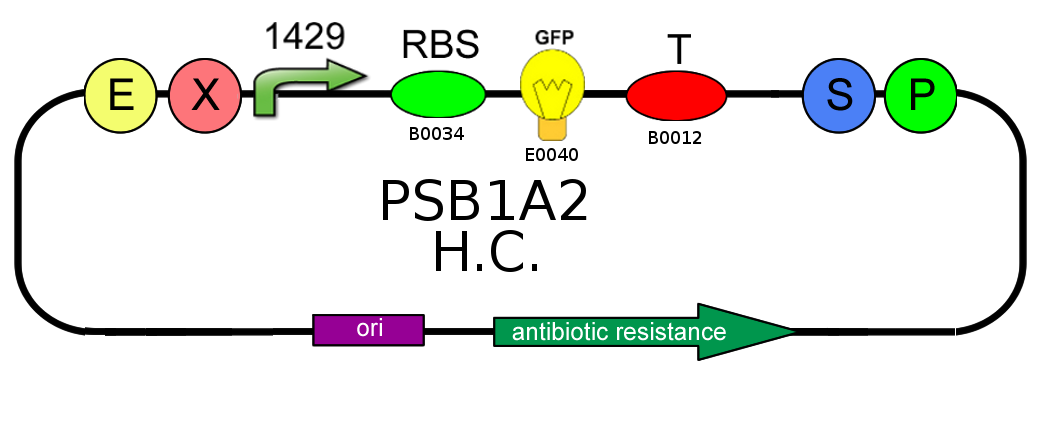
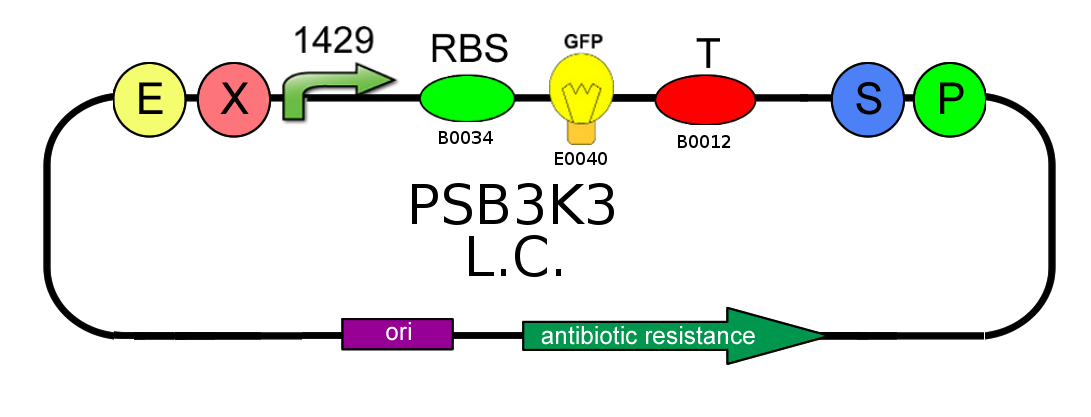
- In order to characterize the difference between the high copy number pSB1A2 and the low to medium copy number pSB3K3 plasmids, we analyzed production of BBa_I13504 (wild type GFP) under BBa_J23118 promoter (1429):
BBa_J23100 vs BBa_J23118
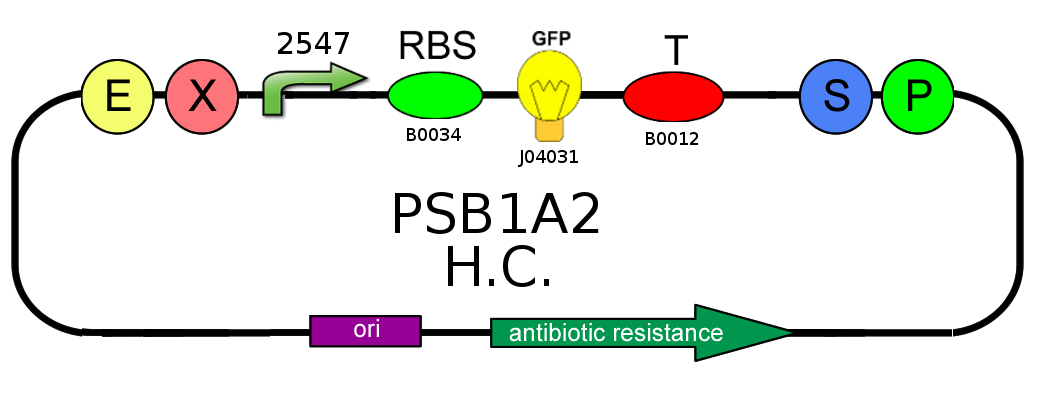
- In order to characterize the ratio between BBa_J23100 (strength 2547) and BBa_J23118 (strength 1429) promoters, we analyzed GFP (BBa_J04031) production on pSB1A2:
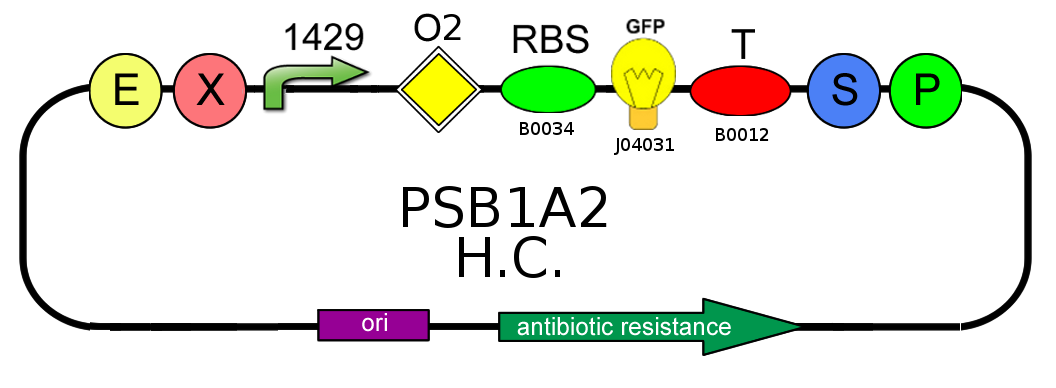
Presence vs Absence of LacI natural operator O2
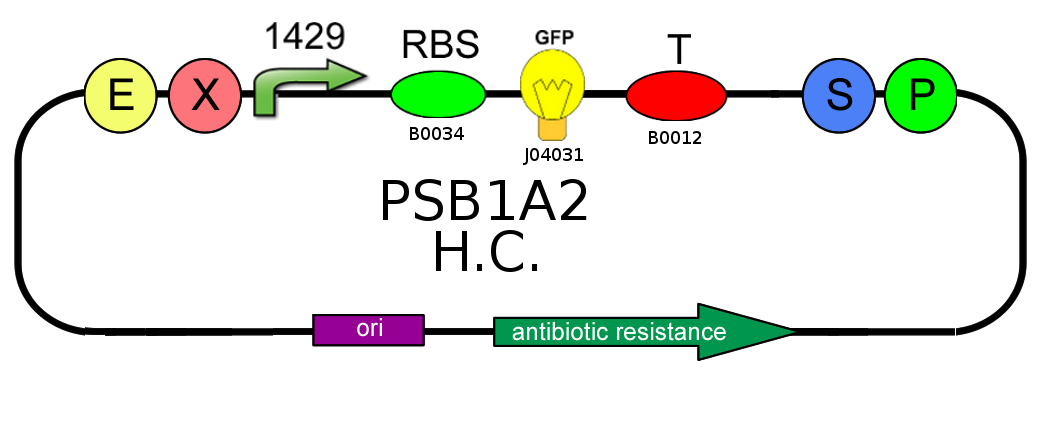
