Team:Duke
From 2009.igem.org
Sahilprasada (Talk | contribs) |
(→CPEC Cloning) |
||
| Line 315: | Line 315: | ||
[[#Protocols | Back to top]] | [[#Protocols | Back to top]] | ||
| - | |||
'''Materials''' | '''Materials''' | ||
* Phusion™ High-Fidelity PCR Kit (FINNZYMES, Cat. No. F-553) | * Phusion™ High-Fidelity PCR Kit (FINNZYMES, Cat. No. F-553) | ||
Revision as of 21:06, 21 October 2009
| Home | Project | Notebook | Team |
We, the determined and motivated Duke University iGEM 2009 team of 6 undergraduate students, 3 graduate students, and 3 professors, have developed a practical, powerful, biotechnological method to deliver on the promise of sustainable green synthetic biology!
Due to the high costs and inefficiency of the process of cloning a gene, we invented a new procedure which lowers costs and increases efficiency. This method, Circular Polymerase Extension Cloning (CPEC), saves time as well, since this method does not involve ligation or restriction enzymes. Our method has been published in a peer-reviewed journal here here
We have applied our method to the production of biologically derived plastics. The rising costs of the current method of producing biodegradable plastics in an environmentally sound manner has hindered its widespread use. However, we have discovered a more efficient pathway to produce these biodegradable plastics.
One-Step Construction of a Bioplastic Production Pathway in E. coli: Abstract
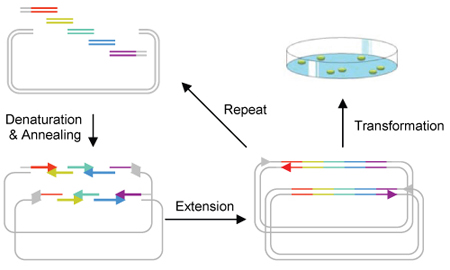
A convenient ligation-free, sequence-independent one-step plasmid assembly and cloning method is developed [Quan J, Tian J (2009) Circular Polymerase Extension Cloning of Complex Gene Libraries and Pathways. PLoS ONE 4(7): e6441]. The strategy, called Circular Polymerase Assembly Cloning (CPEC), relies on polymerase extension to assemble and clone multiple fragments into any vector. Using this method, we are able to quickly assemble a metabolic pathway consisting of multiple enzymes and regulatory elements for the production of a biocompatible as well as biodegradable plastic polymer in E. coli.What is CPEC?
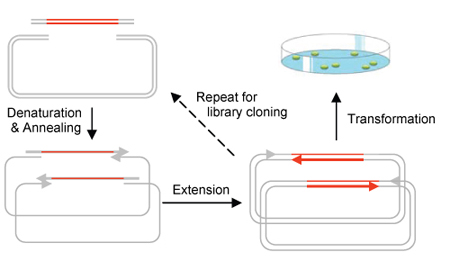
Circular Polymerase Extension Cloning (CPEC) is the development of a much simplified sequence-independent cloning technology based entirely on the polymerase extension mechanism. This method extends overlapping regions between the insert and vector fragments to form a complete circular plasmid. An extremely simple theory, CPEC piggybacks PCR in splicing genes. The gene insert is modified to have ends that overlap with the ends of the linearized vector and both have similar melting temperatures. The insert and vector are placed within a PCR machine in the absence of primers. Denaturation separates the double-stranded insert and vector and the overlapping ends anneal. Polymerase extension mechanism is then used to complete the plasmid. Using this method, we are able to quickly assemble a metabolic pathway consisting of multiple enzymes and regulatory elements for the production of a biocompatible as well as biodegradable plastic polymer in E. coli.
What are the benefits?