- We needed to confirm that LacI natural operator O2 don't influence GFP production when LacI repressor is not present. We evaluate this on pSB1A2, using BBa_J04031 under the control of both BBa_J23118 and BBa_J23100, in presence and in absence of BBa_K079019:
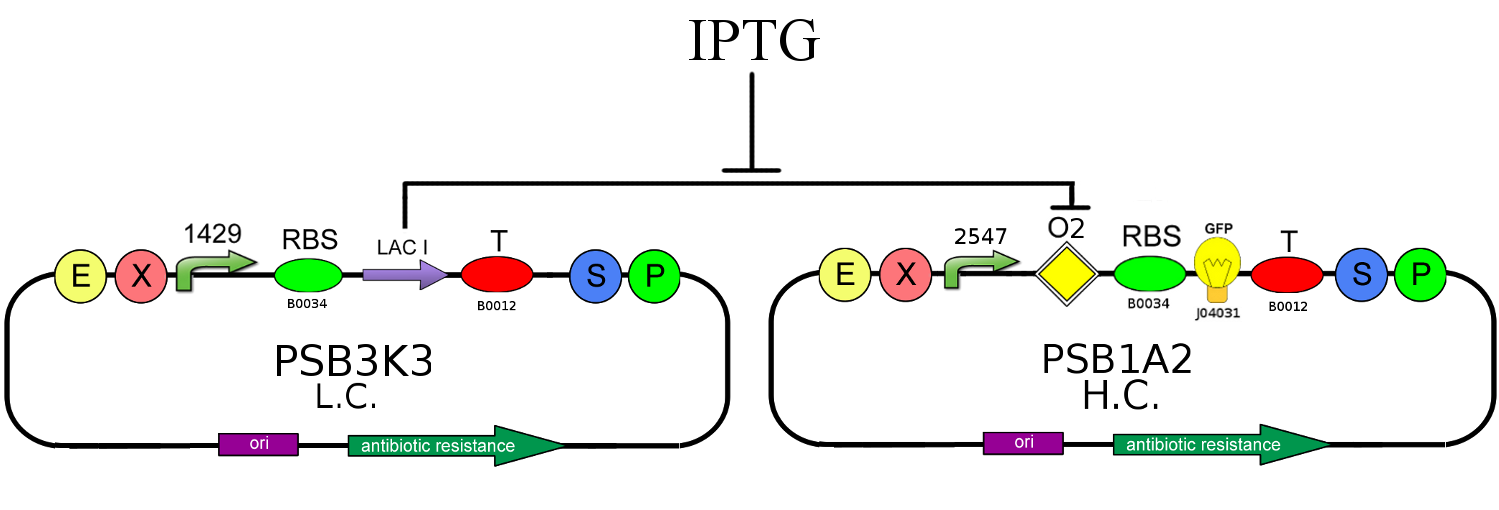
Interaction of LacI repressor with its natural operator O2
- We studied interactions between LacI repressor and its natural operator O2, analyzing this two genetic circuits:
In the first circuit, we used different IPTG concentration in order to evaluate LacI repression strength. In the latter, we verified that LacI doesn't interfere with GFP production if its natural operator O2 is not present.
 "
"