The process of high-throughput cloning is bottle necked at the restriction and ligation stages. A combination of high costs, requirements for restriction site specific enzymes and general inefficiency of the process makes cloning on a large combinatorial gene library inviable. Circular Polymerase Extension Cloning (CPEC) addresses this issue by eliminating the need for restriction and ligation enzymes and thereby streamlining and condensing the procedure into the duration of 5 minutes.One-Step Construction of a Bioplastic Production Pathway using CPEC
We will apply CPEC in the construction of a multi-component plasmid containing biobricks. Previous Duke iGEM projects have yielded the genes in a metabolic pathway that synthesizes poly(3HB-co-4HB), a biodegradable plastic, in E. coli. We will transform those genes into biobricks, with sticky ends, and efficiently combine them in a vector using CPEC.Biodegradable Plastic Synthesis Pathway in E. coli
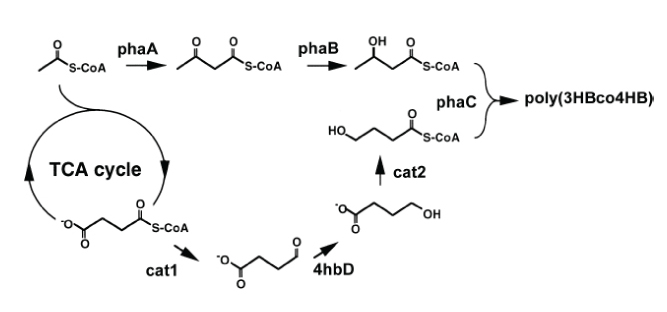
Polyhydroxyalkanoic acids (PHA), naturally occurring storage polymers found in a variety of bacteria, have received increased attention for their potential use as bioplastics that are both biodegradable and reduce reliance on petroleum-based plastics. In particular, the copolymer poly(3-hyroxybutyrate-co-4-hydroxybutyrate), or poly(3HB-co-4HB), which combines the 3HB and 4HB polymers from different bacteria (Figure 2 shows the pathway), has elastic properties ideal for a wide range of thermoplastic applications. The high cost of PHA, however, is the biggest impediment to widespread use of bioplastics. Moreover, poly(3HB-co-4HB) pathways developed so far in E. coli have yielded undesirably low and unpredictable 4HB-to-3HB ratios.
Thus, this project aims to develop a more efficient
biopathway for poly(3HB-co-4HB) while increasing the 4HB monomer composition predictably.
It was hypothesized that optimizing codon permutations of the phaC gene would
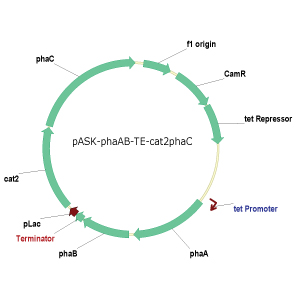
greatly increase affinity of PHA synthase to the 4HB monomer. To date, the phaCAB
and cat2 operons have been cloned into pUC19 and PCR Blunt II-TOPO vectors for
successful independent production of the 3HB and 4HB polymers (Figure 4). Ligation and transformation
into E. coli as six different recombinant constructs will soon be completed
and allow for engineering of the poly(3HB-co4HB) biopathway. Future directions
would be to test the hypothesis to see if phaC can be manipulated to increase
4HB-to-3HB composition in poly(3HB-co-4HB) and to increase efficient production
of the bioplastic by engineering the FtsZ cell division protein to allow for cells
to accumulate larger quantities of PHA granules before dividing. Ultimately, once
an optimal biopathway is found, the goal would be to explore a model for mass
production of PHA bioplastics so that novel applications of bioplastics can be
feasible economically.
Contents |
Calendar
Protocols
CPEC Cloning
Materials
- Phusion™ High-Fidelity PCR Kit (FINNZYMES, Cat. No. F-553)
- Thermocycler
Preparation
5x Phusion HF Buffer 4 ul 10 mM dNTPs 0.4 ul Vector 50 ng/1kb Insert x ng* Phusion DNA Polymerase 0.2 ul ------------------------------- H2O to 20 ul
*The amount of insert is determined so that the molar ratio for vector and insert is 1 to 2.
Procedures
98°C 30sec
10X
98°C 10 sec
Annealing** 30 sec
72°C x sec***
72°C 5min
4°C hold
** Anneal at Tm + 3°C. The Tm should be calculated with the nearest-neighbor method.
***The extension time is usually calculated according to the shortest piece with 15 sec /kb if the cloning is not complicated. For example, if there is only one insert and is shorter than the vector, say, 600 bp, then I will use 15 sec for extension. Refer to the published paper for detailed information.
DNA Purification
Back to top
DNA purification
Materials :
- E.Z.N.A Gel Purification Kit (Omega Bio-Tek, Cat No. D2500-02 )
- Water bath equilibrated to 55-65C
- Microcentrifuge capable of at least 10,000 x g
- Nuclease-free 1.5 ml centrifuge bottles
- Absolute (95%-100%) ethanol
- Protective eye-wear
- Isopropanol (for fragments < 500 bp only)
Protocol:
- Perform agarose gel electrophoresis to fractionate DNA fragments. Any type or grade of agarose may be used. It is strongly recommended, however, that fresh TAE buffer or TBE buffer be used as running buffer. Do not re-use running buffer as its pH will increase and reduce yields.
- When adequate separation of bands has occurred, carefully excise the DNA fragment of interest using a wide, clean scalpel.
- Determine the approximate volume of the gel slice by weighing it in a clean 1.5 ml microfuge tube. Assuming a density of 1 g/ml of gel, the volume of gel is derived as follows: A gel slice of mass 0.3 g will have a volume of 0.3 ml. Add equal volume of Binding Buffer (XP2). Incubate the mixture at 55C-60C for 7 min or until the gel has completely melted. Mix by shaking or inverting the tube every 2-3 minutes. Centrifuge the tube briefly to collect all the liquid to the bottom of the tube.
Note: For DNA fragment less than 500bp, add 1 sample volume of isopropanol after the addition of Binding Buffer (XP2).
- Apply up to 700 ul of the DNA/agarose solution to a HiBind® DNA spin column assembled in a clean 2 ml collection tube (provided) and centrifuge in a microcentrifuge at 8,000-10,000 x g for 1 min at room temperature. Discard the liquid. Re-use the collection tube in Steps 5-8. For volumes greater than 700 ul, load the column and centrifuge successively, 700 ul at a time. Each HiBind® spin-column has a total capacity of 25-30 ug DNA.
- Discard liquid and add 300ul Binding Buffer. Centrifuge at 10,000 x g for 1 minutes.
- Add 700 ul of SPW Buffer diluted with absolute ethanol into the column and wait 2-3 minutes. Centrifuge at 10,000 x g for 1 min at room temperature to wash the sample.
- Discard liquid and repeat step 6 with another 700 ul SPW Buffer.
- Discard liquid and, re-using the collection tube, centrifuge the empty column for 1 min at maxi speed (>13,000 x g) to dry the column matrix. This drying step is critical for good DNA yields.
- Place column into a clean 1.5 ml microcentrifuge tube (not provided). Add 30-50 ul depending on desired concentration of final product) Elution Buffer (or sterile deionized water) directly to the center of the column matrix, then incubate for 1 minute. Centrifuge 1 min at maxi speed (>13,000 x g) to elute DNA. This represents approximately 70% of bound DNA. An optional second elution will yield any residual DNA, though at a lo
PCA (Polymerase Cycle Assembly)
Materials
- Phusion™ High-Fidelity PCR Kit (FINNZYMES, Cat. No. F-553)
- Thermocycler
Preparation
| 5x Phusion HF Buffer | 5 ul |
| 10 mM dNTPs | 0.5 ul |
| Oligo mixture | 125 ng /250 ng /500 ng / |
| Phusion DNA Polymerase | 0.25 ul |
| --------------------- | -------- |
| H2O | to 25 ul |
Procedures
98°C 30sec
40X
98°C 7 sec
70-50°C slow ramp, 0.1°C/sec
50°C 30 sec
72°C 15 sec /kb
72°C 5 min
4°C hold
PCR Product Clean-up for DNA Sequencing
Materials:
- ExoSAP-IT® (usb, Cat. No. 78200)
- Thermocycler
Protocol:
- Remove ExoSAP-IT from -20°C freezer and keep on ice throughout this procedure.
- Mix 5 μl of a post-PCR reaction product with 2 μl of ExoSAP-IT for a combined 7 μl reaction volume.
- Incubate at 37°C for 15 min to degrade remaining primers and nucleotides.
- Incubate at 80°C for 15 min to inactivate ExoSAP-IT.
- The PCR product is now ready for use in DNA sequencing etc.
PCR
Materials
- Phusion™ High-Fidelity PCR Kit (FINNZYMES, Cat. No. F-553)
- Thermocycler
Preperation
| 5x Phusion HF Buffer | 10 ul |
| 10 mM dNTPs | 1 ul |
| DNA template | 1 pg – 10 ng |
| Forward primer (10 uM) | 2.5 ul |
| Reverse primer (10 uM) | 2.5 ul |
| Phusion DNA Polymerase | 0.5 ul |
| -------------------- | ----- |
| H2O | to 50 ul |
Procedure
| 98°C | 30sec | 30X |
| 98°C | 10 sec | |
| Annealing* | 30 sec | |
| 72°C | 15 sec per 1 kb | |
| 72°C | 5min | |
| 4°C | hold |
* Anneal at Tm + 3°C. The Tm should be calculated with the nearest-neighbor method.
Single Colony PCR
Back to top Materials
- Taq DNA Polymerase with Standard Taq Buffer (NEB, Cat. No. M0273)
- 10 mM dNTP Mix (NEB, Cat. No. N0447)
- Thermocycler
Preperation
| 10x Standard Taq Buffer | 2 ul |
| 10 mM dNTPs | 0.4 ul |
| Bacteria culture* | 1 ul |
| Forward primer (10 uM) | 1 ul |
| Reverse primer (10 uM) | 1 ul |
| Taq DNA Polymerase | 0.2 ul |
| H2O | to 20 ul |
*Bacteria culture refers to E. coli cultured in LB solution overnight.
Procedures
| 94°C | 5 min | 25X |
| 94°C | 15 sec | |
| Annealing ** | 30 sec | |
| 72°C | 1 min /1 kb | |
| 72°C | 5min | |
| 4°C | hold |
**Anneal at Tm which is calculated with salt-adjusted method.
Transformation
Materials:
- GC5 Chemical Competent Cells (Genesee Scientific, Cat. No. 42-653)
- SOC Medium (Sigma, Cat. No. S1797)
- LB Agar (Sigma, Cat. No. L3027)
- Petri Dishes (VWR, Cat. No. SC25373-187)
- Cell Spreader (VWR, Cat. No. 89042-018)
- 37°C incubator
- 37°C shaker
- water bath
1. Thaw 1 tube of competent cells on ice;
2. Add 3 ul of cloning product or 1-50 ng of plasmid into competent cells while stirring gently;
3. Keep the tube covered by ice for 30min;
4. Heat-shock the competent cells in water bath for 45 sec at 42°C;
5. Put the tube on ice for 2 minutes;
6. Add 450 ul of SOC medium and then put it in a 37°C shaker for 1 hour;
7. Dilute and spread an appropriate amount on an LB agar plate with the appropriate antibiotics;
8. Place the plate up-side-down in 37°C incubator for 16-18 hours (overnight).
NMR
1. Login and start NMR program.
2. Click Acqi. Load sample. Make sure spin is on and lock is off.
3. Go to lock. Decrease number of sine waves to 1 to obtain step function.
Make sure Zo is 1100, lockpower is below 30, lockgain is 36, lockphase is 352, and spin is 20.
4. Go to shim. Increase lock level.
5. Click main menu. Click set up. Click H1CDCl3.
Type ‘nt=64’ (number of scans), ‘ss=2’ (dummy scans), ‘go’, ‘lb=0.2’ (line broadening).
6. Viewing: Type ‘wft’, ‘dscale’, ‘dfp’ to display peak frequency.
7. Zooming in: Click display. Go to interactive. Type ‘cr=8p, delta=8p’ to set width of zoom.
Use cursor to set boundaries. Click to cut.
8. Printing: Type “pL pscale(0) pltext ppf page’ to print spectra with text and peak frequencies.
Students
Faculty Advisors

| Maggie Jiayuan Quan jq7(at)duke.edu Graduate Student | 
| Faisal Reza faisal.reza(at)duke.edu Graduate Student |
Sponsors

| 
| 
|
 "
"